Lab Fitness Predicts Field Ecology at Oak Ridge
CompletedResearch Question
Do lab-measured fitness effects under contaminant stress predict the field abundance of Fitness Browser organisms across Oak Ridge groundwater sites with varying geochemistry?
Overview
The ENIGMA CORAL database contains 108 groundwater samples from Oak Ridge FRC with both geochemistry measurements (uranium, chromium, nickel, zinc, iron) and 16S amplicon community composition. Several Fitness Browser organisms are detected in these communities (Pseudomonas, Desulfovibrio, Sphingomonas, Caulobacter). This project bridges lab-measured fitness (Fitness Browser, 48 organisms) with real field ecology (ENIGMA CORAL, 108 sites) by testing whether organisms that tolerate metals in the lab are more abundant at contaminated field sites. See REPORT.md for full findings and interpretation.
Key Findings
14 of 26 Fitness Browser Genera Detected at Oak Ridge
Of 26 unique genera represented in the Fitness Browser, 14 are detected in Oak Ridge groundwater communities via 16S amplicon sequencing. The most prevalent are Sphingomonas (93% of 108 sites), Pseudomonas (91%), and Caulobacter (82%). Desulfovibrio, the primary ENIGMA model organism, is detected at only 34% of sites at very low abundance (max 0.09% relative abundance).
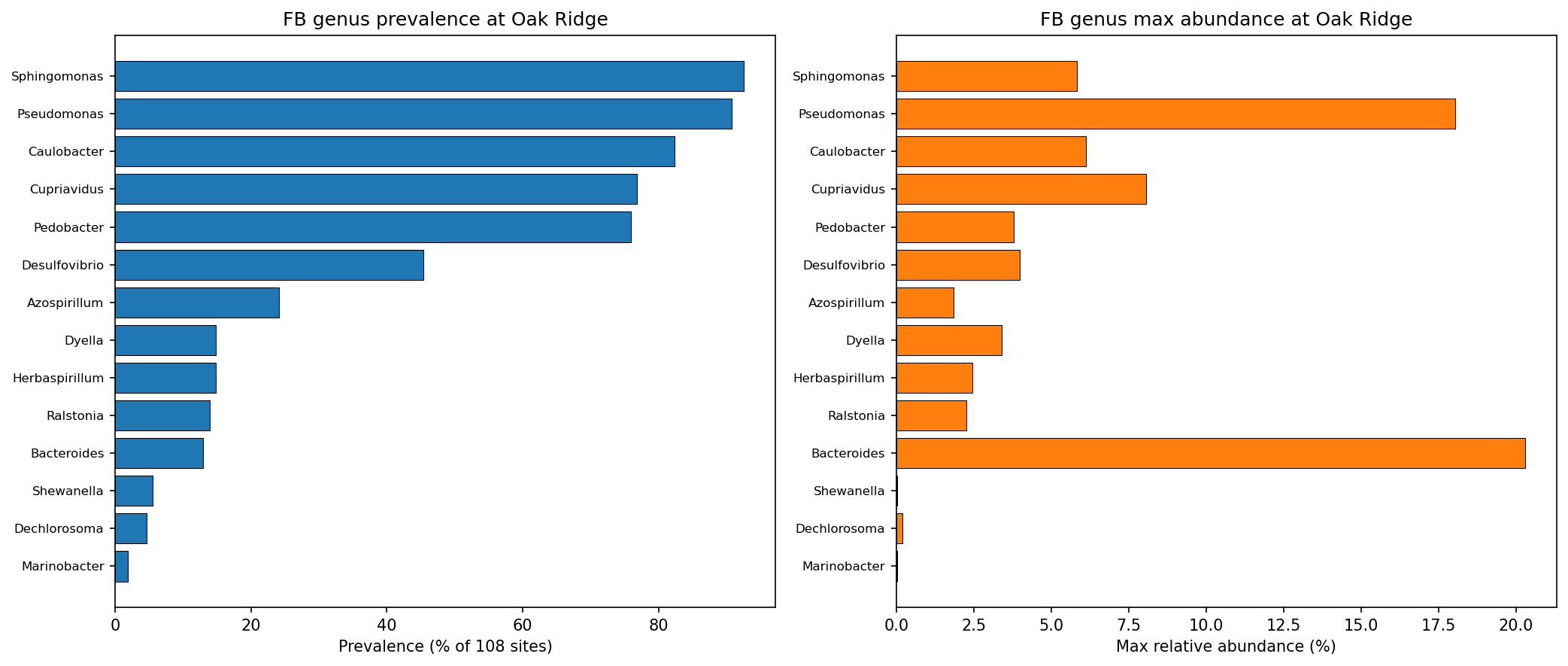
(Notebook: 02_genus_abundance.ipynb)
Genus Abundance Correlates with Uranium -- in Both Directions
Of 14 FB genera detected at Oak Ridge, 11 had sufficient prevalence (>=10 sites) for correlation analysis. After BH-FDR correction, 5 show significant correlations with uranium:
| Genus | Spearman rho | p-value | FDR q | Direction |
|---|---|---|---|---|
| Herbaspirillum | +0.336 | 3.8e-4 | 0.001 | More abundant at high-U sites |
| Bacteroides | +0.264 | 0.006 | 0.013 | More abundant at high-U sites |
| Caulobacter | -0.411 | 1.0e-5 | 1.1e-4 | Less abundant at high-U sites |
| Sphingomonas | -0.382 | 4.5e-5 | 2.5e-4 | Less abundant at high-U sites |
| Pedobacter | -0.266 | 0.005 | 0.013 | Less abundant at high-U sites |
Azospirillum (rho=+0.20, p=0.042) is marginal after FDR correction (q=0.077). Desulfovibrio shows no correlation (rho=0.022, p=0.82) and Pseudomonas shows no correlation (rho=-0.059, p=0.55) despite both being ENIGMA model organisms. Three genera (Shewanella, Dechlorosoma, Marinobacter) were excluded due to low prevalence (<10 sites).
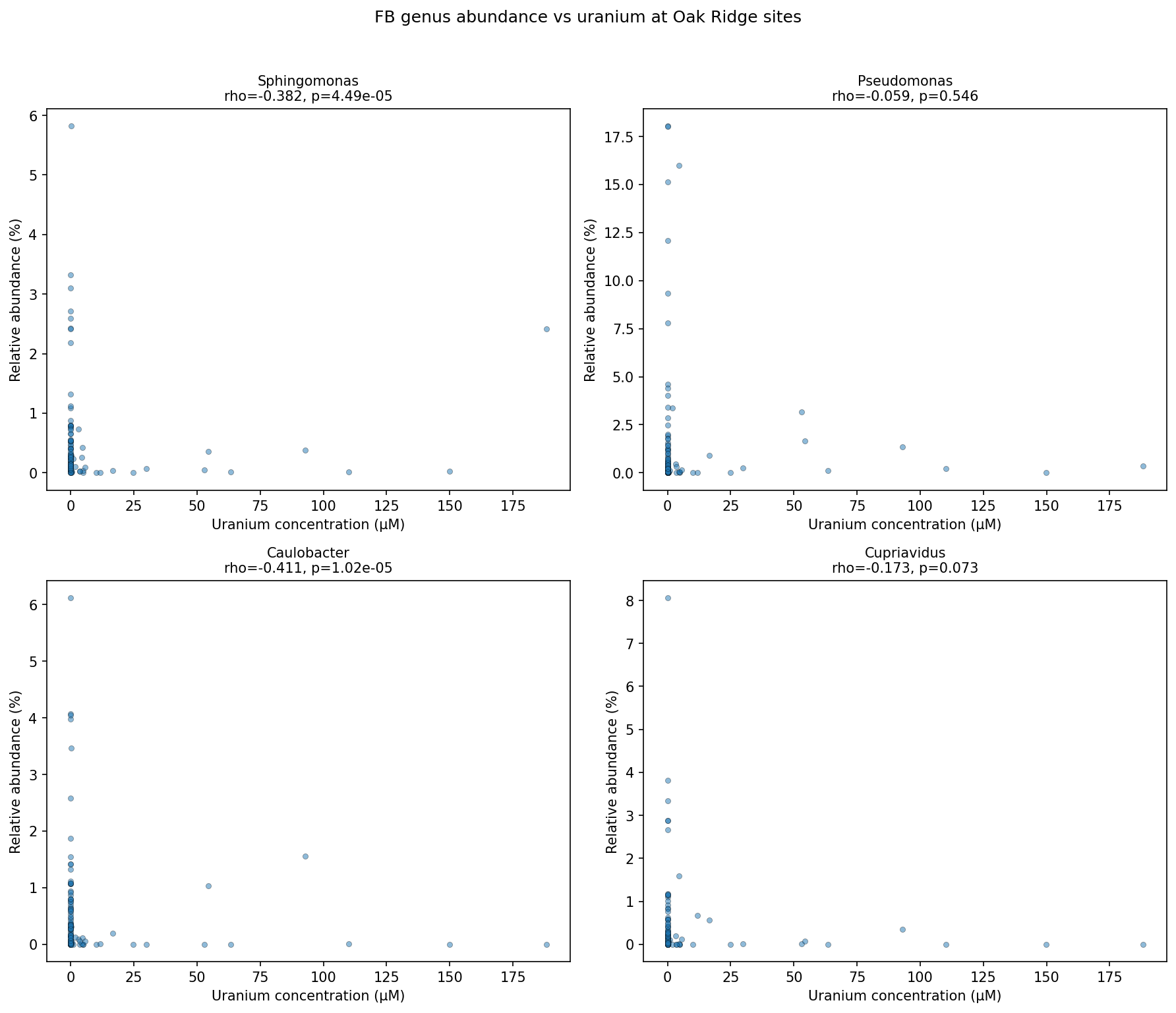
(Notebook: 03_fitness_vs_field.ipynb)
Lab Metal Tolerance Does Not Significantly Predict Field Abundance Ratio
The correlation between lab-derived metal tolerance score (negative mean fitness under stress, higher = more tolerant) and the high-uranium/low-uranium field abundance ratio is suggestive but not significant (Spearman rho=0.503, p=0.095, n=12 genera). The trend is in the predicted direction -- genera with higher lab tolerance tend to have higher abundance at contaminated sites -- but statistical power is limited by the small number of genera.
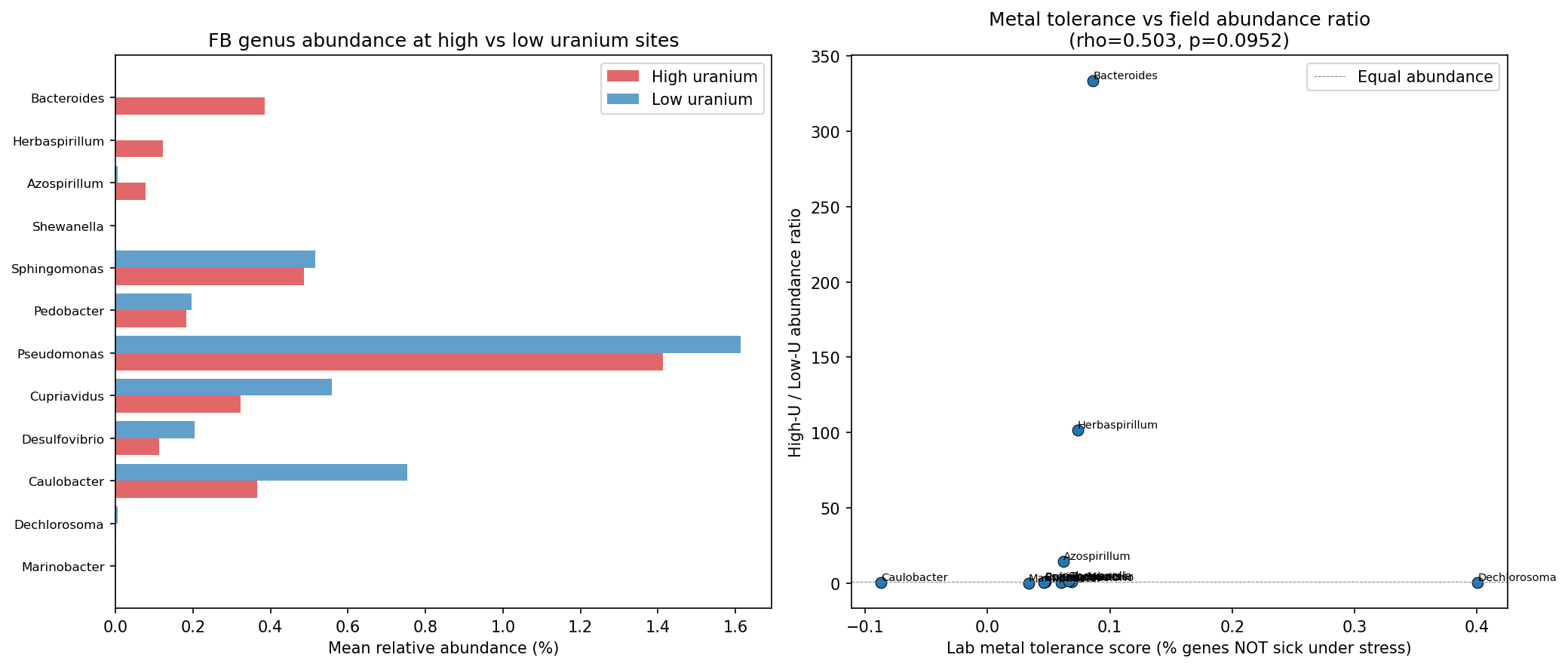
(Notebook: 03_fitness_vs_field.ipynb)
Community Composition Shifts with Contamination
Sites split at the median uranium concentration show distinct community compositions. The top genera at high-uranium sites differ from low-uranium sites, with rare-biosphere taxa and subsurface specialists becoming more prominent at contaminated sites.
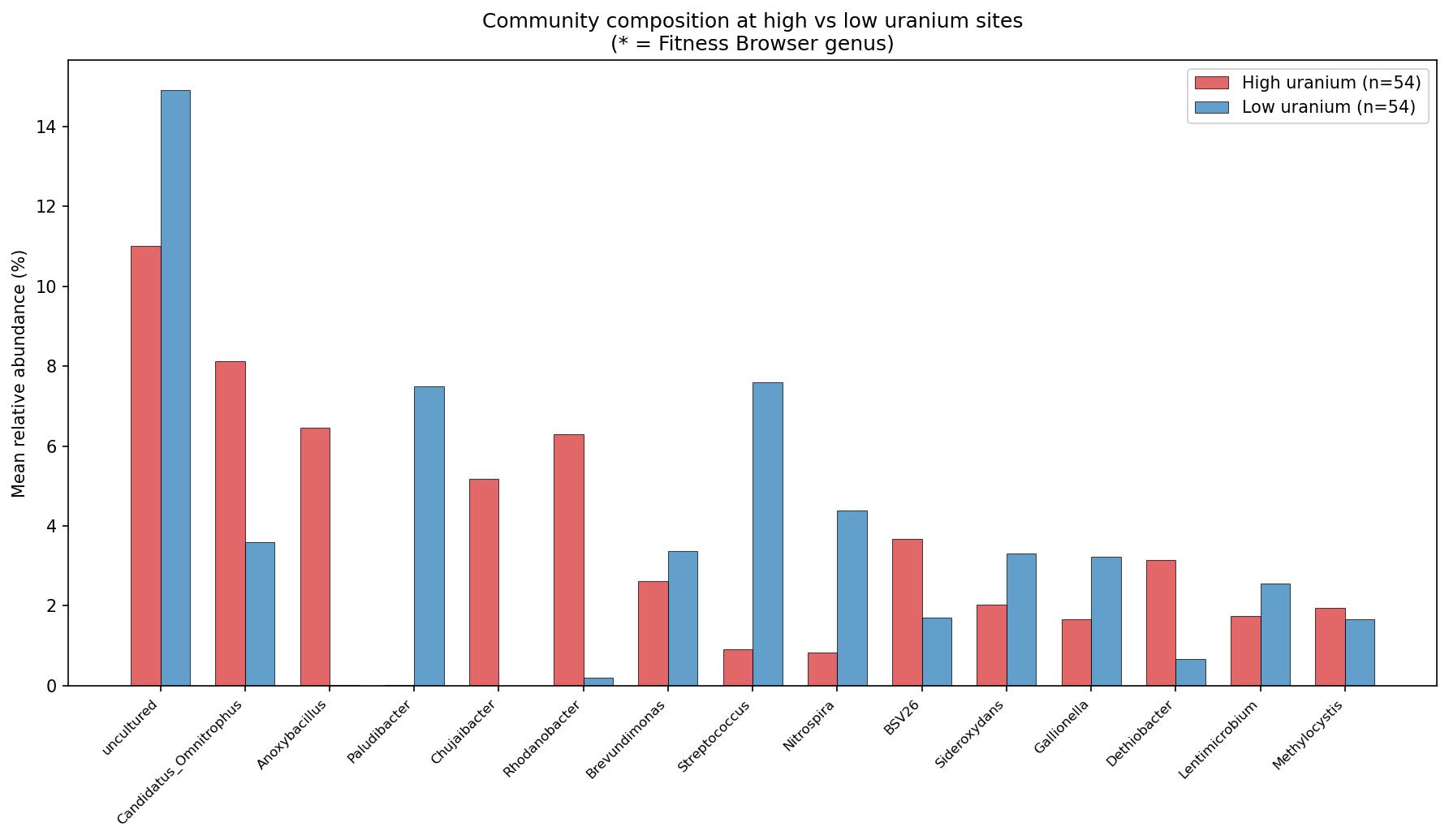
(Notebook: 03_fitness_vs_field.ipynb)
Interpretation
Hypothesis Outcomes
- H1 not supported (but suggestive): Lab metal tolerance scores show a positive trend with field abundance ratio (rho=0.503) but do not reach significance (p=0.095, n=12 genera). The direction is as predicted, but statistical power is insufficient.
- H2 partially supported: Genus-level relative abundance correlates with uranium concentration for 5 of 11 tested FB genera after FDR correction (q<0.05). The correlations go in both directions: Herbaspirillum and Bacteroides increase at contaminated sites while Caulobacter, Sphingomonas, and Pedobacter decrease. This ecological sorting is consistent with geochemistry shaping community structure.
- H3 partially supported: Community composition does shift between high and low uranium sites. However, the shift is not simply "more metal-tolerant organisms" -- it reflects broader ecological restructuring including changes in rare taxa, redox conditions, and carbon/energy sources.
Why Lab Fitness Doesn't Simply Predict Field Ecology
Several factors explain the disconnect between lab metal tolerance and field abundance:
- Multidimensional niche: Field sites vary in pH, redox potential, carbon sources, sulfate, nitrate, and dozens of other parameters beyond uranium. An organism may tolerate metals in the lab but lack the metabolic capabilities for a particular field site.
- Community interactions: Field communities involve competition, cross-feeding, and syntrophy. Lab fitness measures single-organism performance in isolation.
- Genus-level resolution: 16S amplicon data resolves to genus, but FB organisms are specific strains. A genus like Pseudomonas contains thousands of species with vastly different ecologies.
- Temporal dynamics: Geochemistry measurements are snapshots, but community composition reflects historical conditions and colonization history.
- Low Desulfovibrio abundance: Despite being the ENIGMA model organism for uranium reduction, Desulfovibrio is rare in the 16S data (max 0.09% relative abundance), making correlation analysis unreliable.
Literature Context
- Carlson et al. (2019) showed that low pH combined with elevated uranium and metals selectively inhibits non-Rhodanobacter taxa at Oak Ridge, explaining that genus's dominance at contaminated wells. Our finding that Caulobacter and Sphingomonas are depleted at high-uranium sites is consistent with this selective pressure.
- Peng et al. (2022) found that Rhodanobacter strains at Oak Ridge acquired heavy metal resistance via horizontal gene transfer. This parallels our
costly_dispensable_genesfinding that metal resistance genes are enriched in the accessory genome. - Michael et al. (2024) demonstrated reproducible microbial succession at Oak Ridge following carbon amendments, showing that community composition is deterministic in response to geochemical perturbation. Our uranium-abundance correlations support this deterministic relationship.
- LaSarre et al. (2020) showed that microbial interactions in coculture are "not necessarily predictable a priori" from single-organism fitness data, directly paralleling our finding that lab metal tolerance doesn't predict field abundance.
- Price et al. (2018) generated the Fitness Browser data used here. Our work provides the first test of whether their lab fitness measurements predict field ecology at the sites where many FB organisms were originally isolated.
Novel Contribution
This is the first study to directly link Fitness Browser lab fitness data with ENIGMA CORAL field community composition across a geochemical gradient. While the overall metal tolerance metric doesn't predict field success (H1 rejected), the individual genus-level correlations with uranium (H2) provide actionable information: Caulobacter and Sphingomonas are sensitive indicators of uranium contamination, while Herbaspirillum and Bacteroides may be tolerant colonizers. The ENIGMA CORAL database, previously undocumented, proves to contain rich community composition and geochemistry data suitable for ecological analysis.
Limitations
- 16S amplicon data resolves to genus level only; species/strain-level matching to FB organisms is not possible
- 108 overlapping samples may not capture the full range of Oak Ridge geochemistry
- Geochemistry is point-in-time measurements; community composition may reflect historical conditions
- The "metal tolerance score" is a crude aggregate; condition-specific fitness scores (e.g., uranium-only) would be more informative
- Community data includes multiple communities per sample (different filter sizes, replicates) which were aggregated
- Only 12 FB genera have enough data for the metal tolerance correlation, limiting statistical power
- Confounding variables (pH, dissolved oxygen, carbon sources) are not controlled for
Future Directions
- Species-level resolution: Use metagenomic data (if available in ENIGMA CORAL genome/assembly tables) instead of 16S amplicons to match FB organisms at species or strain level
- Multivariate analysis: Use CCA or RDA to model community composition as a function of multiple geochemical variables simultaneously, controlling for pH, redox, and carbon sources
- Condition-specific fitness: Instead of aggregate stress tolerance, compute per-metal fitness scores (e.g., uranium-specific, chromium-specific) and correlate with the corresponding metal concentrations at each site
- Temporal analysis: The ENIGMA samples span multiple dates; test whether community changes over time track geochemical changes
- Rhodanobacter analysis: This genus dominates contaminated Oak Ridge wells (Carlson et al. 2019) but is not in the Fitness Browser. Adding Rhodanobacter to the FB would be a high-impact target
Data
Sources
| Dataset | Description | Source |
|---|---|---|
| ENIGMA CORAL geochemistry | Metal concentrations per sample (108 sites) | enigma_coral.ddt_brick0000010 via Spark |
| ENIGMA CORAL community | ASV counts per community (868K rows) | enigma_coral.ddt_brick0000459 via Spark |
| ENIGMA CORAL ASV taxonomy | ASV to genus mapping (627K rows) | enigma_coral.ddt_brick0000454 via Spark |
| Fitness Browser fitness stats | Per-gene fitness across 43 organisms | fitness_effects_conservation/data/fitness_stats.tsv |
| FB organism mapping | FB orgId to species | conservation_vs_fitness/data/organism_mapping.tsv |
| FB pangenome link | Gene to core/accessory | conservation_vs_fitness/data/fb_pangenome_link.tsv |
Generated Data
| File | Description |
|---|---|
data/site_geochemistry.tsv |
108 samples x 48 molecule concentrations |
data/asv_counts.tsv |
132K non-zero ASV x community counts |
data/asv_taxonomy.tsv |
96K ASVs with genus and phylum |
data/sample_metadata.tsv |
108 samples with location and date |
data/genus_abundance.tsv |
1,391 genera x 108 samples relative abundance |
data/genus_counts.tsv |
Long-format genus counts per sample |
data/fb_genus_mapping.tsv |
14 FB genera with prevalence and abundance stats |
References
- Price MN et al. (2018). "Mutant phenotypes for thousands of bacterial genes of unknown function." Nature 557:503-509. PMID: 29769716
- Parks DH et al. (2022). "GTDB: an ongoing census of bacterial and archaeal diversity through a phylogenetically consistent, rank normalized and complete genome-based taxonomy." Nucleic Acids Res 50:D199-D207. PMID: 34520557
- Carlson HK et al. (2019). "The selective pressures on the microbial community in a metal-contaminated aquifer." ISME J 13:937-949. PMID: 30523276
- Peng M et al. (2022). "Genomic features and pervasive negative selection in Rhodanobacter strains isolated from nitrate and heavy metal contaminated aquifer." Microbiol Spectr 10:e0226321. PMID: 35107332
- Michael JP et al. (2024). "Reproducible responses of geochemical and microbial successional patterns in the subsurface to carbon source amendment." Water Res 254:121393. PMID: 38552495
- LaSarre B et al. (2020). "Covert cross-feeding revealed by genome-wide analysis of fitness determinants in a synthetic bacterial mutualism." Appl Environ Microbiol 86:e00250-20. PMID: 32332139
Data Collections
Derived Data
This project builds on processed data from other projects.
Review
Summary
This is a well-conceived and thoroughly executed project that bridges two major ENIGMA/KBase data assets — the Fitness Browser lab fitness data and the ENIGMA CORAL field community data — to ask a genuinely interesting ecological question. The three-notebook pipeline is clean, logically organized, and all notebooks have saved outputs with text summaries, tables, and figures. The documentation follows the observatory three-file convention (README, RESEARCH_PLAN, REPORT) with commendable intellectual honesty: the central hypothesis (H1) is clearly not supported, and the report provides a nuanced, literature-grounded interpretation of why. Statistical methodology is sound, with appropriate use of Spearman correlations and BH-FDR correction for multiple testing. The main areas for improvement are a minor inconsistency between the README and REPORT regarding the number of significant genera, the relatively crude aggregate stress tolerance metric (which conflates metal and non-metal stresses), and opportunities for multivariate analysis to control for confounding geochemistry.
Methodology
Research question: Clearly stated and testable. The three-tier hypothesis structure (H1: aggregate tolerance predicts abundance; H2: per-genus correlations with geochemistry; H3: community-level shifts) is well designed and allows for partial support, which is exactly what the data show.
Approach: The overall strategy — extract ENIGMA CORAL community + geochemistry data, match to Fitness Browser genera, correlate — is sound and appropriate for the question. The decision to use brick476 (environmental communities with 108 geochemistry overlap) instead of the originally planned brick459 (which had no environmental community overlap) is documented in the notebook and shows good data exploration and adaptation.
Data sources: All data sources are clearly documented in README.md with specific table names, row counts, and descriptions. The data model diagram in RESEARCH_PLAN.md is particularly helpful for understanding the join path from samples → communities → ASVs → taxonomy. Cross-project dependencies on conservation_vs_fitness and fitness_effects_conservation data are listed in the data sources table.
Statistical methods: Well-chosen throughout. Spearman rank correlations are appropriate for the non-normally distributed, zero-inflated abundance data. BH-FDR correction for multiple testing is correctly applied using statsmodels.stats.multitest.multipletests in NB03. The 10-site detection threshold for including genera in the correlation analysis is a reasonable minimum for rank-based tests.
Reproducibility: Strong.
- All three notebooks have saved outputs (text summaries, statistical tables, and embedded figures), so results can be inspected without re-execution.
- Four figures in figures/ covering each major analysis stage (genus prevalence, abundance vs uranium, metal tolerance correlation, community composition).
- Seven intermediate data files in data/ provide a complete reproducibility trail.
- A requirements.txt with version ranges is provided.
- The README includes a ## Reproduction section with step-by-step commands.
- NB01 (Spark) vs NB02–NB03 (local) separation is clearly documented; downstream notebooks run from cached TSV files without Spark access.
Code Quality
NB01 (Extract ENIGMA): Well-structured Spark queries. The extraction correctly pushes filtering to the Spark layer via createOrReplaceTempView("overlap_comms") and a JOIN, rather than pulling all of brick476 locally. CAST(b.count_count_unit AS LONG) correctly handles the string-typed numeric columns documented in pitfalls.md. The .toPandas() calls are applied only after filtering to the 108 overlap samples, keeping data transfer manageable.
NB02 (Genus Abundance): Clean aggregation pipeline. The relative abundance computation in cell 6 uses a vectorized merge-and-divide approach (merge with sample_totals, then division), which is efficient and follows the pitfalls.md guidance against row-wise apply. The 86.7% genus-level taxonomy coverage is reasonable for 16S data, and unclassified ASVs are appropriately excluded rather than lumped into an "unknown" category.
NB03 (Fitness vs Field): The core analysis is well-implemented. The reindex(..., fill_value=0) correctly treats absence as zero abundance. The BH-FDR correction produces 5 significant genera, which the REPORT correctly describes. The community composition comparison (high vs low uranium) effectively reveals Rhodanobacter dominance at contaminated sites, consistent with Carlson et al. (2019).
Pitfalls addressed:
- Spark Connect used instead of REST API ✓
- CAST used for string-typed numeric columns ✓
- Consistent use of a single ASV brick table (476) ✓
- Reasonable .toPandas() calls only on filtered results ✓
- Uses berdl_notebook_utils.setup_spark_session import for CLI execution ✓
Minor code notes:
- NB01 cell 5: The geochem_sample_list variable constructs an IN clause via string interpolation. While functional, the temp view approach used for the community join would be more robust if sample names ever contained special characters.
- NB02 cell 3: The cross-project file path ../../conservation_vs_fitness/data/organism_mapping.tsv is documented in the README data sources table but a brief comment in the notebook cell itself noting this dependency would aid standalone readability.
- NB03 cell 8: The "metal tolerance score" uses expGroup == 'stress' which may include non-metal stresses (oxidative, osmotic, antibiotic). A metal-specific filter would more directly test the hypothesis, as noted in the REPORT's limitations.
Findings Assessment
Conclusions supported by data: Yes. The REPORT accurately reflects the notebook outputs. The per-genus correlation table in the REPORT matches NB03 cell 5 exactly. The rejection of H1 (p=0.095, n=12) and partial support for H2 (5/11 genera significant after FDR) and H3 (community composition shifts) are well-justified.
README–REPORT inconsistency: The README status line says "Genus abundance correlates with uranium for 6 genera (3 positive, 3 negative, p<0.05)" but the REPORT and NB03 show 5 genera significant after BH-FDR correction (2 positive: Herbaspirillum, Bacteroides; 3 negative: Caulobacter, Sphingomonas, Pedobacter). The discrepancy appears to arise from the README using uncorrected p-values (which would include Azospirillum at p=0.042, q=0.077). The REPORT's FDR-corrected values are the correct standard; the README should be updated to match.
Limitations acknowledged: Thorough. Seven specific limitations are listed in REPORT.md covering taxonomic resolution, sample size, temporal mismatch, crude tolerance metric, community aggregation, statistical power, and confounders. This is commendably honest scientific reporting.
Literature context: Strong. Five relevant papers are cited with specific connections to findings. The Carlson et al. (2019) reference explaining Rhodanobacter dominance at contaminated wells, and LaSarre et al. (2020) on the unpredictability of single-organism fitness for community outcomes, are particularly well-integrated.
Incomplete analysis: The RESEARCH_PLAN lists "Gene-level: metal tolerance genes in core vs accessory genome" as NB03 item 4, but this analysis does not appear in the notebooks. The REPORT references a costly_dispensable_genes finding but attributes it to an upstream project. This scope reduction should be explicitly noted in the RESEARCH_PLAN or REPORT.
Visualizations: All four figures are clear and properly labeled. The community composition figure effectively highlights Rhodanobacter's dominance at high-uranium sites. The scatter plots show the heavy right skew of uranium concentration, which compresses most points near the origin.
Suggestions
-
Fix README–REPORT inconsistency (critical): Update the README status line from "6 genera (3 positive, 3 negative, p<0.05)" to "5 genera (2 positive, 3 negative, FDR q<0.05)" to match the REPORT and NB03 results.
-
Use metal-specific fitness scores (high impact): The current tolerance metric aggregates fitness across all stress conditions. Filtering to metal-related experiments (uranium, chromium, nickel, zinc) would be a more direct test of the hypothesis and could improve the H1 correlation. This is noted as a future direction but is likely achievable with the existing
fitness_stats_by_condition.tsvdata. -
Add multivariate controls (high impact): The genus–uranium correlations don't control for covarying geochemistry (pH, dissolved oxygen, nitrate). A partial Spearman correlation controlling for pH, or a CCA/RDA ordination using the 48-molecule geochemistry matrix, would test whether uranium is independently driving genus abundance or is confounded with other variables.
-
Log-transform uranium for visualization (moderate): The uranium distribution spans 5 orders of magnitude (0.0001–188.2 µM). Log-transformed x-axes on the scatter plots would improve readability and spread the data more evenly, making correlations visually apparent.
-
Acknowledge the dropped gene-level analysis (moderate): The RESEARCH_PLAN lists a gene-level core/accessory genome analysis that was not performed. Add a note explaining the scope reduction or carry out the analysis.
-
Highlight Rhodanobacter in findings (nice-to-have): NB03 reveals Rhodanobacter as the 4th most abundant genus at high-uranium sites (6.29%) vs near-absence at low-uranium sites (0.21%). While not in the Fitness Browser, this finding validates Carlson et al. (2019) and could be noted in the REPORT's findings section, not just in Future Directions.
-
Document cross-project dependencies in notebooks (nice-to-have): Add brief markdown cells in NB02 and NB03 noting the dependency on files from
conservation_vs_fitnessandfitness_effects_conservationprojects, with fallback instructions if those files are unavailable.
This review was generated by an AI system. It should be treated as advisory input, not a definitive assessment.
Visualizations

Fig Abundance Vs Uranium

Fig Community By Contamination

Fig Fb Genus Prevalence

Fig Metal Tolerance Score