About Microbial Discovery Forge
AI co-scientist and research observatory for BERDL-scale microbial discovery.
Architecture
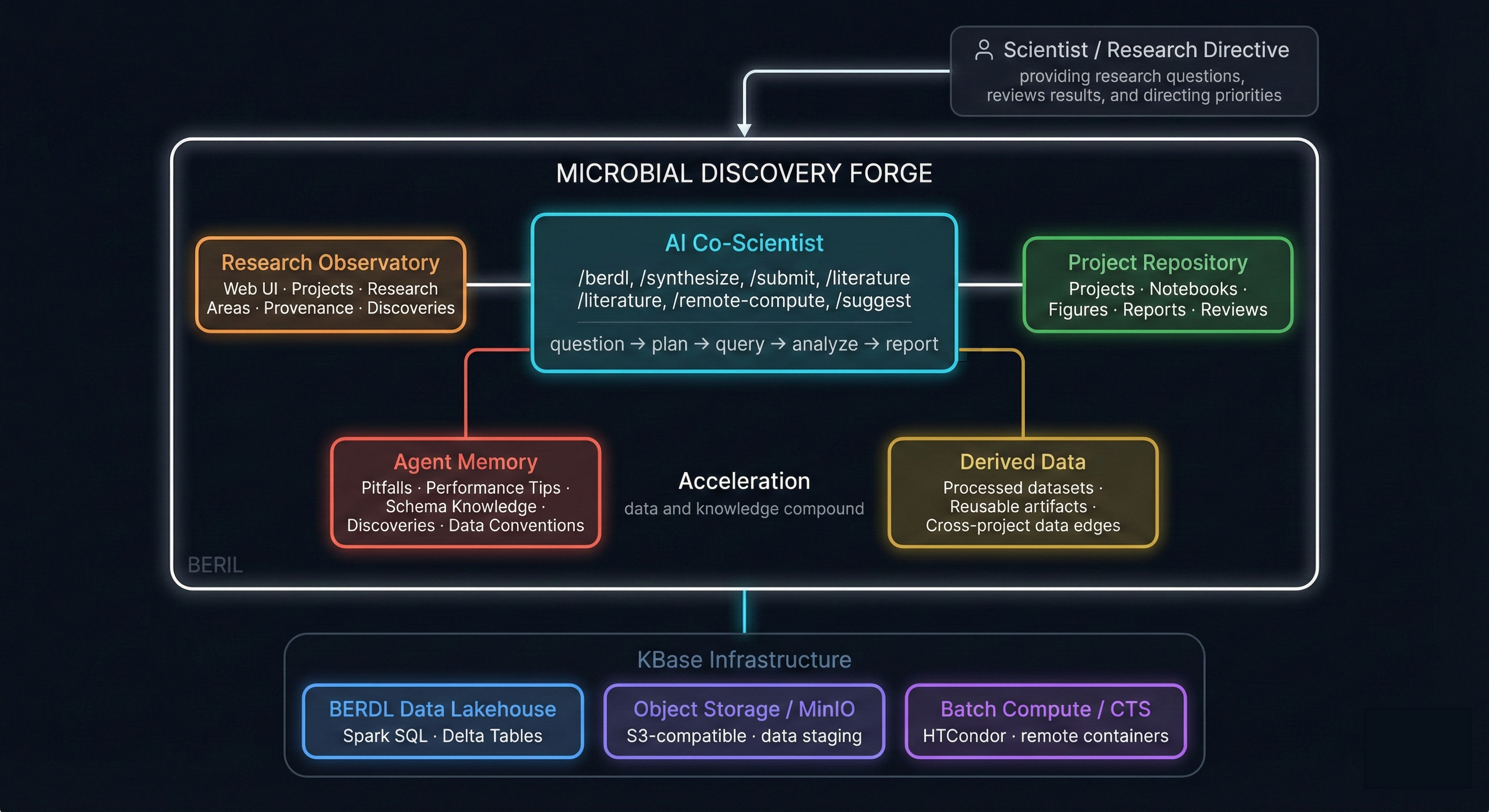
Two Experiences
AI Co-Scientist
A human-in-the-loop AI research workflow that plans analyses, executes them on BERDL, synthesizes findings with literature context, and captures lessons learned as reusable skills and memory.
Research Observatory
Explore curated data collections, browse completed research projects with full reports, and discover findings documented with figures, citations, and reproducible notebooks.
Microbial Discovery Forge Team

Paramvir S. Dehal
Project lead & primary contact; PI BERIL and AI/ML Team Lead KBase; Scientific lead, primary developer of Microbial Discovery Forge

Ontologies and metadata/data harmonization guidance

Engineering support

Skills development and evaluations contributions

Evaluations contributions

Project management support

Data collections
Partner / Enabling Effort

Vision/leadership for BERDL (KBase)

Driving BERDL data resources and curation

BERDL architect

BERIL co-PI

BERIL co-PI
Contributing Projects and Resources
Infrastructure
BERIL
The BER Intelligent Layer is the broader program providing AI integration capabilities for DOE BER data resources. The Research Observatory is developed under the BERIL project umbrella.
KBase
The Department of Energy Systems Biology Knowledgebase provides the platform ecosystem, community infrastructure, and the AI/ML team that supports this work.
BERDL
The BER Data Lakehouse is the underlying data resource hosting 35+ databases across 9 tenants of curated scientific datasets. Developed by the KBase team, BERDL provides the data foundation for all Observatory analyses.
Data Partners
NMDC
The National Microbiome Data Collaborative provides multi-omics microbiome data including annotations, metabolomics, and proteomics. Used in Observatory projects for environmental and functional analyses.
ENIGMA
The Ecosystems and Networks Integrated with Genes and Molecular Assemblies SFA provides environmental microbiology data from Oak Ridge field sites, including genomes, communities, and strain isolates.
JGI
The Joint Genome Institute provides upstream genome sequencing, assembly, and annotation pipelines. JGI's GOLD and IMG databases supply the foundational genomic data underlying BERDL collections.
PhageFoundry
The Phage Foundry provides species-specific genome browsers for phage-host interaction research, with curated data for Acinetobacter, Klebsiella, Pseudomonas, and other priority pathogens.
What is BERDL?
The KBase BER Data Lakehouse (BERDL) is a Delta Lakehouse hosting 35 databases across 9 tenants of curated scientific datasets for computational biology research. It provides:
- Multiple data collections including pangenomes, mutant fitness data, biochemistry, multi-omics, marine ecology, phage research, and more
- Spark SQL access for large-scale queries
- JupyterHub integration for interactive analysis
- REST API for programmatic access
Primary BERDL Collections
Pangenome Collection
kbase_ke_pangenome
Pangenome data for 293,059 genomes across 27,690 microbial species derived from GTDB r214. Includes core/accessory gene classification, functional annotations, and ANI relationships.
Fitness Browser
kescience_fitnessbrowser
Gene fitness data from transposon mutant experiments across 40+ bacterial organisms. Identify essential genes and condition-specific fitness effects.
KBase Genomes
kbase_genomes
Structural genomics data including contigs, features, and protein sequences from the KBase genome repository.
ModelSEED Biochemistry
kbase_msd_biochemistry
Biochemistry reference data for metabolic modeling. Reactions, compounds, and pathway mappings from ModelSEED.
Phenotype Collection
kbase_phenotype
Experimental phenotype data including growth conditions, measurements, and phenotypic outcomes.
AI Integration
BERIL provides skills and plugins for AI assistants that enable:
- Schema exploration - Understand available tables and columns
- Query generation - Generate SQL queries for common analysis patterns
- Data interpretation - Help interpret results in biological context
- Cross-collection analysis - Combine data from multiple collections
AI assistants with BERIL skills can help researchers explore the data lakehouse more efficiently, reducing the overhead of learning new schemas and query patterns.
How to Cite
If you use the Microbial Discovery Forge or its findings in your work, please cite:
Paramvir S. Dehal. Microbial Discovery Forge — v0.1, 2026. Built on the KBase BER Data Lakehouse (BERDL).
Getting Started
For Researchers
- Browse existing projects for inspiration
- Explore collections to understand available data
- Read query pitfalls before writing SQL
- Access JupyterHub at hub.berdl.kbase.us
For Contributors
For contributions, please contact psdehal@lbl.gov.
- Pick a research idea or propose your own
- Create a project folder in
projects/ - Document findings in
docs/discoveries.md - Share pitfalls and learnings as you go