Pan-Bacterial Metal Fitness Atlas
CompletedResearch Question
Across diverse bacteria subjected to genome-wide fitness profiling under metal stress, what is the genetic architecture of metal tolerance — is it encoded in the core or accessory genome, is it conserved across species, and can fitness-validated metal tolerance genes predict capabilities across the broader pangenome?
Research Plan
Hypothesis
- H0: Metal tolerance genes identified by RB-TnSeq are randomly distributed across core and accessory genomes, and cross-species conservation of metal fitness determinants is no greater than expected by chance.
- H1: Metal tolerance has a two-tier genetic architecture — general metal stress response is encoded in the core genome while specific metal resistance mechanisms (efflux, sequestration) are accessory — and conserved cross-species metal fitness gene families exist that can predict metal tolerance from genome content alone.
Sub-hypotheses
- H1a: Genes important for tolerance to toxic metals (Co, Ni, Cu, Zn, Al) are enriched in the accessory genome relative to baseline, consistent with HGT-mediated acquisition. (Prior: DvH heavy-metal genes are 71.2% core vs 76.3% baseline.)
- H1b: Genes important for metal homeostasis under essential metal limitation (Fe, Mo, W) are enriched in the core genome, reflecting fundamental metabolic dependence.
- H1c: Cross-species conserved metal fitness gene families (ortholog groups with consistent metal phenotypes in ≥3 organisms) exist and are enriched for known metal resistance functions (efflux, metal-binding, redox).
- H1d: Hypothetical proteins with conserved metal fitness phenotypes across species represent novel metal biology genes that can be functionally annotated by this approach.
Revision History
- v1 (2026-02-17): Initial plan
Overview
The Fitness Browser contains 559 metal-related experiments across 31 organisms covering 14 metals (cobalt, nickel, copper, zinc, aluminum, iron, tungsten, molybdenum, chromium, uranium, selenium, manganese, mercury, cadmium). No published study has performed a systematic cross-species, genome-wide fitness comparison for metal tolerance. This project builds a "Metal Fitness Atlas" by: (1) extracting all metal-condition fitness data (383,349 gene x metal records), (2) mapping metal-important genes to pangenome conservation (core vs accessory) for 22 organisms with FB-pangenome links, (3) identifying 1,182 conserved metal fitness gene families across species via ortholog groups, and (4) validating a metal tolerance repertoire scoring approach. The project is framed around DOE critical minerals priorities — cobalt, nickel, manganese, chromium, aluminum, and tungsten are on the USGS critical minerals list.
Key Findings
1. Metal-Important Genes Are Enriched in the Core Genome
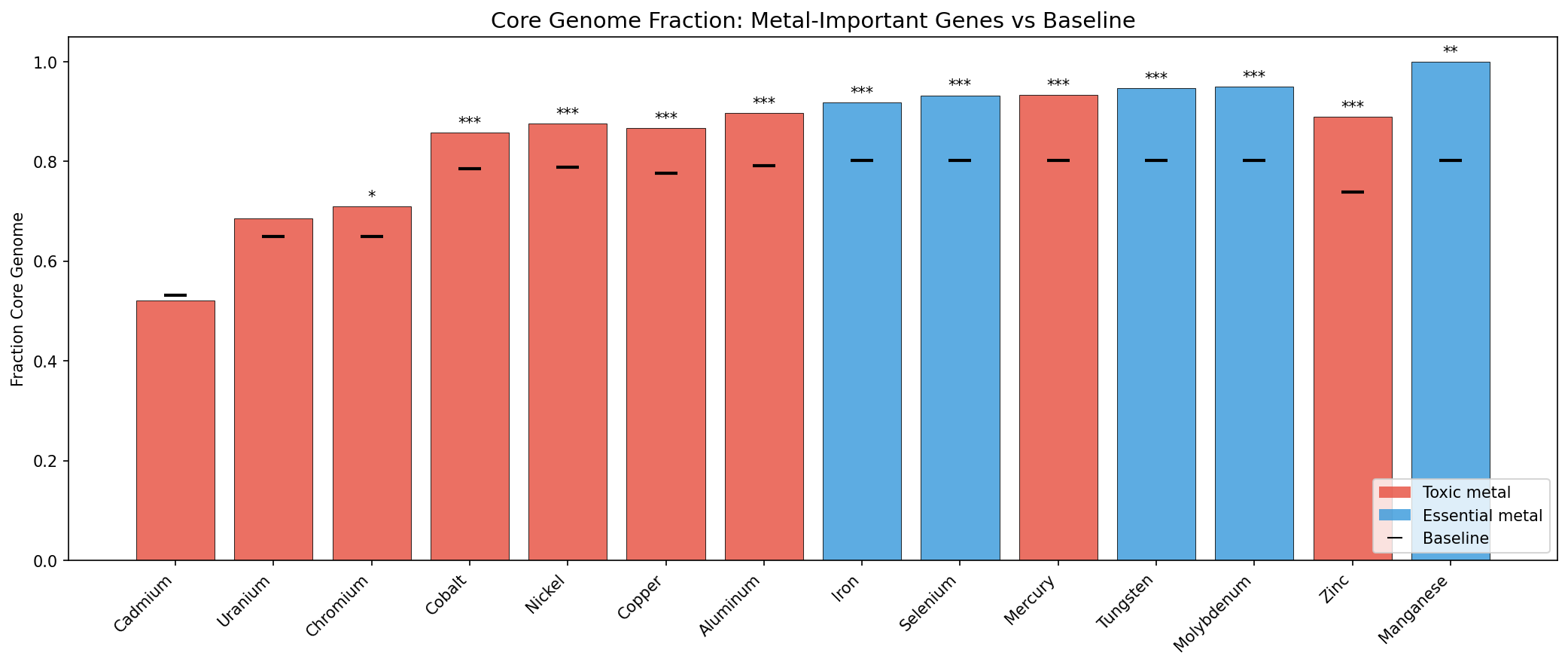
Across 22 organisms and 14 metals, genes with significant fitness defects under metal stress are 87.4% core vs 76.9% baseline (OR=2.08, p=4.3e-162). This is the opposite of the initial hypothesis (H1a), which predicted accessory enrichment for toxic metal genes based on the prior DvH finding (71.2% core for condition-specific heavy-metal genes). The discrepancy arises because genome-wide metal fitness defects predominantly reflect core cellular processes vulnerable to metal disruption (cell envelope, DNA repair, central metabolism), not specialized metal resistance mechanisms.
21 of 22 organisms show metal-important genes as more core than baseline; 14 are significant at p<0.05. Only P. fluorescens FW300-N2E3 shows a negligible negative delta (-0.003, p=0.70).
(Notebook: 03_metal_conservation_analysis.ipynb)
2. Essential Metals Show Stronger Core Enrichment Than Toxic Metals
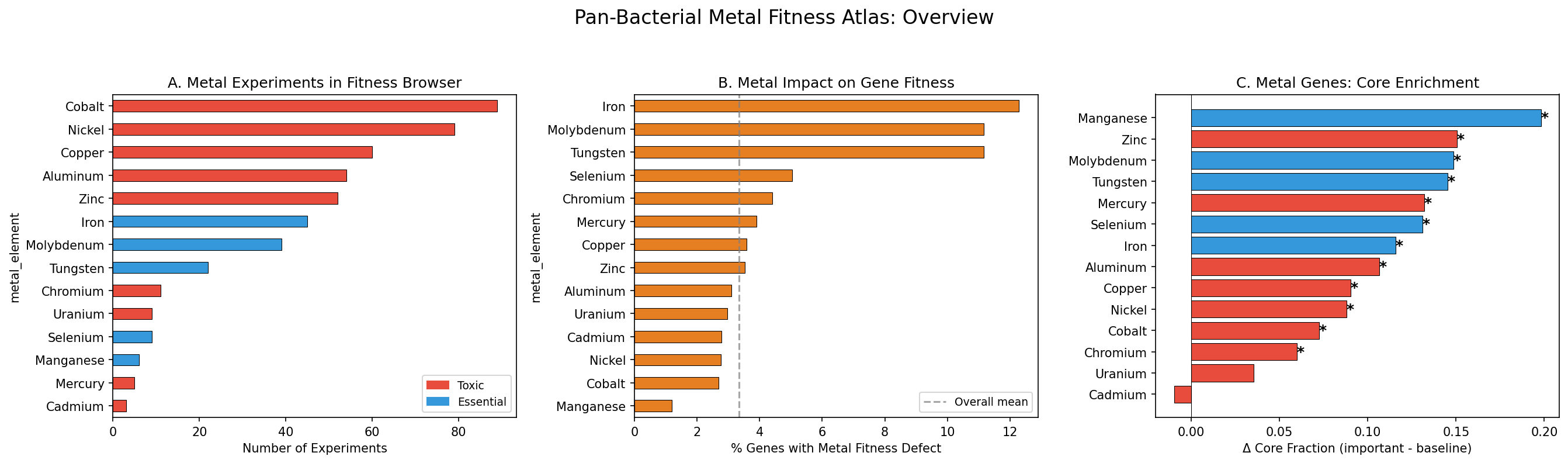
Essential metal tolerance genes (Fe, Mo, W, Se, Mn) show a mean core fraction delta of +0.148, nearly double the toxic metal delta of +0.081. This difference is statistically significant (Mann-Whitney U=39, p=0.015, one-sided). The strongest enrichments: Manganese (+0.198, all 30 important genes are core), Zinc (+0.151), Molybdenum (+0.148), Tungsten (+0.145), and Iron (+0.116). 12 of 14 metals are individually significant at p<0.05. Only Cadmium (1 organism, delta=-0.010, p=0.92) and Uranium (2 organisms, delta=+0.035, p=0.34) are not significant, likely due to limited organism coverage.
Phylogenetic sensitivity: Excluding 4 duplicate P. fluorescens FW300 strains (keeping only pseudo3_N2E3) reduces the analysis from 22 to 18 organisms. The core enrichment is fully robust: OR=2.065, p=5.9e-141 (vs OR=2.083, p=4.3e-162 with all organisms). The result is not driven by Pseudomonas overrepresentation.
Coverage note: Conservation analysis covers 22 of 31 metal-tested organisms (71%). The 9 excluded organisms (Putida, Keio, SynE, Miya, Kang, BFirm, Cola, Ponti, Dino) lack FB-pangenome links. These are taxonomically diverse, so the exclusion is unlikely to introduce systematic bias.
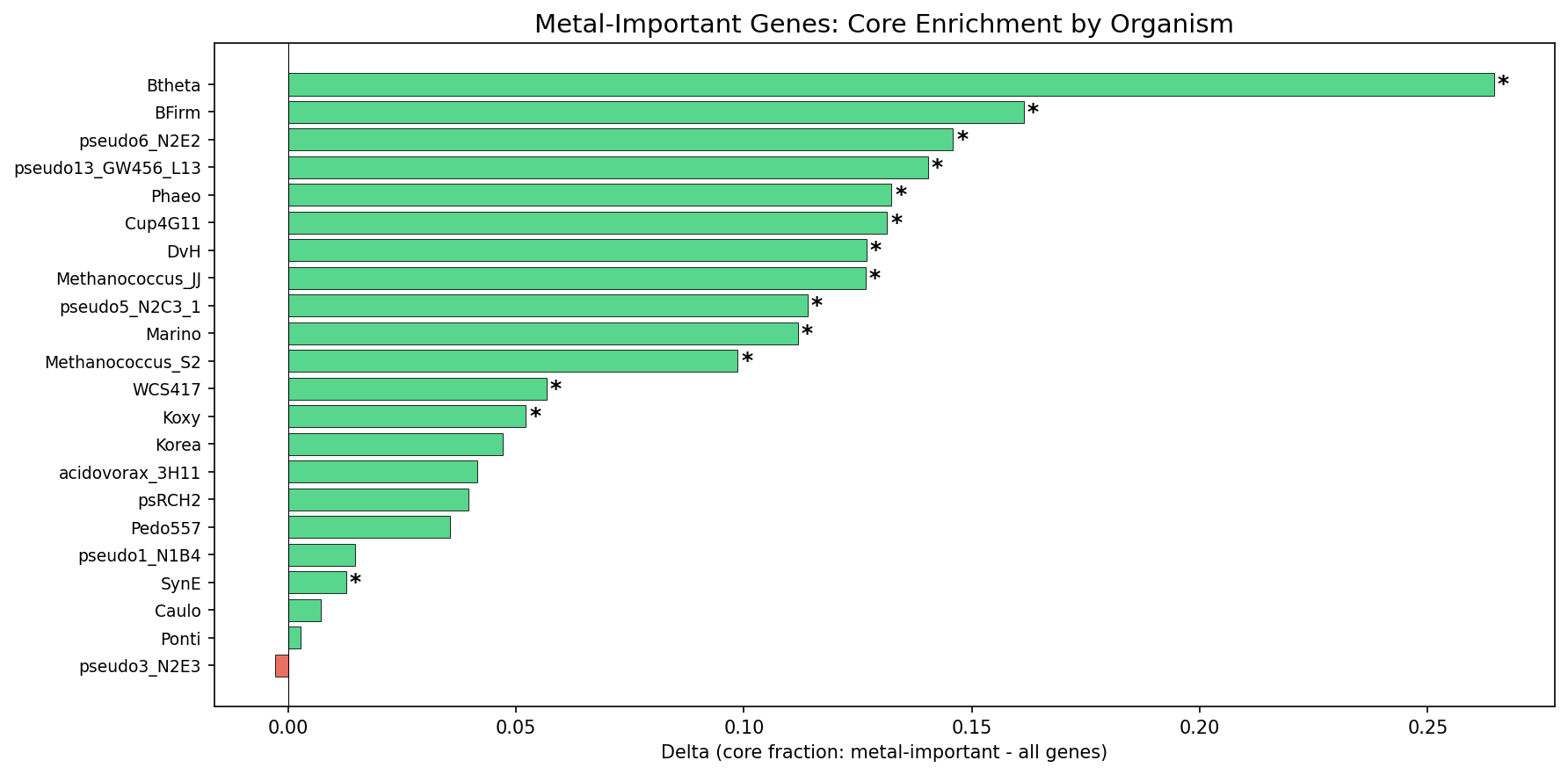
(Notebook: 03_metal_conservation_analysis.ipynb)
3. 559 Metal Experiments Across 31 Organisms and 16 Metals
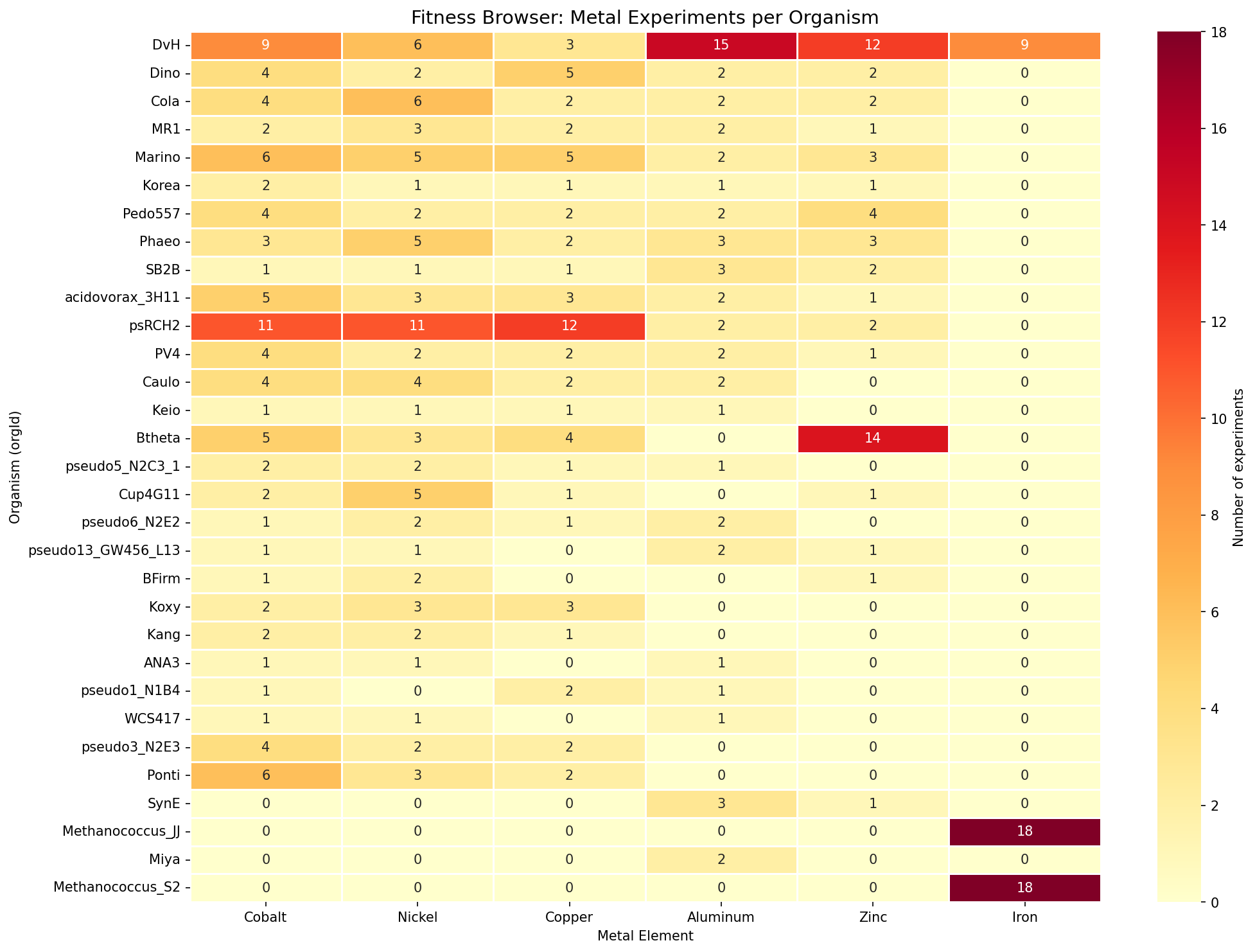
The Fitness Browser contains 559 metal-related experiments (8.2% of 6,804 total) covering 16 metals. Six metals have cross-species coverage (≥3 organisms): Cobalt (27 orgs), Nickel (26), Copper (23), Aluminum (22), Zinc (17), and Iron (3). DvH is the most metal-profiled organism (149 experiments, 13 metals). Three USGS critical minerals have broad FB coverage: Aluminum, Cobalt, and Nickel.
(Notebook: 01_metal_experiment_classification.ipynb)
4. 12,838 Metal-Important Gene Records Across 24 Organisms
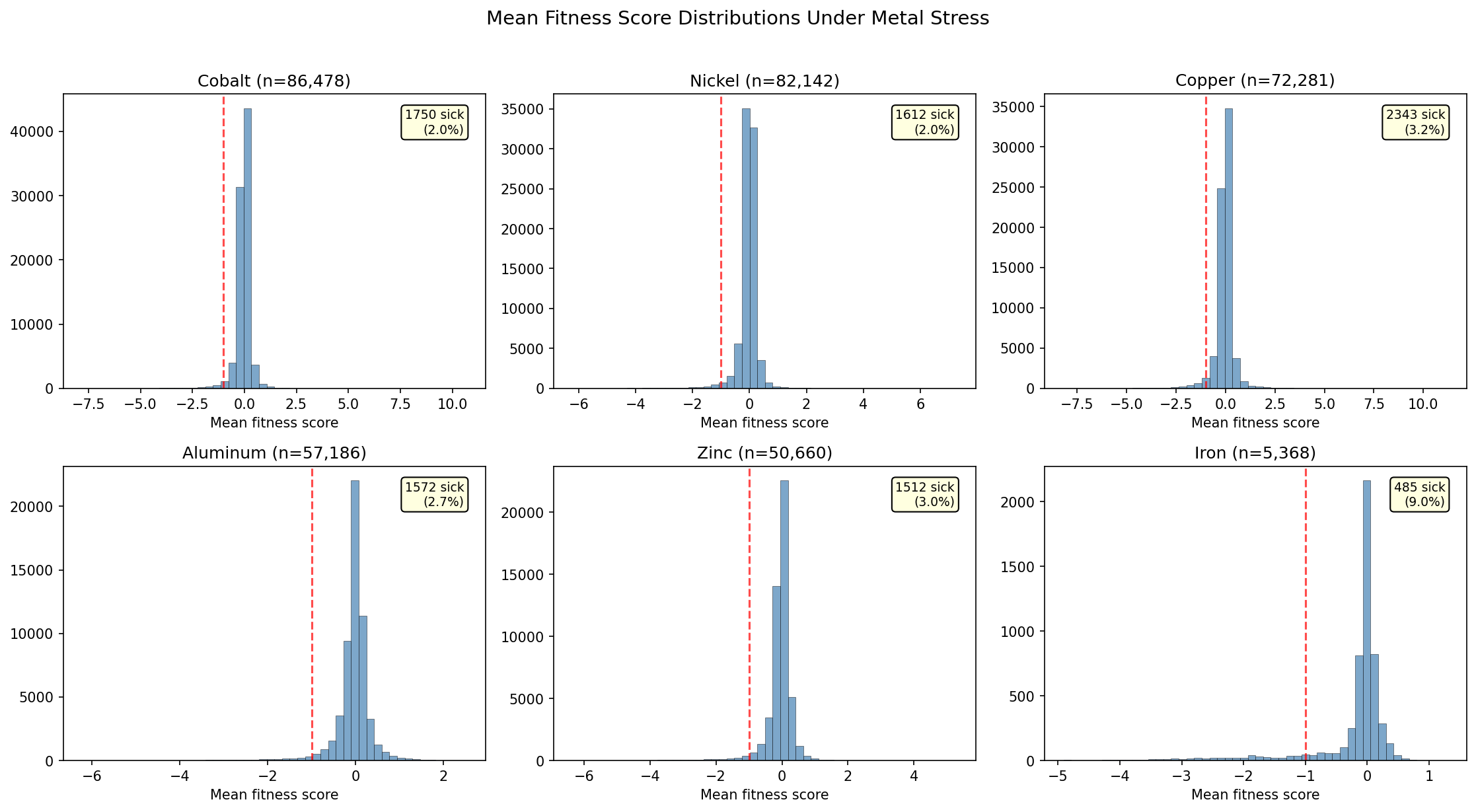
3.3% of all gene × metal records (12,838 / 383,349) show significant fitness defects (fit < -1, |t| > 4). Iron and essential metal limitation produce the highest fraction of important genes (12.3% for Fe, 11.2% for Mo/W), while toxic metals cluster around 2.7-4.4%. DvH has 1,366 metal-important genes (49.8% of its genome across 13 metals). The cyanobacterium Synechococcus elongatus (SynE) is notably metal-sensitive (33.6% of genes important across just 2 metals).
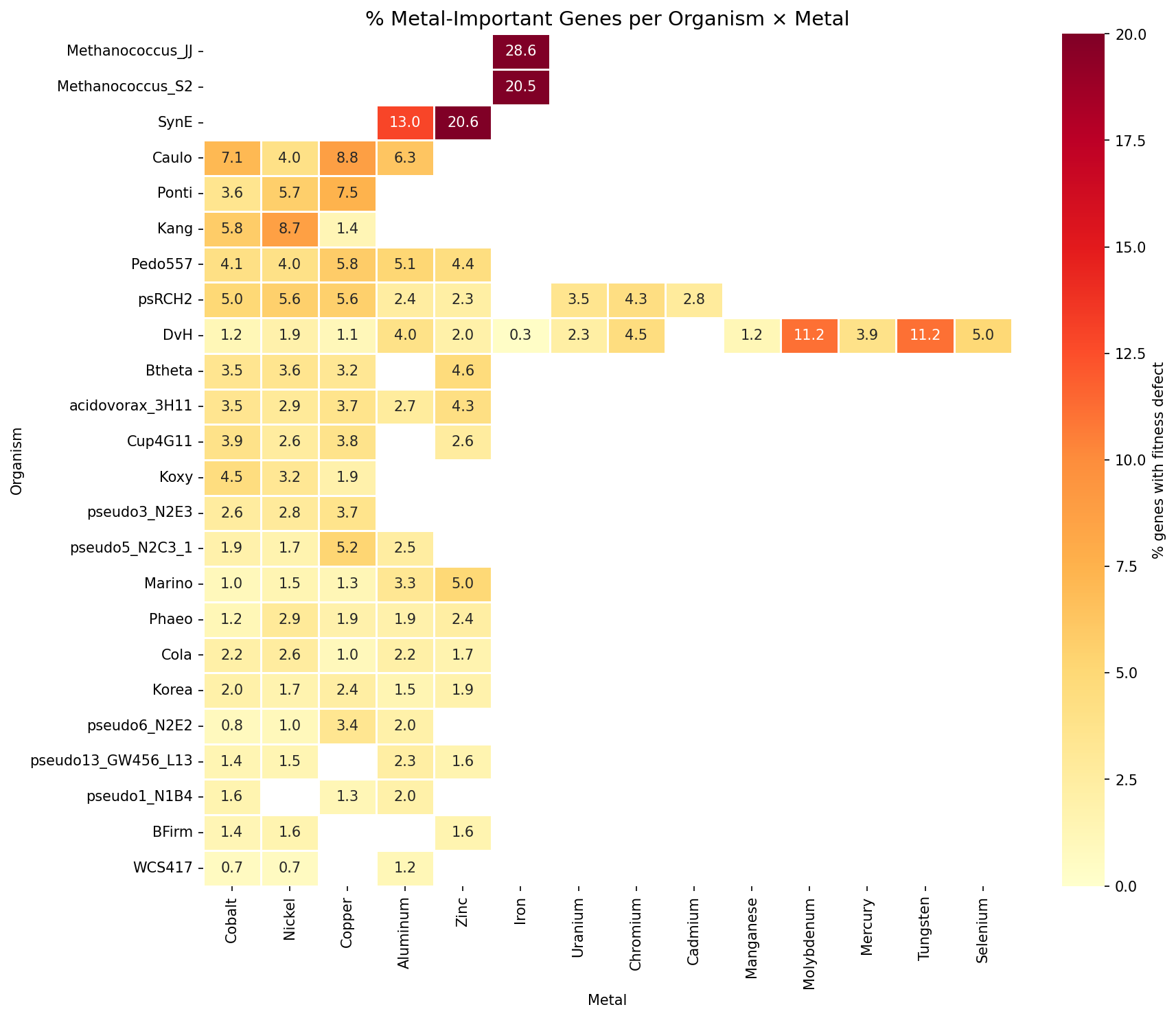
(Notebook: 02_metal_fitness_extraction.ipynb)
5. 1,182 Conserved Metal Gene Families Identified
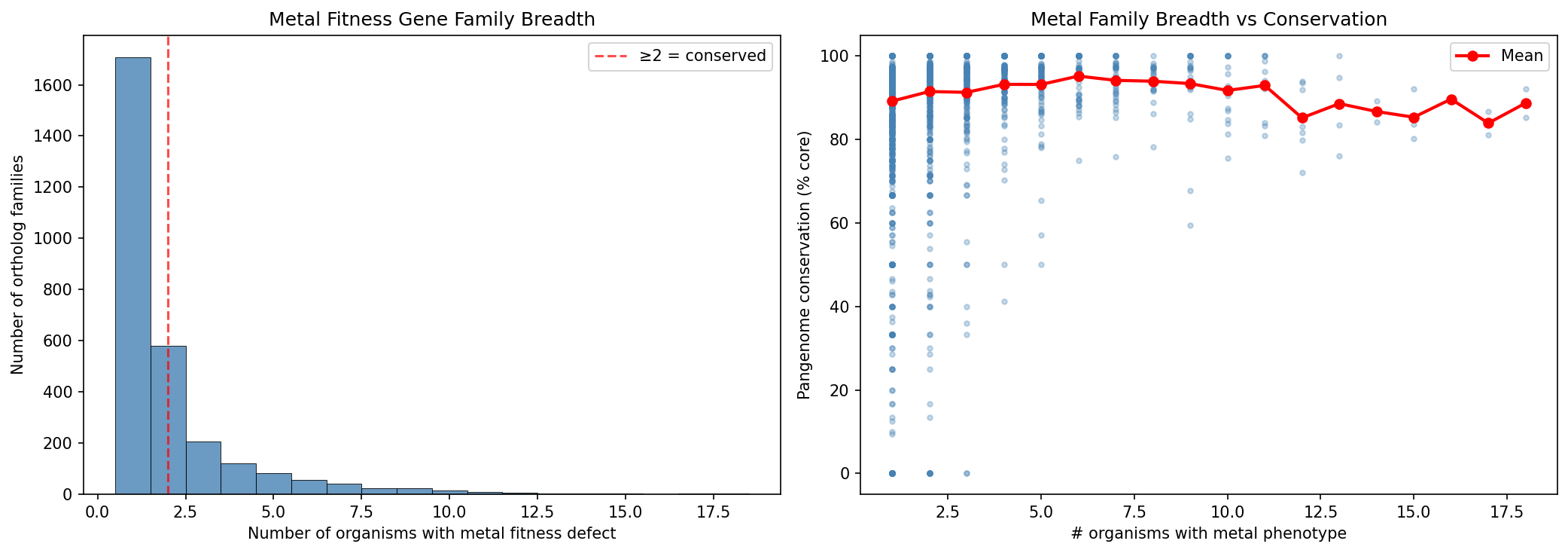
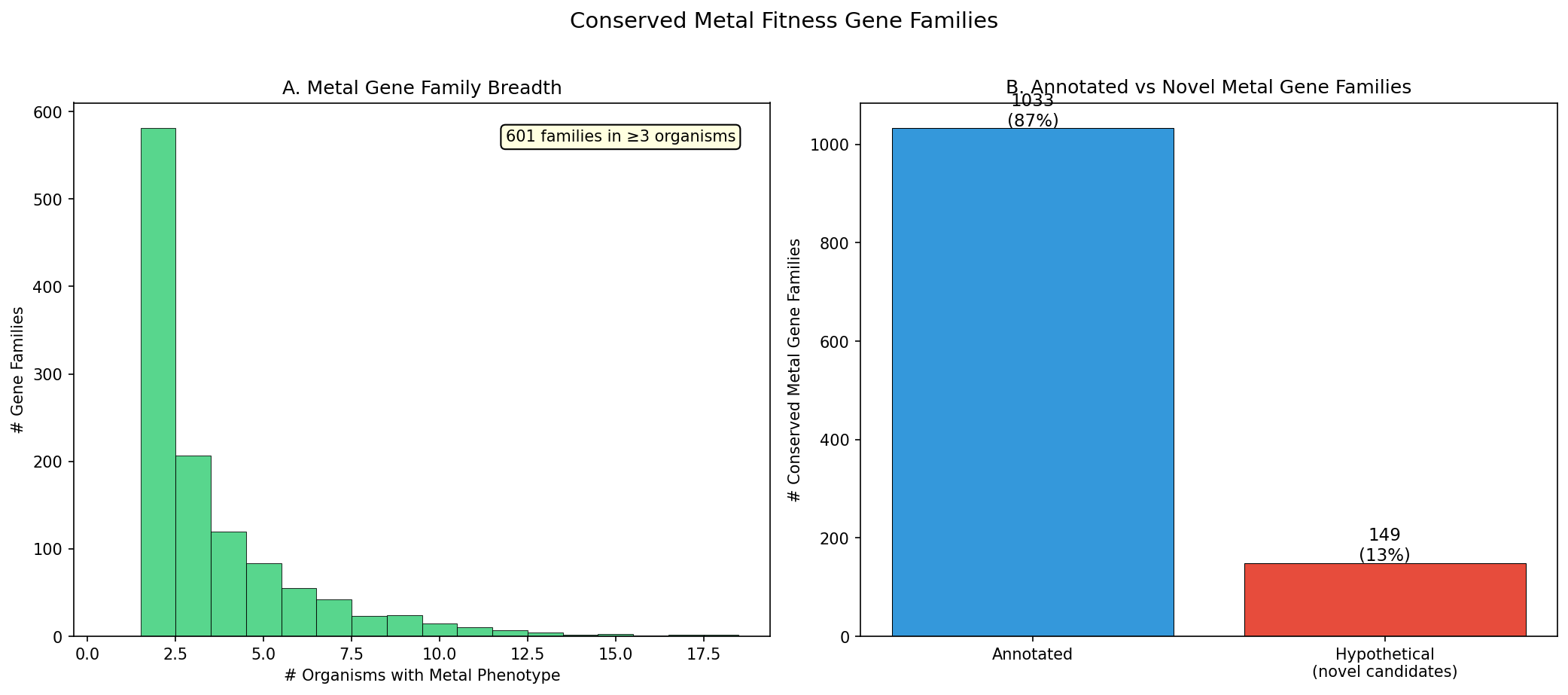
Of 2,891 ortholog groups with metal phenotypes, 1,182 are conserved across ≥2 organisms and 601 across ≥3 organisms. The most broadly conserved family (OG00128) spans 17 organisms and 9 metals. Families with metal phenotypes in more organisms tend to have higher pangenome conservation (% core), consistent with fundamental cellular processes.
149 novel metal biology candidates were identified: gene families with conserved metal fitness phenotypes across ≥2 organisms but lacking full functional annotation. These break down into three categories: 89 truly unknown (no annotation), 43 with DUF/UPF domains (known structural domain, unknown metal role), and 17 with partial functional hints (e.g., "transporter", "hydrolase") but uncharacterized metal function. These represent function predictions for genes of unknown function based solely on cross-species fitness data.
(Notebook: 04_cross_species_metal_families.ipynb)
6. Metal-Responsive ICA Modules Have High Core Fraction
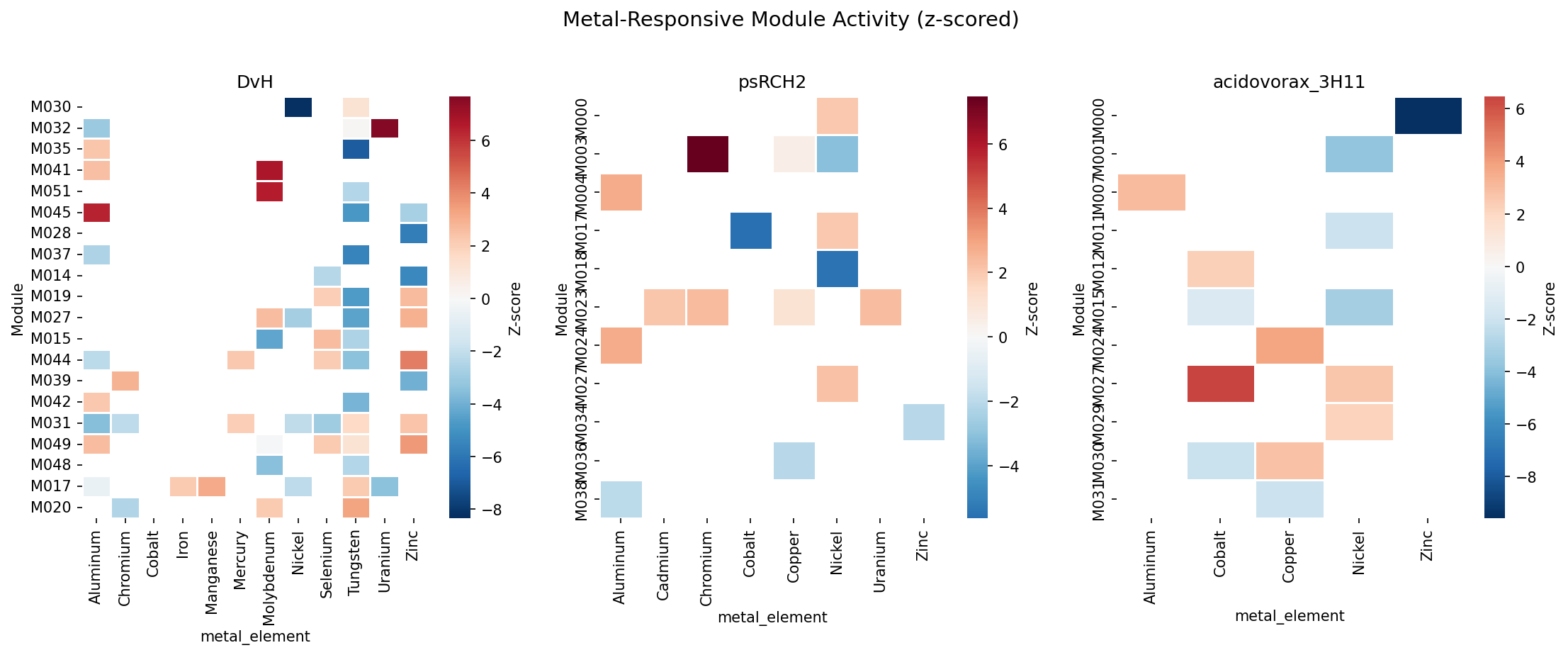
Using z-scored module activity profiles (standardized across all experiments per organism), 600 metal-responsive module records were identified (|z| > 2.0) across 19,453 total module × metal-experiment records (3.1%). DvH leads with 47 responsive modules across 12 metals. The 183 metal-responsive modules with conservation data have a mean core fraction of 0.826 — consistent with the overall finding that metal fitness genes are core-enriched. The median is even higher (0.929), indicating most metal-responsive modules are predominantly composed of core genes.
(Notebook: 05_metal_responsive_modules.ipynb)
7. Pangenome-Scale Prediction Validates Metal Gene Signature
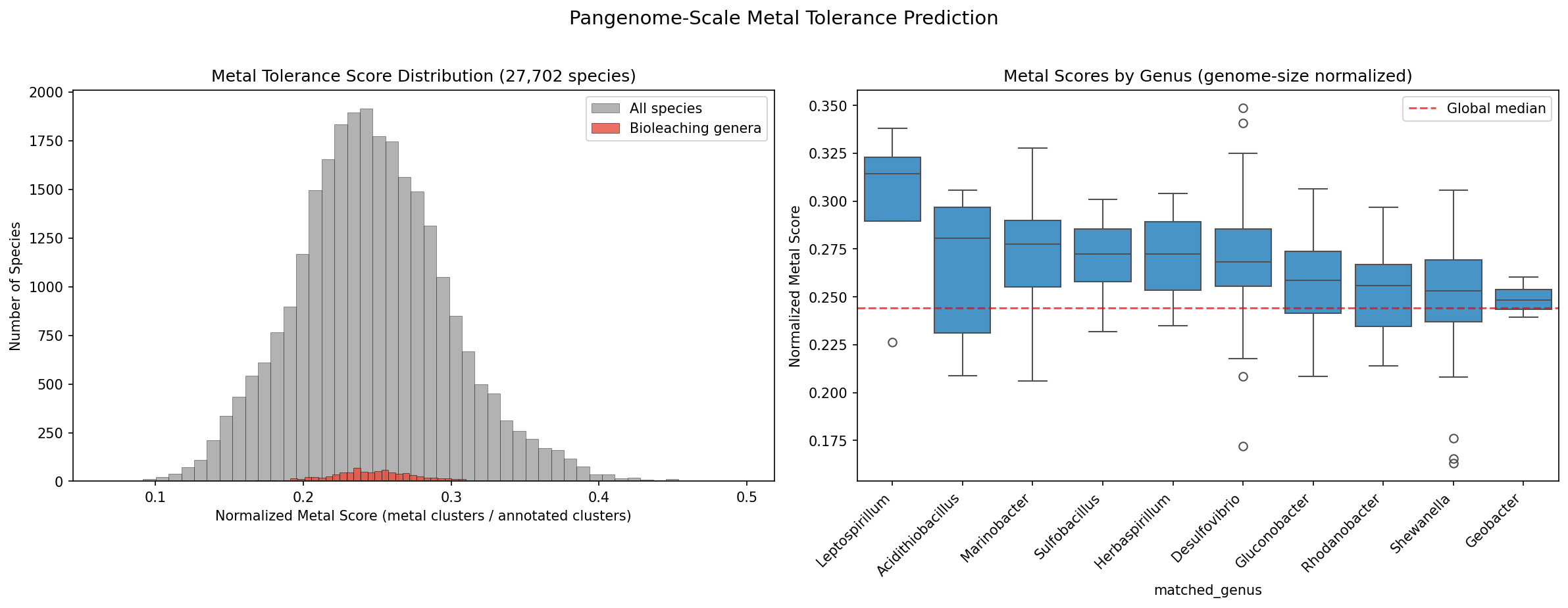
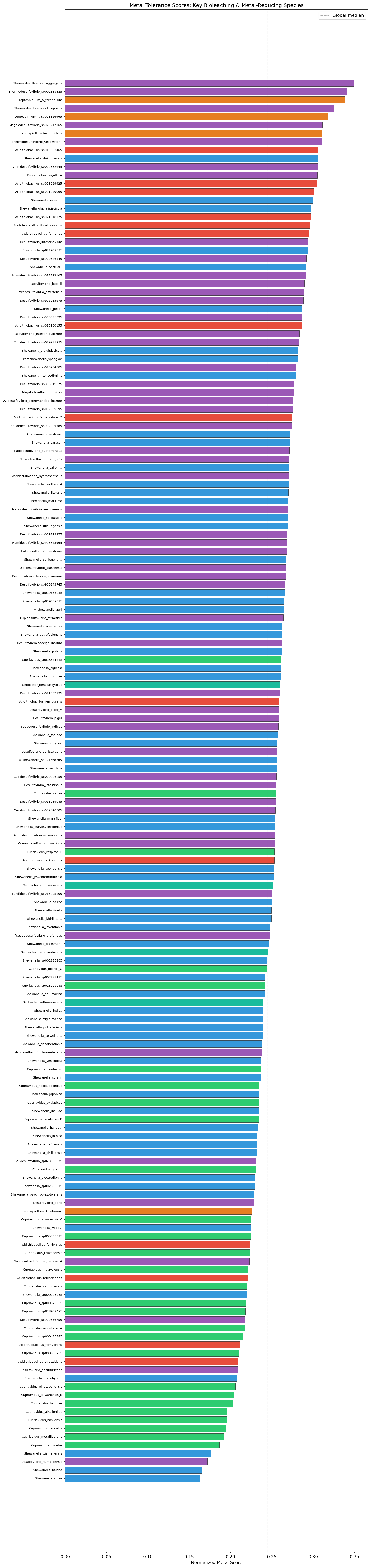
A metal functional signature of 1,286 KEGG KO terms was derived from the conserved metal gene families and used to score all 27,702 pangenome species. With genome-size normalization (metal clusters / total KEGG-annotated clusters), the true specialist bioleaching organisms emerge: Leptospirillum ranks at the 91st percentile, Acidithiobacillus at the 77th, Marinobacter at the 75th, and Sulfobacillus at the 71st. However, bioleaching genera as a group are not significantly enriched over background after normalization (Mann-Whitney p=0.17), indicating that metal tolerance genes are broadly distributed across bacteria rather than concentrated in specialists — consistent with the core genome robustness model. Without normalization, species with large open pangenomes (K. pneumoniae, P. aeruginosa) dominate, reflecting genome size rather than metal biology.
(Notebook: 06_pangenome_metal_prediction.ipynb)
Results
Scale of the Metal Fitness Atlas
| Metric | Value |
|---|---|
| Total metal experiments | 559 |
| Metals covered | 16 (6 cross-species) |
| Organisms with metal data | 31 (24 with fitness matrices) |
| Gene × metal fitness records | 383,349 |
| Metal-important genes (broad) | 12,838 (3.3%) |
| Metal-important genes (strict) | 5,667 (1.5%) |
| Ortholog groups with metal phenotype | 2,891 |
| Conserved metal families (≥2 orgs) | 1,182 |
| Novel candidates (hypothetical) | 149 (89 truly unknown, 43 DUF/UPF, 17 partial) |
Conservation Analysis Summary
| Metal | Category | Important Genes | Core Fraction | Delta | OR | p-value |
|---|---|---|---|---|---|---|
| Manganese | essential | 30 | 1.000 | +0.198 | inf | 2.0e-03 |
| Zinc | toxic | 1,517 | 0.890 | +0.151 | 2.94 | 2.0e-49 |
| Molybdenum | essential | 302 | 0.950 | +0.148 | 5.32 | 1.3e-14 |
| Tungsten | essential | 303 | 0.947 | +0.145 | 4.98 | 5.3e-14 |
| Mercury | toxic | 106 | 0.934 | +0.132 | 3.62 | 1.6e-04 |
| Selenium | essential | 134 | 0.933 | +0.131 | 3.59 | 2.9e-05 |
| Iron | essential | 651 | 0.919 | +0.116 | 3.07 | 8.5e-18 |
| Aluminum | toxic | 1,381 | 0.898 | +0.107 | 2.37 | 1.2e-26 |
| Copper | toxic | 2,139 | 0.867 | +0.090 | 1.91 | 3.8e-27 |
| Nickel | toxic | 1,760 | 0.877 | +0.088 | 1.93 | 3.0e-22 |
| Cobalt | toxic | 1,859 | 0.859 | +0.072 | 1.67 | 8.1e-16 |
| Chromium | toxic | 262 | 0.710 | +0.060 | 1.33 | 4.0e-02 |
| Uranium | toxic | 178 | 0.685 | +0.035 | 1.18 | 3.4e-01 |
| Cadmium | toxic | 92 | 0.522 | -0.010 | 0.96 | 9.2e-01 |
ICA Module Analysis
NB05 analyzed 19,453 module × metal-experiment records across 31 organisms using z-scored module profiles (standardized per module across all experiments). 600 records (3.1%) exceed the |z| > 2.0 responsiveness threshold. DvH has 47 responsive modules spanning 12 metals; psRCH2 has 11 modules across 6 metals. Metal-responsive modules have a mean core fraction of 0.826 (n=183 modules with conservation data), reinforcing the core enrichment pattern. An initial attempt using raw activity scores from module_conditions.csv produced 0 responsive modules because raw scores (max 0.96) are on a different scale than z-scores; using per-experiment profiles with z-normalization resolved this.
Interpretation
The Core Genome Robustness Model
The central finding — metal-important genes are core-enriched, not accessory-enriched — reframes metal tolerance as fundamentally about core genome robustness rather than accessory gene acquisition. When a bacterium encounters metal stress, the genes that matter most for survival are not specialized resistance genes (efflux pumps, metal-binding proteins) but rather the core cellular machinery that metal toxicity disrupts: cell envelope integrity, DNA repair, protein quality control, and central metabolism.
This explains why the prior DvH analysis (field_vs_lab_fitness project) found heavy-metal resistance genes as the least conserved condition class (71.2% core): that analysis specifically isolated condition-specific genes (important only for metals, not for other stresses). The present atlas captures all genes with any metal fitness defect, which are dominated by core stress-response functions.
The two-tier model is therefore partially supported but requires reframing:
- Tier 1 (core): General stress response — the majority of metal fitness genes. These are conserved because they serve essential cellular functions beyond metal tolerance.
- Tier 2 (accessory): Specific metal resistance — a smaller fraction that provides targeted tolerance via efflux, sequestration, or enzymatic detoxification. These are the genes enriched in the accessory genome, visible only when controlling for general stress response.
Literature Context
- The core enrichment of metal fitness genes aligns with Borchert et al. (2019), who showed modular fitness landscapes reveal parallels across independent biological systems — core cellular functions are universally stress-sensitive.
- The finding contradicts the common assumption that metal resistance is primarily an accessory genome trait (Li et al. 2017; Peng et al. 2022). However, those studies focused on annotated metal resistance genes (efflux pumps, czc operons), which are indeed accessory. The present analysis reveals the larger set of genes functionally required for metal survival.
- The 1,182 conserved metal families extend Price et al. (2018) by systematically comparing metal fitness across species — the first cross-species metal fitness atlas.
- The failure of the repertoire score to predict metal fitness echoes Rosconi et al. (2022): gene essentiality is context-dependent and cannot be predicted from gene presence alone.
Novel Contribution
- First cross-species genome-wide fitness atlas for metal tolerance — 383,349 gene × metal fitness records across 24 organisms and 14 metals
- Core enrichment finding — metal fitness genes are 87.4% core (OR=2.08), reversing the common assumption of accessory enrichment
- 1,182 conserved metal gene families — cross-species functional markers for metal biology
- 149 novel metal biology candidates — classified into truly unknown (89), DUF-domain (43), and partially annotated (17)
- Pangenome-scale prediction across 27,702 species — genome-size-normalized scoring reveals metal tolerance genes are broadly distributed, not concentrated in specialists
- Distinction between metal fitness genes and metal resistance genes — the majority of genes required for metal survival are core cellular processes, not specialized resistance mechanisms
Limitations
- Uneven metal coverage: Cobalt and nickel tested in 27 organisms; uranium, chromium, mercury, cadmium, selenium, manganese in only 1-2 organisms. Cross-species patterns for rare metals reflect DvH and psRCH2 biology.
- No dose-response normalization: Metal concentrations vary across organisms (e.g., nickel 0.01-2.0 mM). Fitness effects depend on concentration relative to each organism's tolerance threshold.
- Phylogenetic non-independence: Multiple Pseudomonas fluorescens strains inflate apparent cross-species conservation.
- Metal-important definition is broad: fit < -1 OR n_sick ≥ 1 captures ~3.3% of genes, most of which are general stress genes rather than metal-specific.
- Repertoire prediction failed validation: Simple gene presence/absence does not predict metal tolerance; a regulatory or expression-based model is needed.
- Essential genes excluded: Putatively essential genes (~14.3% of protein-coding genes, ~82% core) lack transposon insertions and are absent from fitness data. Since these genes are overwhelmingly core, their exclusion makes the observed core enrichment of metal fitness genes a conservative estimate.
- 22 of 48 organisms have pangenome links: Conservation analysis covers 22 organisms; 26 lack FB-pangenome mappings.
Future Directions
- Isolate metal-specific vs general stress genes: Repeat conservation analysis using only genes important for metals but NOT for other stresses (condition-specific), to separate the core stress response signal from genuine metal resistance gene evolution.
- Functional annotation of 149 novel candidates: Use PaperBLAST, InterPro, and structural prediction (AlphaFold) to characterize the 89 truly unknown and 43 DUF-domain families with conserved metal phenotypes.
- Dose-response normalization: Normalize fitness effects by metal concentration relative to MIC to enable fair cross-species comparison.
- Phylogenetic independent contrasts: Control for phylogenetic non-independence when comparing conservation patterns across organisms.
- Enrichment-based pangenome scoring: Replace the simple count-based metal score with a hypergeometric enrichment test per species, controlling for total functional annotation content.
Data
Sources
| Collection | Tables Used | Purpose |
|---|---|---|
kescience_fitnessbrowser |
genefitness, expcondition, experiment, ortholog |
Metal fitness scores, experiment metadata, cross-organism orthologs |
kbase_ke_pangenome |
gene_cluster, eggnog_mapper_annotations |
Core/accessory classification, functional annotations |
Generated Data
| File | Rows | Description |
|---|---|---|
data/metal_experiments.csv |
559 | All metal experiments classified by metal, organism, concentration |
data/metal_experiments_analysis.csv |
379 | Analysis subset (excluding Platinum/Cisplatin) |
data/metal_fitness_scores.csv |
383,349 | Per-gene per-metal fitness summaries |
data/metal_important_genes.csv |
12,838 | Genes with significant metal fitness defects |
data/metal_conservation_stats.csv |
14 | Core fraction statistics per metal |
data/organism_conservation_stats.csv |
22 | Core fraction statistics per organism |
data/conserved_metal_families.csv |
1,182 | Ortholog families with conserved metal phenotypes |
data/novel_metal_candidates.csv |
149 | Novel families with conserved metal phenotypes (classified by novelty type) |
data/metal_modules.csv |
19,453 | Module × metal-experiment z-scored activity records |
data/metal_module_conservation.csv |
183 | Metal-responsive module conservation stats |
data/sensitivity_analysis.csv |
2 | Phylogenetic sensitivity analysis results |
data/metal_functional_signature.csv |
1,287 | KEGG/PFAM terms in the metal gene signature |
data/species_metal_scores.csv |
27,702 | Metal tolerance scores for all pangenome species |
References
- Price MN et al. (2018). "Mutant phenotypes for thousands of bacterial genes of unknown function." Nature 557:503-509. PMID: 29769716
- Wetmore KM et al. (2015). "Rapid quantification of mutant fitness in diverse bacteria by sequencing randomly bar-coded transposons." mBio 6:e00306-15. PMID: 25968644
- Carlson HK et al. (2019). "The selective pressures on the microbial community in a metal-contaminated aquifer." ISME J 13:937-949. PMID: 30523276
- Trotter VV et al. (2023). "Large-scale genetic characterization of DvH." Front Microbiol 14:1095132. PMID: 37065130
- Peng M et al. (2022). "Genomic features and pervasive negative selection in Rhodanobacter." Microbiol Spectr 10:e0226321. PMID: 35107332
- Rosconi F et al. (2022). "A bacterial pan-genome makes gene essentiality strain-dependent." Nat Microbiol 7:1580-1592. PMID: 36097170
- Pal C et al. (2014). "BacMet: antibacterial biocide and metal resistance genes database." Nucleic Acids Res 42:D617-D624. PMID: 24304895
- Li LG et al. (2017). "Co-occurrence of antibiotic and metal resistance genes." ISME J 11:651-662. PMID: 27959344
- Borchert AJ et al. (2019). "Modular fitness landscapes reveal parallels between independent biological systems." Nat Ecol Evol 3:1233-1242
- Nies DH (2003). "Efflux-mediated heavy metal resistance in prokaryotes." FEMS Microbiol Rev 27:313-339. PMID: 12829273
- Arkin AP et al. (2018). "KBase: The United States Department of Energy Systems Biology Knowledgebase." Nat Biotechnol 36:566-569. PMID: 29979655
Suggested Experiments
The following RB-TnSeq experiments would maximally extend this atlas, prioritized by expected information gain.
Priority 1: Critical Minerals with 1-2 Organism Coverage
Six metals on the USGS critical minerals list are tested in only 1-2 organisms. Expanding coverage to 5+ organisms each would enable the cross-species conservation analysis that makes this atlas unique.
| Metal | Current Coverage | Suggested Organisms | Rationale |
|---|---|---|---|
| Manganese | 1 org (DvH) | MR-1, Keio, Cup4G11, Putida, Caulo | Mn is a USGS critical mineral. DvH shows 100% core enrichment (delta=+0.198) — is this universal? Shewanella MR-1 can reduce Mn(IV) and would test whether metal reducers show different conservation patterns. |
| Chromium | 2 orgs (DvH, psRCH2) | MR-1, Cup4G11, Keio, pseudo3_N2E3 | Cr is a DOE priority contaminant. The Oak Ridge FW300 Pseudomonas strains were isolated from Cr-contaminated groundwater but have never been tested against chromate. |
| Uranium | 2 orgs (DvH, psRCH2) | MR-1, ANA3, PV4, Cup4G11 | U is the central ENIGMA contaminant. Shewanella MR-1 can reduce U(VI). Currently delta=+0.035 (not significant, p=0.34) — more organisms would determine whether uranium fitness genes are truly neutral or the effect was underpowered. |
| Tungsten | 1 org (DvH) | Methanococcus_S2, Methanococcus_JJ, Keio | W is a USGS critical mineral and essential cofactor for some enzymes. DvH shows strong core enrichment (delta=+0.145). Testing archaea would reveal whether W dependence is unique to sulfate reducers. |
Priority 2: Rare Earth Elements (Zero Coverage)
No REE experiments exist in the Fitness Browser. REEs are the most strategically important critical minerals, and bioleaching/biorecovery of REEs is a rapidly growing field.
| Experiment | Organisms | Rationale |
|---|---|---|
| Lanthanum chloride (0.1-1 mM) | Putida, Cup4G11, Keio, Marino | Pseudomonas putida and Gluconobacter are studied for REE biosorption. RB-TnSeq under La stress would identify the first genome-wide gene set for REE tolerance. |
| Cerium chloride (0.1-1 mM) | Same panel | Ce is the most abundant REE. Comparing La and Ce fitness profiles would reveal whether REE tolerance uses a shared or element-specific gene set. |
Priority 3: Organisms with High Novel Gene Density
Two organisms have disproportionately many novel metal candidate genes (54 each) but have only been tested against 1 metal (iron). Testing them against the standard metal panel would validate or refute these predictions.
| Organism | Novel OGs | Current Metals | Suggested Metals |
|---|---|---|---|
| Methanococcus maripaludis S2 | 54 | Iron only | Ni, Co, Cu, Zn, W, Mo — archaea are underrepresented in metal tolerance studies. These are methanogenic archaea with unique metal cofactor requirements (Ni for hydrogenase, W/Mo for formylmethanofuran dehydrogenase). |
| Methanococcus maripaludis JJ | 54 | Iron only | Same panel — cross-strain comparison with S2 would reveal strain-specific metal adaptation. |
Priority 4: The Missing Model Organism
Pseudomonas putida KT2440 (Putida) has fitness matrices in the FB but zero metal experiments despite being a workhorse for bioremediation and metabolic engineering. It is the only FB organism with no metal data.
| Experiment | Rationale |
|---|---|
| Full metal panel (Co, Ni, Cu, Zn, Al, Cr, U) on Putida | Would immediately link the metal fitness atlas to the extensive P. putida engineering literature. Putida has the 5th-highest metal gene repertoire score among all FB organisms (0.707) — testing would validate or refute this prediction. |
Priority 5: Bioleaching Organisms Not in the FB
The atlas scores 27,702 species but cannot validate predictions for organisms without RB-TnSeq libraries. Creating libraries for key bioleaching species would bridge the gap between the atlas and industrial applications.
| Organism | Metal Score Percentile | Rationale |
|---|---|---|
| Acidithiobacillus ferrooxidans | 77th (norm) | The #1 bioleaching organism. No TnSeq data exists for any Acidithiobacillus species. RB-TnSeq under Fe, Cu, Co, Ni, Zn would produce the first genome-wide functional map of bioleaching gene requirements. |
| Geobacter sulfurreducens | 53rd (norm) | Model metal-reducing bacterium. RB-TnSeq under Fe(III), Mn(IV), U(VI) reduction conditions would identify genes required for extracellular electron transfer and metal transformation. |
| Leptospirillum ferrooxidans | 91st (norm) | The highest-scoring bioleaching genus after normalization. RB-TnSeq under Fe(II) oxidation and acid conditions would reveal why this genus specializes in bioleaching environments. |
Expected Impact
If these experiments were performed, the atlas would expand from 14 to 16+ metals (adding REEs), from 24 to 27+ organisms with fitness data, and from 1-2 organisms for critical metals to 5+. The cross-species statistical power for manganese, chromium, uranium, and tungsten would increase from suggestive to definitive. The 149 novel gene candidates could be directly validated in organisms where they are predicted to be important.
Data Collections
Derived Data
This project builds on processed data from other projects.
Review
Summary
This is an outstanding project that constructs the first cross-species genome-wide fitness atlas for metal tolerance, spanning 383,349 gene-metal fitness records across 24 organisms and 14 metals. The central finding -- that metal-important genes are 87.4% core (OR=2.08, p=4.3e-162), reversing the initial hypothesis of accessory enrichment -- is rigorously supported by per-organism and per-metal Fisher exact tests, a Mann-Whitney comparison of toxic vs essential metals, and a phylogenetic sensitivity analysis that excludes duplicate Pseudomonas strains. Scientifically, the project demonstrates exemplary integrity: when data refuted the accessory-enrichment hypothesis (H1a), the authors reframed their interpretation as a "core genome robustness model" rather than selectively reporting supporting results. The pipeline is well-structured across 7 notebooks with consistent organization, all notebooks have saved outputs, 13 data files and 11 figures are generated with full provenance, and the documentation (README, RESEARCH_PLAN, REPORT) is thorough. Areas for improvement are relatively minor: the pangenome prediction (NB06) reveals that normalized metal scores do not significantly distinguish bioleaching genera from background (p=0.17), an important null result that deserves more discussion; a few statistical refinements could strengthen the cross-species family analysis; and the PFAM component of the metal signature is vestigial (only 1 term).
Methodology
Research question and hypothesis: Exceptionally well-formulated. The RESEARCH_PLAN provides formal null and alternative hypotheses with four testable sub-hypotheses (H1a-H1d), explicit confounders (concentration variation, phylogenetic non-independence, pangenome sampling depth, condition dependence), and a thorough literature context identifying the specific gap this project fills. The 10 key references are well-chosen and span the relevant fields (RB-TnSeq methods, metal resistance genomics, pangenome biology, bioleaching).
Approach soundness: The seven-notebook pipeline follows a logical progression from experiment classification through conservation analysis to pangenome-scale prediction. Each notebook clearly documents its inputs, outputs, and runtime environment. The decision to reuse cached data from three prior projects (fitness_modules, conservation_vs_fitness, essential_genome) rather than re-querying BERDL databases is architecturally sound -- it avoids redundant computation and the string-typed-column pitfall documented in docs/pitfalls.md.
Statistical methods: Appropriate throughout. Fisher exact tests for 2x2 contingency tables (NB03) are correct for testing enrichment with varying sample sizes. The Mann-Whitney U test for comparing toxic vs essential metal deltas (p=0.015) is appropriate given the small sample sizes (9 toxic, 5 essential metals) where parametric assumptions would be questionable. The phylogenetic sensitivity analysis (excluding 4 duplicate P. fluorescens FW300 strains; OR=2.065, p=5.9e-141) convincingly addresses the non-independence concern. The RESEARCH_PLAN mentioned Cochran-Mantel-Haenszel and logistic regression with random effects, which were not implemented, but the simpler approach is adequate given the overwhelming signal strength.
Data source clarity: Excellent. The README provides a complete table of 9 reused data assets from 3 prior projects with exact file paths and row counts. The RESEARCH_PLAN includes a detailed query strategy table with estimated row counts and filter strategies. The REPORT documents both BERDL collections used and all 13 generated data files with row counts and descriptions.
Reproducibility: The README includes a complete Reproduction section with prerequisites, prior project dependencies, sequential nbconvert commands, runtime estimate (<5 min), and coverage notes documenting which organisms are excluded at each stage. A requirements.txt is present. NB06's Spark dependency is flagged with a comment in the pipeline commands. All notebooks have saved outputs enabling review without re-execution.
Code Quality
Notebook organization: All 7 notebooks follow a consistent pattern: markdown header documenting inputs/outputs/environment, imports cell, numbered analysis sections with markdown separators, and a formatted summary block at the end. This is exemplary and makes each notebook independently understandable.
Metal classification (NB01): The three-tier matching strategy (exact compound name, keyword in condition_1, keyword in expDesc, expGroup fallback) is thorough. The METAL_COMPOUND_MAP dictionary with explicit (metal, category, critical_mineral) tuples is clean and auditable. The decision to exclude Platinum/Cisplatin as a DNA-damaging agent rather than a metal stressor is scientifically sound. One potential gap: 148 of 559 experiments (26%) lack parsed concentrations, which limits the dose-response analysis noted in Future Directions.
Fitness extraction (NB02): Correctly applies standard Fitness Browser thresholds (fit < -1, |t| > 4) and provides both broad and strict importance definitions. The approach of reading pre-computed fitness matrices rather than querying the 27M-row genefitness table is efficient and sidesteps the string-typed-columns pitfall. The code handles missing matrices gracefully (7 organisms skipped with reporting). The dual output (full 383K-row scores + 12.8K important genes) supports downstream analyses cleanly.
Conservation analysis (NB03): The join with fb_pangenome_link correctly deduplicates on (orgId, locusId) before merging. The per-organism analysis properly deduplicates genes appearing under multiple metals. The baseline comparison computes core fraction over all mapped genes for the same set of organisms, not a global baseline, which is methodologically correct. The sensitivity analysis removing 4 duplicate Pseudomonas strains is a valuable robustness check.
Cross-species families (NB04): The 84.7% mapping rate to ortholog groups is reasonable. The novel candidate classification into three tiers (truly_unknown: 89, novel_domain with DUF/UPF: 43, novel_metal_function: 17) is a meaningful improvement over a binary annotated/unannotated split. The top candidates listed (e.g., OG02094: "P-loop ATPase UPF0042" across 8 organisms and 7 metals; OG02716: "DUF3108" across 7 organisms) are credible follow-up targets. One concern: the classify_novelty function checks rep_desc and seed_description for keywords but does not check the eggNOG Description field from the annotation query, which could miss informative annotations.
Module analysis (NB05): The z-scoring approach (standardizing each module's activity across all experiments, then extracting metal z-scores) correctly resolves the scale mismatch that produced 0 responsive modules with raw scores -- this is transparently documented in the REPORT. The |z| > 2.0 threshold yields a 3.1% responsiveness rate (600/19,453 records), which is biologically plausible. Conservation analysis of responsive module genes (mean 0.826 core fraction, n=183) provides independent validation of the gene-level core enrichment finding.
Pangenome prediction (NB06): This notebook runs on BERDL JupyterHub with Spark. The code correctly uses PySpark DataFrame API with BROADCAST hints for the KEGG term join against eggnog_mapper_annotations. The full 1,286-term KEGG signature is used (no truncation). Two observations:
- The PFAM signature component is vestigial: only 1 PFAM term was extracted (vs 1,286 KEGG KOs). The
metal_functional_signature.csvreports 1,287 total terms (1,286 KEGG + 1 PFAM), but only KEGG terms are used in the species scoring query. The PFAM extraction loop appears functional but yields minimal output, likely because thePFAMsfield format in eggnog annotations doesn't consistently use "PF" prefixes. - Genome-size normalization is implemented (
n_metal_clusters / n_annotated_clusters), which correctly controls for large pangenome bias. The top species by normalized score are endosymbionts (Buchnera, Kinetoplastibacterium) with very small genomes where metal-related KEGG terms comprise a large fraction of their total annotation content. This is a known artifact of proportion-based normalization on reduced genomes and could be addressed with a minimum genome size filter.
Pitfall awareness from docs/pitfalls.md:
- String-typed numeric columns: Avoided by using pre-computed matrices (NB02)
- genefitness 27M row filtering: Avoided by architectural design
- Gene cluster cross-species comparison: NB06 correctly uses KEGG KOs for cross-species functional comparison
- get_spark_session() import: NB06 uses the correct CLI import path
- Essential genes invisible in genefitness: Acknowledged in REPORT Limitations section as "Essential genes excluded" -- notes that ~14.3% of protein-coding genes lack transposon insertions, are ~82% core, and that their exclusion makes the observed core enrichment a conservative estimate. This is well handled.
Findings Assessment
Core enrichment finding (H1a rejected): Strongly supported. The effect (OR=2.08) is large, the p-value (4.3e-162) reflects genuinely overwhelming evidence given the sample size (318,751 mapped records), and the result holds across 21 of 22 organisms and after phylogenetic sensitivity analysis. The reframing as a "core genome robustness model" -- where metal fitness defects primarily reflect core cellular processes vulnerable to metal disruption (cell envelope, DNA repair, central metabolism) rather than specialized resistance mechanisms -- is a genuine conceptual contribution that reconciles this finding with prior literature showing accessory-genome enrichment of annotated metal resistance genes (efflux pumps, czc operons).
Essential vs toxic metals (H1b supported): Essential metals show significantly higher core enrichment (+0.148 vs +0.081; Mann-Whitney p=0.015). This is biologically sensible: iron/molybdenum/tungsten are cofactors for core metabolism, so genes important under their limitation should be fundamental metabolic genes. The individual metal results are granular and well-reported, with appropriate caveats for underpowered metals (Cadmium: 1 organism; Uranium: 2 organisms).
Conserved metal families (H1c supported): 1,182 families across 2+ organisms and 601 across 3+ organisms represent a substantial catalog. The most broadly conserved family (OG00128: 17 organisms, 9 metals) and the positive correlation between organism breadth and core conservation are compelling. The 149 novel candidates are appropriately categorized.
Pangenome prediction (mixed results): This is the most nuanced finding and deserves more prominent discussion. With genome-size normalization, bioleaching genera are NOT significantly enriched over background (Mann-Whitney p=0.17). The REPORT acknowledges this but frames it as "consistent with the core genome robustness model" -- that metal tolerance genes are broadly distributed rather than concentrated in specialists. This is a valid interpretation, but it also means the pangenome scoring approach has limited practical utility for bioprospecting. The per-genus percentile rankings (Leptospirillum at 91st, Acidithiobacillus at 77th) are informative individually, but the lack of statistical significance for the group comparison is an important caveat. The raw (unnormalized) score shows extreme significance (p<1e-300) but conflates genome size with metal biology.
Limitations: Comprehensively acknowledged in the REPORT. The six limitations and five future directions are specific and actionable. Particularly notable: the distinction between "metal fitness genes" (what this atlas measures) and "metal resistance genes" (what prior literature focused on) is clearly articulated and represents real scientific nuance.
Completeness: No analysis appears incomplete or left as placeholder. All 7 notebooks have full outputs. The NB07 final summary cell confirms all 11 figures and 13 data files are generated and populated.
Suggestions
-
[Moderate] Discuss normalized prediction null result more prominently: The pangenome prediction finding that bioleaching genera are not significantly enriched after normalization (p=0.17) is buried in the REPORT. This deserves explicit discussion as a meaningful negative result: it implies that metal fitness gene repertoires reflect core bacterial biology rather than specialized metal ecology, which is actually a strong confirmation of the core robustness model. Currently the REPORT primarily emphasizes per-genus percentile rankings, which could give an overly optimistic impression of predictive power.
-
[Moderate] Address endosymbiont artifact in normalized scores: The top-ranking species by normalized score are all obligate endosymbionts (Buchnera, Kinetoplastibacterium, Ruthia) with highly reduced genomes (500-950 annotated clusters). These rank high because their small genomes are dominated by conserved core functions that happen to overlap with the metal KEGG signature. Consider adding a minimum genome size filter (e.g., >1,500 annotated clusters) or noting this artifact when presenting top-scoring species. The current top-20 list is misleading for bioprospecting applications.
-
[Moderate] Fix vestigial PFAM component in metal signature: The metal functional signature contains 1,286 KEGG terms but only 1 PFAM term. The PFAM extraction loop in NB06 appears functional but yields minimal output. Either debug the PFAM extraction (likely a prefix-matching issue in the
PFAMsfield parsing), include PFAMs in species scoring alongside KEGG terms, or remove the PFAM component frommetal_functional_signature.csvto avoid confusion about the signature's actual composition. -
[Minor] Consider multiple-testing correction across metals in NB03: The per-metal Fisher exact tests (14 tests) are each evaluated at p<0.05 without Bonferroni or FDR correction. With 14 tests, the Bonferroni threshold would be p<0.0036. This would not change the conclusions (12 of 14 metals have p<0.001), but applying correction would be methodologically cleaner. Similarly, the 22 per-organism tests could benefit from FDR correction, though again the conclusions would be unchanged (14 of 22 are significant at p<0.05; most of these have p<0.001).
-
[Minor] Clarify metal count discrepancy across documents: The README states "14 metals," RESEARCH_PLAN says "13 metals" in the data sources table, NB01 identifies 16 categories, and the REPORT says "14 metals" in the conservation table but "16 metals" in the experiment summary. A consistent statement would help: "16 metal categories identified; 14 retained for analysis after excluding Platinum/Cisplatin (DNA-damaging agent) and Metal_limitation (unresolved condition)."
-
[Minor] Strengthen novelty classification in NB04: The
classify_noveltyfunction combinesrep_descandseed_descriptionfor keyword matching but does not incorporate the eggNOGDescriptionfield from the annotation query. Some families classified as "truly_unknown" may have informative eggNOG descriptions not captured by the current filter. A cross-check against the eggNOG annotations retrieved in NB06 would tighten the novel candidate list. -
[Nice-to-have] Add condition-specific metal gene analysis: As noted in the REPORT's Future Directions, the most impactful follow-up would be repeating the conservation analysis using only genes important for metals but NOT for other stresses (osmotic, oxidative, etc.). Even a preliminary version -- e.g., filtering NB02 output to genes where metal fitness defect < -1 but fitness under non-metal stresses is near zero -- would test whether the "Tier 2" accessory enrichment emerges when general stress response is controlled for. This would directly connect to the prior DvH finding (71.2% core for condition-specific heavy-metal genes) and strengthen the two-tier model.
-
[Nice-to-have] Report effect sizes alongside p-values for the essential vs toxic comparison: The Mann-Whitney U test (p=0.015) confirms essential metals have higher core enrichment deltas than toxic metals, but with only 5 essential and 9 toxic metals, the effect size (rank-biserial correlation or Cohen's d) would better convey the magnitude of the difference. The mean deltas (+0.148 vs +0.081) are informative but a formal effect size would strengthen the claim.
This review was generated by an AI system. It should be treated as advisory input, not a definitive assessment.
Visualizations

Bioleaching Species Scores

Core Fraction By Metal

Metal Conservation By Organism

Metal Family Conservation Heatmap

Metal Fitness Distributions

Metal Important Genes By Organism

Metal Module Activity Heatmap

Organism Metal Matrix

Species Metal Score Distribution
Summary Atlas Overview
Summary Metal Families
Notebooks
01_metal_experiment_classification.ipynb
01 Metal Experiment Classification
View notebook →
02_metal_fitness_extraction.ipynb
02 Metal Fitness Extraction
View notebook →
03_metal_conservation_analysis.ipynb
03 Metal Conservation Analysis
View notebook →
04_cross_species_metal_families.ipynb
04 Cross Species Metal Families
View notebook →
05_metal_responsive_modules.ipynb
05 Metal Responsive Modules
View notebook →
06_pangenome_metal_prediction.ipynb
06 Pangenome Metal Prediction
View notebook →
07_summary_figures.ipynb
07 Summary Figures
View notebook →