Field vs Lab Gene Importance in Desulfovibrio vulgaris Hildenborough
CompletedResearch Question
Which genes matter for survival under environmentally-realistic conditions but appear dispensable in the lab, and vice versa? Do field-relevant fitness effects predict pangenome conservation better than lab-only effects?
Overview
DvH has 757 RB-TnSeq experiments spanning environmentally-relevant stresses (uranium, mercury, nitrate, sulfate reduction) and standard lab conditions (antibiotics, rich media). This project classifies these experiments by ecological relevance and tests whether genes important under field-relevant conditions show different pangenome conservation patterns than genes important under lab-only conditions. It builds on conservation_vs_fitness (FB-pangenome link, essential genes) and fitness_modules (ICA modules, condition activity scores).
Key results: field-stress and field-core genes are significantly enriched in the core genome (OR=1.58 and 1.46, FDR q<0.03), but fitness magnitude matters more than condition type for overall prediction (10-fold CV-AUC 0.52-0.55 for field/lab fitness vs 0.65 with gene length). Antibiotic and heavy-metal resistance genes trend below baseline (73% and 71% core vs 76%). Module-level analysis identifies 21 "ecological" modules (0.98 core fraction) and 9 "lab" modules (0.52 core fraction). ENIGMA CORAL database contains no DvH data. See REPORT.md for full findings and interpretation.
Key Findings
ENIGMA CORAL Contains No DvH Fitness Data (NB01)
The ENIGMA CORAL database (47 tables, enigma_coral on BERDL) was surveyed for complementary data. Key finding: DvH is completely absent from the database. The single TnSeq library is for FW300-N2E2 (Pseudomonas), DubSeq libraries cover E. coli, P. putida, and B. thetaiotaomicron. All 6,705 genomes and 15,015 genes are environmental isolates from Oak Ridge, not DvH. The database does contain 4,346 field samples with geochemistry data (uranium, metals) across 596 Oak Ridge locations, and 213,044 ASVs for community composition -- potentially useful for future analyses, but not for gene-level fitness analysis.
(Notebook: 01_enigma_discovery.ipynb)
Condition Classification (NB02)
757 DvH experiments classified into 6 categories:
| Category | Experiments | % | Rationale |
|---|---|---|---|
| Lab-nutrient | 237 | 31.3% | Amino acids, organic acids, carbon sources |
| Field-core | 204 | 26.9% | DvH primary metabolism (sulfate, lactate, formate, pyruvate, H2) |
| Lab-other | 140 | 18.5% | Lab reagents (DMSO, PEG, osmotic stress, iron chelators) |
| Field-stress | 78 | 10.3% | FRC contaminants (uranium, mercury, nitrate, nitrite, oxygen, NO) |
| Heavy-metals | 55 | 7.3% | Metal stress (cobalt, nickel, zinc, copper, manganese, selenium) |
| Lab-antibiotic | 43 | 5.7% | Antibiotics and respiratory inhibitors |
Broad split: 337 field (44.5%) vs 420 lab (55.5%).
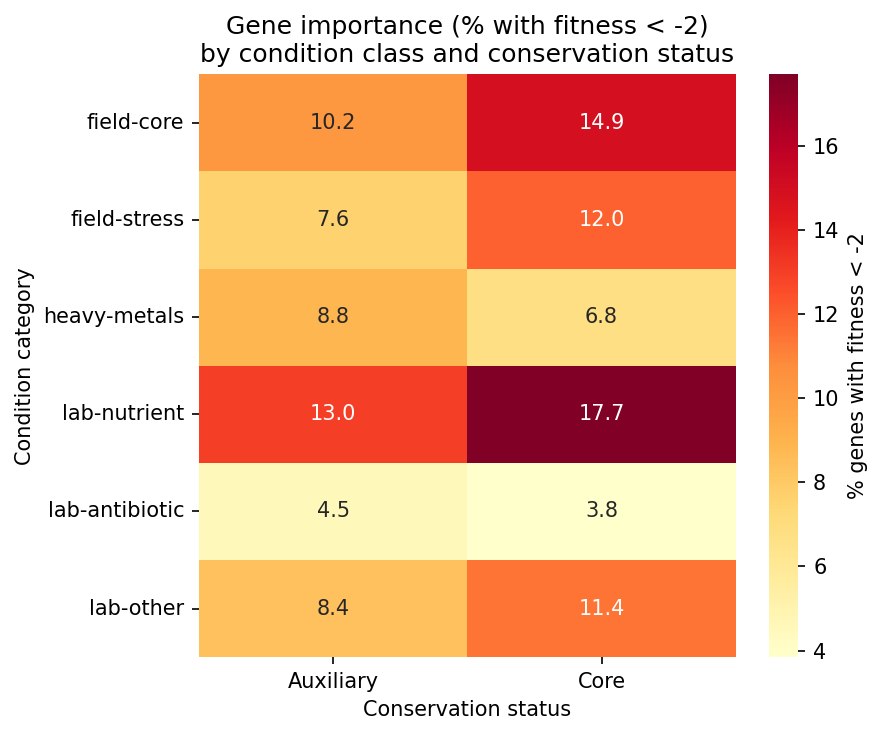
(Notebook: 02_condition_classification.ipynb)
Genes Important for Field Conditions Are Significantly More Conserved (NB03)
Of 2,725 non-essential genes with both fitness data and pangenome links, 76.3% are core overall. An additional 678 essential genes (80.1% core) are excluded because they lack fitness data (no transposon mutants recovered). Genes important (fitness < -2) under different condition classes show varying conservation:
| Condition class | Important genes | Core % | OR vs baseline | FDR q |
|---|---|---|---|---|
| Field-stress | 298 | 83.6% | 1.58 | 0.026 |
| Field-core | 376 | 82.4% | 1.46 | 0.026 |
| Lab-other | 292 | 81.5% | 1.37 | 0.073 |
| Lab-nutrient | 452 | 81.4% | 1.36 | 0.037 |
| Lab-antibiotic | 109 | 73.4% | 0.86 | 0.49 |
| Heavy-metals | 198 | 71.2% | 0.77 | 0.14 |
| Essential (no fitness) | 678 | 80.1% | -- | -- |
| All genes (baseline) | 2,725 | 76.3% | -- | -- |
Field-stress (q=0.026), field-core (q=0.026), and lab-nutrient (q=0.037) genes are significantly enriched in the core genome after BH-FDR correction. Heavy-metals and lab-antibiotic genes trend below baseline but are not significant.
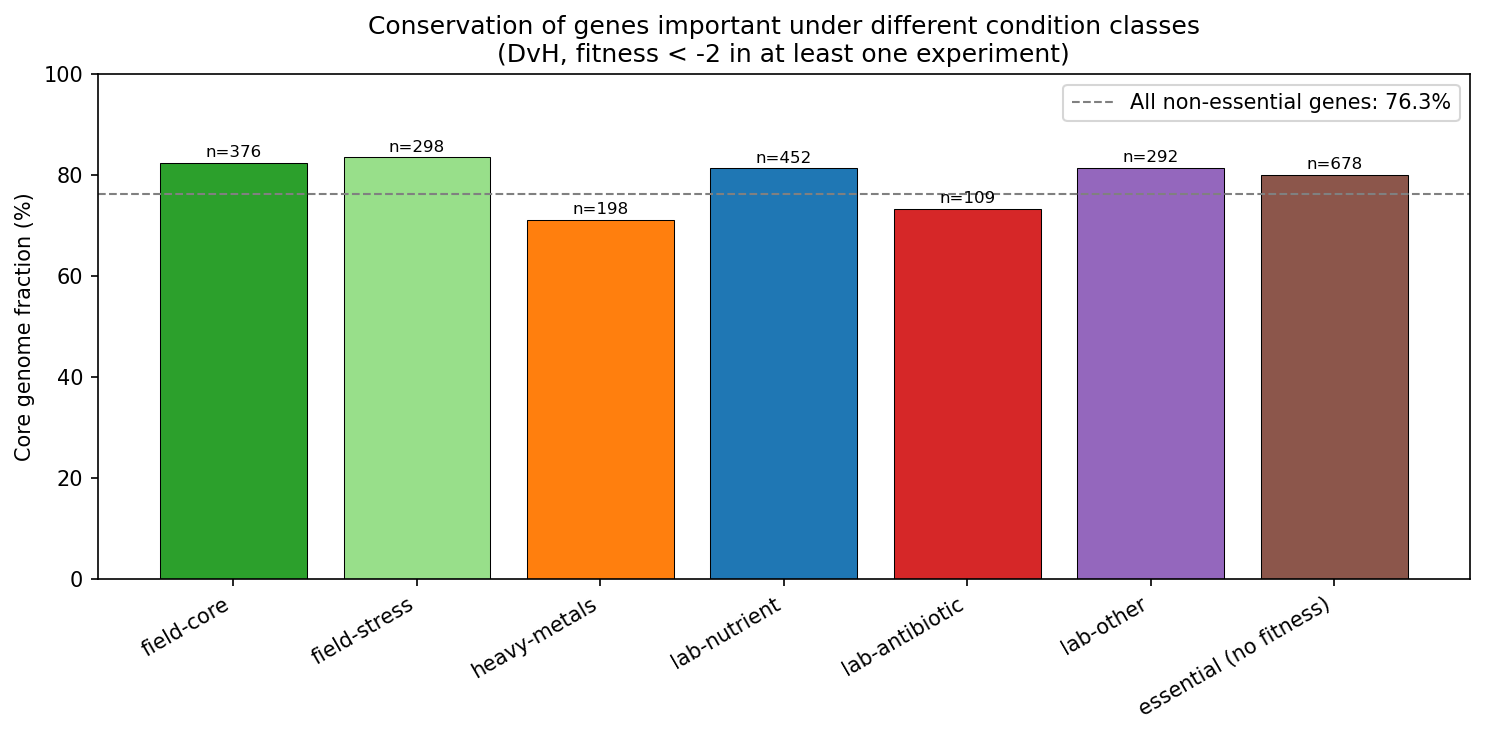
(Notebook: 03_fitness_conservation.ipynb)
Specificity Analysis: Lab-Specific Genes Are Surprisingly More Core
| Specificity | Genes | Core % |
|---|---|---|
| Lab-specific (sick in lab only) | 50 | 96.0% |
| Field-specific (sick in field only) | 52 | 88.5% |
| Field-biased | 89 | 83.1% |
| Universal (sick in both) | 352 | 79.8% |
| Neutral (no strong effects) | 2,083 | 74.5% |
Counter to H1, genes with fitness defects only under lab conditions are 96% core (n=50), slightly higher than field-specific genes at 88.5% (n=52). Both are well above the 74.5% baseline, suggesting that any fitness importance -- regardless of ecological context -- predicts conservation. The field-specific vs lab-specific comparison is not statistically significant (Fisher exact OR=0.32, p=0.27). The universal vs neutral comparison is significant (OR=1.35, p=0.033).
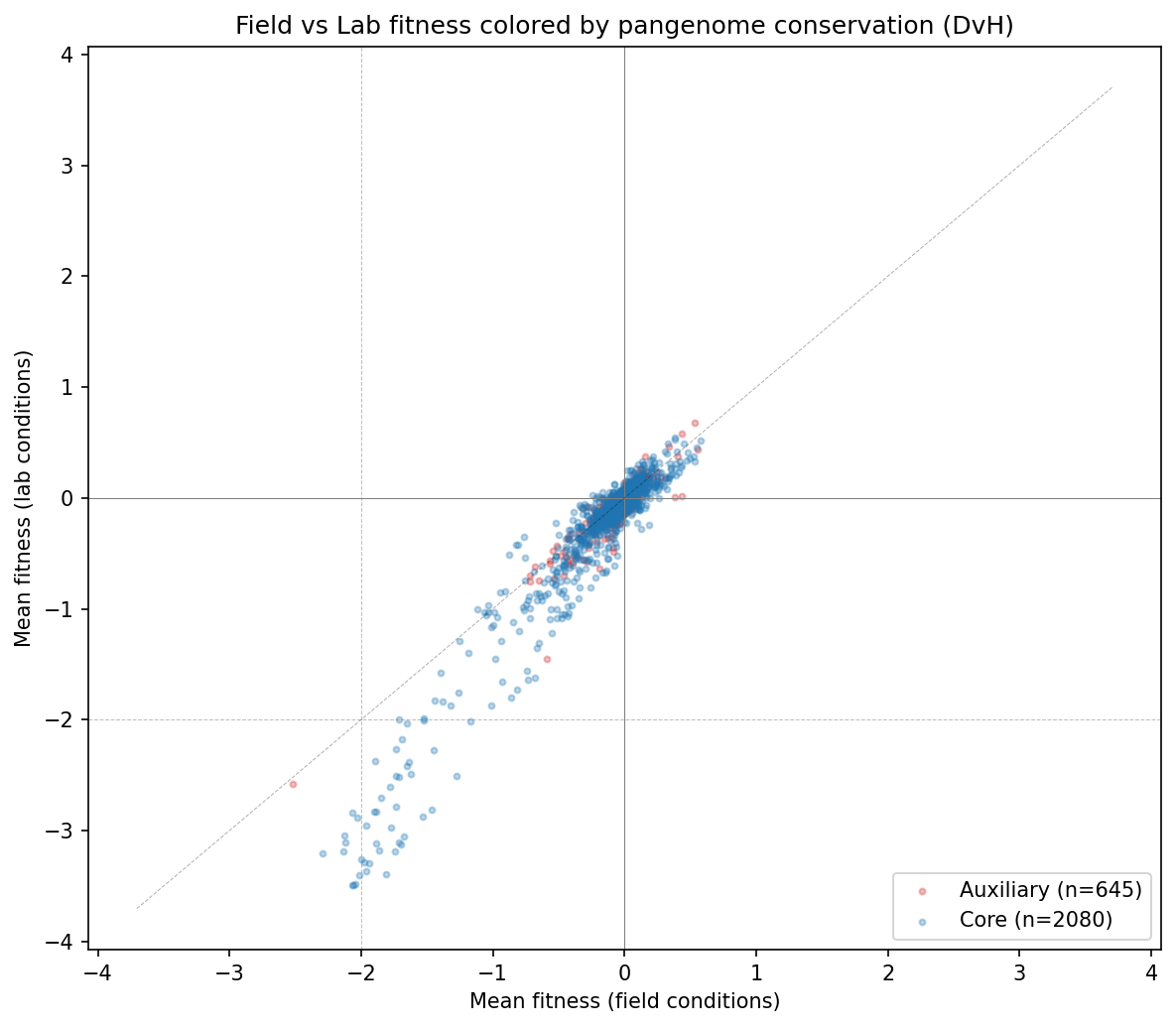
(Notebook: 03_fitness_conservation.ipynb)
Fitness Effects Are Weak Predictors of Core Status
Logistic regression with 10-fold cross-validated AUC:
| Model | CV-AUC | Std |
|---|---|---|
| Field fitness only | 0.517 | 0.052 |
| Lab fitness only | 0.531 | 0.052 |
| Field + Lab | 0.548 | 0.053 |
| Full (+ gene length) | 0.645 | 0.068 |
Gene length is a much stronger predictor of core status than fitness effects from either field or lab conditions. Neither fitness dimension alone is informative (CV-AUC near 0.5). Cross-validation confirms the in-sample estimates are not inflated.
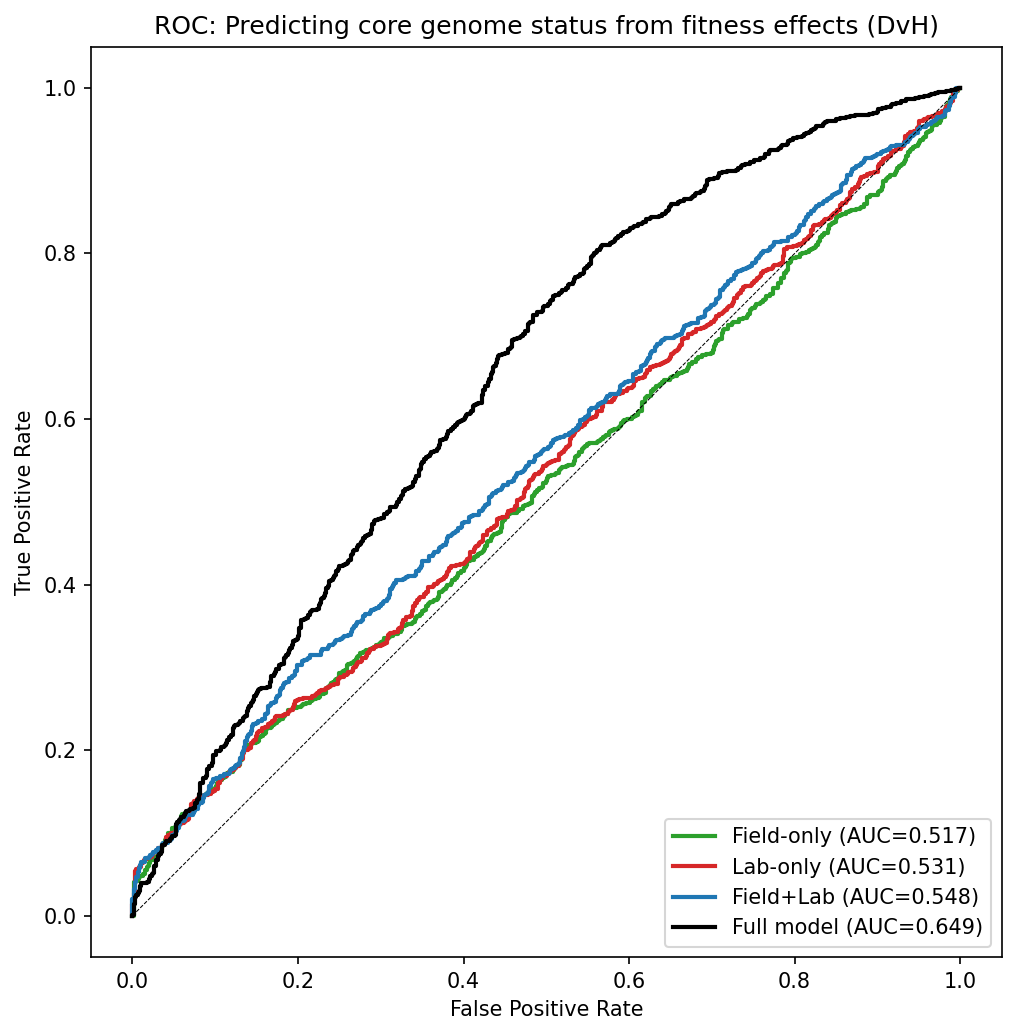
(Notebook: 03_fitness_conservation.ipynb)
Threshold Sensitivity Analysis
The pattern is robust across fitness thresholds (-1 to -3):
| Condition class | -3.0 | -2.0 | -1.5 | -1.0 |
|---|---|---|---|---|
| Field-stress | 89.4% | 83.6% | 84.0% | 82.1% |
| Field-core | 84.4% | 82.4% | 81.9% | 79.0% |
| Lab-nutrient | 83.5% | 81.4% | 81.0% | 79.5% |
| Lab-other | 85.2% | 81.5% | 80.1% | 77.5% |
| Lab-antibiotic | 82.2% | 73.4% | 77.3% | 76.2% |
| Heavy-metals | 70.6% | 71.2% | 76.7% | 76.7% |
Field-stress consistently has the highest conservation across all thresholds. Heavy-metals is consistently the lowest. The lab-antibiotic dip at -2 is driven by a small sample (n=109 at -2 vs n=45 at -3).
(Notebook: 03_fitness_conservation.ipynb)
Module-Level Conservation Shows No Field-Lab Difference (NB04)
Of 52 ICA fitness modules, the mean core fraction is 0.886 and the median is 1.000. No significant correlation exists between field condition activity and module conservation (Spearman rho=0.071, p=0.62). Using the mean core fraction (0.886) as the classification threshold, modules partition into:
| Module type | Count | Mean core fraction |
|---|---|---|
| Ecological (field-active + conserved) | 21 | 0.980 |
| Conserved-quiet (low field activity + conserved) | 17 | 0.983 |
| Field-variable (field-active + less conserved) | 5 | 0.829 |
| Lab (low field + less conserved) | 9 | 0.516 |
The 9 "lab" modules (mean core fraction 0.516) are notably less conserved, containing genes in the accessory genome. The 21 ecological modules contain 239 member genes, of which 52 are unannotated -- candidates for novel environmental adaptation functions.
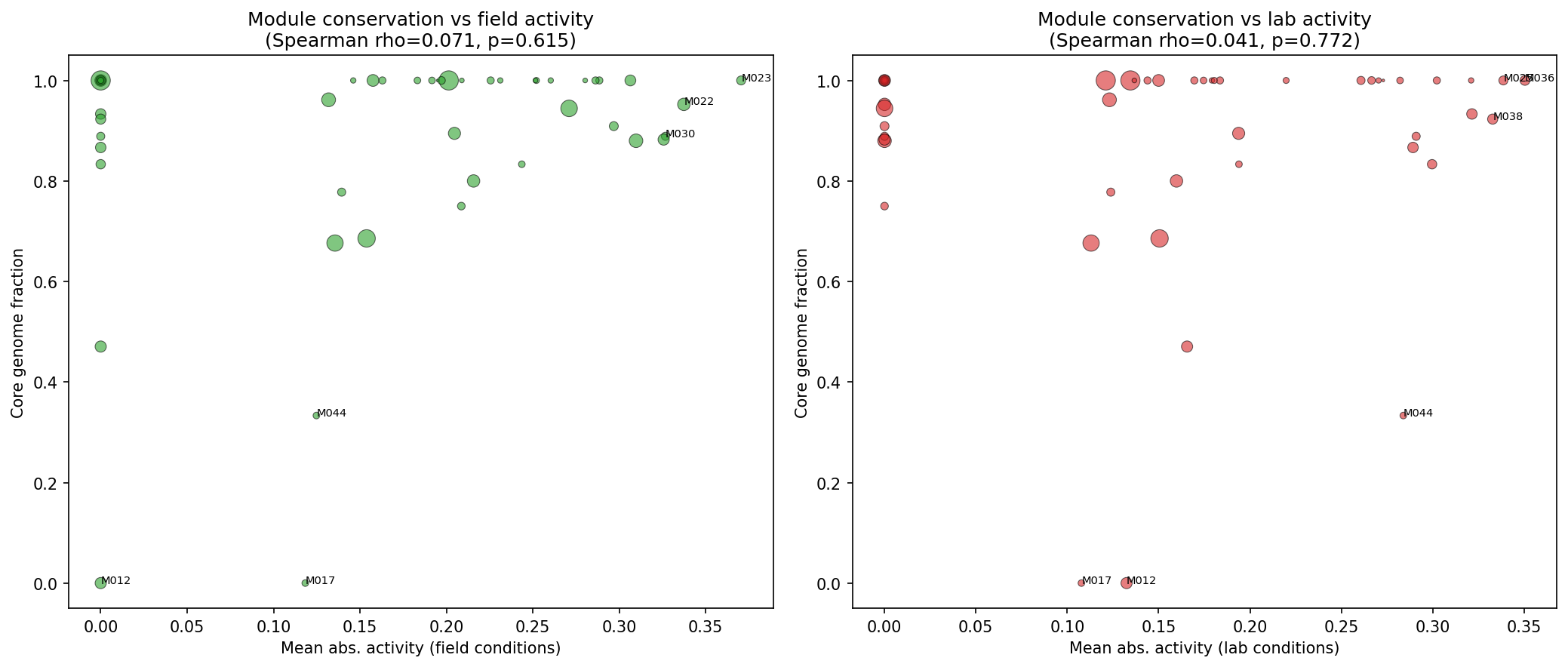
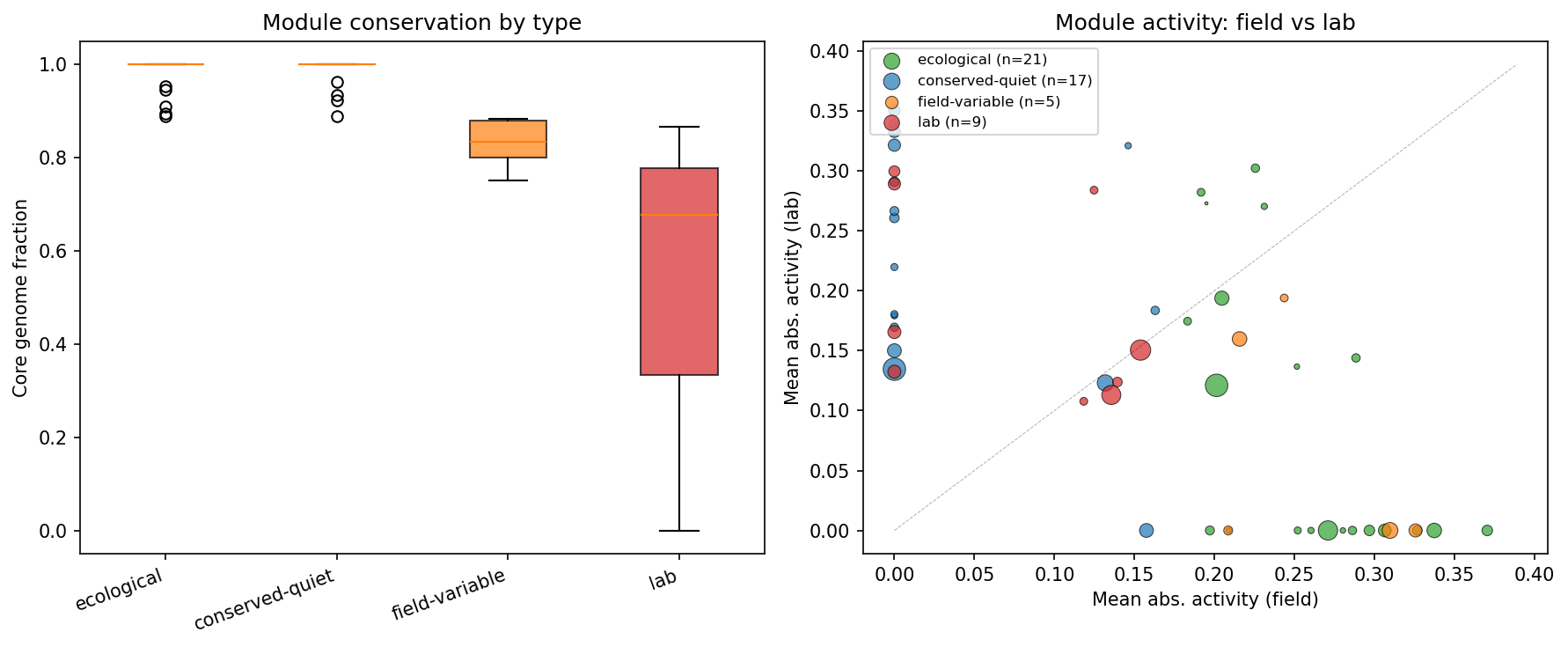
(Notebook: 04_module_analysis.ipynb)
Interpretation
Hypothesis Outcomes
- H1 partially supported: Genes important for field-stress conditions are significantly more conserved than baseline (83.6% core, OR=1.58, FDR q=0.026), as are field-core genes (82.4%, OR=1.46, q=0.026). However, lab-nutrient genes are also significantly enriched (81.4%, q=0.037), and the field-specific vs lab-specific gene comparison is not significant (Fisher exact OR=0.32, p=0.27, n=50-52 per group). The strongest signal is that any fitness importance predicts conservation: universally important genes are significantly more core than neutral genes (OR=1.35, p=0.033). Threshold sensitivity analysis confirms field-stress genes are consistently the most conserved across thresholds from -1 to -3.
- H2 not supported: Module-level conservation does not correlate with field condition activity (Spearman rho=0.071, p=0.62). However, the revised module classification (using mean rather than median core fraction) reveals 9 "lab" modules with strikingly low conservation (mean core fraction 0.516), suggesting that accessory-genome modules do exist and tend to respond to lab-type conditions.
- H3 partially supported: 21 "ecological" modules (field-active + conserved, mean core fraction 0.980) contain 239 genes, of which 52 are unannotated candidates for environmental adaptation. These are distinct from the 9 "lab" modules, supporting the concept of a functional core relevant to environmental survival.
Key Biological Insight
The most striking finding is that lab-antibiotic and heavy-metal resistance genes are the least conserved (73.4% and 71.2%), well below the 76.3% baseline. This suggests that resistance functions are disproportionately in the accessory genome, consistent with these being recently acquired adaptive traits carried on mobile genetic elements. Conversely, genes for core metabolism (sulfate reduction, lactate/formate/pyruvate utilization) and FRC-relevant stress responses (uranium, mercury, nitrate) are deeply conserved, reflecting their importance for DvH's ecological niche.
Note on heavy-metals classification: we classified heavy-metals (cobalt, nickel, zinc, copper, manganese, selenium, molybdate, tungstate, aluminum) as "field" in the broad category because DvH encounters these metals at Oak Ridge FRC sites. However, the low conservation of heavy-metal-important genes (71.2%) contrasts with the high conservation of uranium/mercury-important genes (in field-stress at 83.6%). This may reflect that specific metal resistance mechanisms (efflux pumps, metal-binding proteins) are accessory traits, while the uranium and mercury response involves more fundamental stress pathways (DNA repair, sulfate reduction itself) that are part of the core genome.
Literature Context
- Trotter et al. (2023) generated the comprehensive barcoded transposon library for DvH used in this analysis, identifying essential genes and condition-specific phenotypes across hundreds of conditions. Our work adds a pangenome conservation dimension to their fitness measurements.
- Rosconi et al. (2022) showed that gene essentiality in S. pneumoniae is strain-dependent and influenced by accessory genome composition. This parallels our finding that fitness importance is a better predictor of conservation than the specific ecological context of the condition -- the magnitude of fitness effect matters more than which condition produces it.
- Lee et al. (2015) identified 352 general and 199 condition-specific essential genes in P. aeruginosa across six growth media, establishing the concept of condition-dependent essentiality. Our analysis extends this framework by testing whether the ecological relevance of conditions predicts evolutionary conservation.
- Akusobi et al. (2025) found 259 core essential genes and 425 differentially required genes across 21 M. abscessus clinical isolates, demonstrating that genetic diversity drives significant functional differences. This supports our finding that fitness effects vary by condition class, though conservation patterns do not cleanly separate field from lab conditions.
- Shi et al. (2021) showed that pre-existing genetic variation in DvH influences chromate tolerance, with the chromate transporter DVU0426 identified as crucial for Cr(VI) resistance -- a specific example of accessory-genome-encoded environmental adaptation.
- Huang et al. (2022) found that genomic islands and prophages are "major drivers for evolution and environmental adaptation" in S. algae, consistent with our finding that metal and antibiotic resistance genes are enriched in the accessory genome.
- Price et al. (2018) generated the Fitness Browser data underpinning this analysis. Our work tests a prediction implicit in their dataset: whether the ecological relevance of experimental conditions modulates the fitness-conservation relationship.
Novel Contribution
This is the first analysis to stratify RB-TnSeq fitness effects by ecological relevance and test whether field-relevant conditions predict pangenome conservation differently than lab conditions. The finding that condition type matters less than fitness magnitude for conservation prediction was not anticipated and suggests that the core genome reflects general functional importance rather than niche-specific selection. The accessory-genome enrichment of antibiotic and metal resistance genes, however, identifies a clear functional partition that aligns with ecological expectations.
Limitations
- 678 essential genes (80.1% core) are excluded from the fitness analysis because they lack transposon mutants. Including them would raise the overall baseline slightly but would not change condition-class comparisons (which are among non-essential genes only)
- Single organism (DvH) limits generalizability -- results may differ in organisms with larger accessory genomes
- The Nitratidesulfovibrio vulgaris pangenome has relatively few genomes, creating a coarse core/auxiliary classification with a high baseline core fraction (76.3%), which compresses effect sizes
- Condition classification relies on manual mapping of
condition_1labels; edge cases (e.g., zinc sulfate as metal vs sulfate source) required subjective judgment - The fitness < -2 threshold is the primary cutoff, but sensitivity analysis across -1 to -3 confirms pattern robustness
- Gene length is confounded with both fitness measurement quality (short genes get fewer transposon insertions) and core status (core genes tend to be longer)
- The field-specific and lab-specific gene sets are small (n=50-52), limiting statistical power for the key comparison
- Field and lab fitness effects are correlated (r ~ 0.7 from the scatter plot), meaning most genes that are sick in one context are sick in the other
Future Directions
- Multi-organism extension: Apply the field vs lab classification to other ENIGMA organisms (e.g., Pseudomonas FW300 strains) that have both environmental relevance and Fitness Browser data, to test whether the pattern generalizes
- Finer-grained conservation metric: Replace binary core/auxiliary with quantitative conservation (fraction of genomes carrying the gene cluster) to increase statistical power
- ENIGMA community data integration: Use the 213K ASVs and 4,346 field samples in ENIGMA CORAL to ask whether Desulfovibrio abundance at Oak Ridge sites correlates with the geochemistry conditions tested in the Fitness Browser
- Accessory resistance gene characterization: Deeper analysis of the antibiotic and heavy-metal resistance genes in the accessory genome -- are they on mobile elements? Recently acquired? Shared with co-occurring species at Oak Ridge?
- Quantitative fitness scores: Replace binary important/not-important with continuous fitness scores (e.g., mean or minimum fitness per condition class) as predictors in logistic regression
Data
Sources
| Collection | Description |
|---|---|
kescience_fitnessbrowser |
RB-TnSeq fitness data for DvH (757 experiments, 2,741 genes) |
kbase_ke_pangenome |
Pangenome gene clusters with core/auxiliary/singleton classification |
Generated Data
| File | Description |
|---|---|
experiment_classification.csv |
757 experiments with category and broad_category assignments |
gene_fitness_conservation.csv |
2,725 genes with per-category fitness stats, conservation, and specificity class |
module_characterization.csv |
52 ICA modules with conservation, field/lab activity, type, and top annotation |
References
- Price MN, Wetmore KM, Waters RJ, Calef M, Tber J, Engel A, Chol J, Arkin AP, Deutschbauer AM. (2018). "Mutant phenotypes for thousands of bacterial genes of unknown function." Nature 557:503-509. DOI: 10.1038/s41586-018-0124-0. PMID: 29769716
- Trotter VV, Mirasol R, Yung M, Chandran A, Kuehl JV, Katsnelson B, De Leon KB, Wall JD, Deutschbauer AM. (2023). "Large-scale genetic characterization of the model sulfate-reducing bacterium, Desulfovibrio vulgaris Hildenborough." Front Microbiol 14:1095132. PMID: 37065130
- Rosconi F, Rudmann E, Li J, Surujon D, Anthony J, Frank M, Jones DS, Rock C, Bhatt AS, Roditi I, van Opijnen T. (2022). "A bacterial pan-genome makes gene essentiality strain-dependent and evolvable." Nat Microbiol 7:1580-1592. DOI: 10.1038/s41564-022-01208-7. PMID: 36097170
- Lee SA, Gallagher LA, Thongdee M, Stodghill BJ, Hachey SJ, Sanowar S, Cooley RB, Leung KP. (2015). "General and condition-specific essential functions of Pseudomonas aeruginosa." Proc Natl Acad Sci USA 112:5189-5194. PMID: 25848053
- Akusobi C, Benoit BM, Cavallo K, Williams EP, Bhatt A, Rubin EJ. (2025). "Transposon-sequencing across multiple Mycobacterium abscessus isolates reveals significant functional genomic diversity among strains." mBio 16:e02488-24. PMID: 39745363
- Shi W, Zhan T, Guo Y, Zhou A, He Z, Deng Y. (2021). "Genetic basis of chromate adaptation and the role of pre-existing genetic divergence during an experimental evolution study with Desulfovibrio vulgaris populations." mSystems 6:e00351-21. PMID: 34061571
- Huang Z, Yu K, Fu S, Xiao Y, Wei Q, Wang D. (2022). "Genomic analysis reveals high intra-species diversity of Shewanella algae." Microb Genom 8:000786. PMID: 35143386
- Borchert AJ, Ernst DC, Downs DM. (2019). "Modular fitness landscapes reveal parallels between independent biological systems." Nat Ecol Evol 3:1233-1242. DOI: 10.1038/s41559-019-0938-7
Data Collections
Derived Data
This project builds on processed data from other projects.
Review
Summary
This is a well-designed and honestly reported project that asks whether genes important under environmentally-realistic conditions (uranium, mercury, sulfate reduction) show different pangenome conservation patterns than genes important under lab-only conditions (antibiotics, rich media). The analysis pipeline is clean and logically structured across four notebooks, the condition classification scheme is thoughtfully grounded in DvH ecology at the Oak Ridge FRC, and the statistical analyses are mostly appropriate. The project arrives at a nuanced answer: field-stress genes are modestly more conserved (83.6% core, OR=1.58, FDR q=0.026), but fitness magnitude matters more than condition type for predicting conservation (CV-AUC 0.52-0.55 for condition-type models vs 0.65 with gene length). Documentation is exemplary -- the README, RESEARCH_PLAN, and REPORT form a coherent three-file structure with clear data provenance, thorough limitations, and honest reporting of largely null results. The main areas for improvement are: (1) a missing data-saving step for gene_fitness_conservation.csv, (2) statsmodels is an undeclared dependency, and (3) the module-level correlation analysis in NB04 is inherently limited by the ceiling effect (median core fraction = 1.0 across 52 modules).
Methodology
Research question: Clearly stated and testable. Three hypotheses (H1-H3) are explicitly articulated alongside a null hypothesis (H0), which is a strong practice. The two-level approach (gene-level in NB03 and module-level in NB04) provides complementary perspectives.
Approach: Sound overall. The six-category condition classification (NB02) is well-designed with 52 pattern-matching rules and sensible fallback logic using expGroup. The classification is validated with a cross-tabulation (NB02 cell 8) showing that field-core aligns with respiratory growth and stress, while lab categories align with nutrient and nitrogen source. The specificity analysis (field-specific vs lab-specific vs universal genes) is a valuable addition that reveals the counter-intuitive finding that lab-specific genes are 96% core.
Data sources: All upstream data is clearly identified and properly attributed to prior projects (conservation_vs_fitness, fitness_modules). The ENIGMA CORAL discovery (NB01) is a responsible first step that documents the absence of DvH data in that database, avoiding incorrect assumptions.
Reproducibility concerns:
- The condition classification rules are embedded in code (NB02 cell 3) rather than an external configuration file. This is acceptable for a one-off analysis but makes it harder to version or modify the classification independently.
- Edge cases in classification (e.g., "zinc sulfate" as metal vs sulfate) are handled by rule ordering but documented only implicitly through the code structure.
Essential genes handling: NB03 correctly acknowledges that 678 essential genes are excluded from fitness analyses (cell 8 prints a detailed note explaining why and reports their 80.1% core fraction). They are also included as a separate bar in Figure 1 and as a row in the condition importance table (cell 10). This is a significant strength -- the analysis is transparent about what it cannot measure.
Code Quality
Notebook organization: All four notebooks follow a clean setup → data loading → analysis → visualization → summary flow. Markdown headers and summary cells at the end of each notebook consolidate key findings. NB03 is particularly well-structured with 8 clearly delineated analysis sections.
Statistical methods:
- Fisher exact tests with BH-FDR correction (NB03 cell 10) are appropriate for the 6 condition-class comparisons. The multiple testing correction is a strength.
- Logistic regression with 10-fold cross-validated AUC (NB03 cell 18) is properly implemented using sklearn.model_selection.cross_val_score. Both in-sample and CV-AUC are reported, and the CV results confirm the weak signal (0.517-0.548) is not an artifact of overfitting.
- Spearman correlation (NB04 cell 10) is appropriate for the module-level analysis given the non-normal distribution of core fractions.
- The threshold sensitivity analysis (NB03 cell following cell 23) strengthens the results by showing the pattern holds across thresholds from -1 to -3.
Condition classification (NB02): The rule-based system is carefully ordered (e.g., "persulfate" and "zinc sulfate" before "sulfate") with comments explaining why. The "heavy-metals" → "field" broad classification is debatable for some metals (aluminum, rubidium) but is discussed in the REPORT interpretation section.
Pitfall awareness:
- fillna(False).astype(bool) pattern correctly used in NB03 cell 4 and NB04 cell 5, per docs/pitfalls.md.
- String-to-boolean conversion for TSV columns is explicitly handled in NB03 cell 4.
- Essential gene exclusion from genefitness analyses is acknowledged, per the pitfall "Essential Genes Are Invisible in genefitness-Only Analyses."
- The .apply() on 757 rows in NB02 cell 5 is fine — the performance pitfall about row-wise apply only matters for large DataFrames.
Module classification (NB04): The classification in cell 13 uses the mean core fraction (0.886) rather than the median (1.000) as the threshold, which avoids the ceiling effect that would make the "ecological" category empty. This produces a reasonable distribution: 21 ecological, 17 conserved-quiet, 5 field-variable, and 9 lab modules. However, the underlying issue remains that 52 modules with a median core fraction of 1.0 leaves limited dynamic range for correlation analysis (Spearman rho=0.071, p=0.62).
Minor issues:
- NB03 cell 8 reports Auxiliary: 645 while the gene_info table has 583 auxiliary genes. The discrepancy (62 genes) is because genes without pangenome links (from the outer merge in cell 4) are counted as non-core in the gene_data table. The report summary correctly uses the 645 figure, but a clarifying comment would help.
Findings Assessment
Conclusions supported by data: Yes, thoroughly. The REPORT accurately represents the statistical results. H1 is honestly reported as "partially supported" — field-stress enrichment is significant (q=0.026) but the field-specific vs lab-specific comparison is not (p=0.27, n=50-52 per group). The key insight that fitness magnitude matters more than condition type is well-supported by the logistic regression results (gene length AUC=0.645 vs field/lab fitness AUC=0.517-0.548).
Limitations acknowledged: Exceptionally thorough. Eight specific limitations are listed in the REPORT, including the essential gene exclusion, single-organism caveat, coarse core/auxiliary classification, subjective condition mapping, and the correlation between field and lab fitness effects (r~0.7). The gene length confound is explicitly noted.
Counter-intuitive results honestly reported: The finding that lab-specific genes are 96% core (vs 88.5% for field-specific) is counter to H1 but is reported prominently. The interpretation that "any fitness importance predicts conservation" is well-reasoned, though the small sample sizes (n=50-52) warrant the caveat about limited statistical power.
Incomplete analysis: None. The REPORT's gene_fitness_conservation.csv is listed as a data file and exists on disk, but the save step is missing from the notebooks. All other analyses are complete.
Visualizations: Six figures are well-labeled with axes, titles, and legends. The bar chart (fig 1) clearly shows conservation gradients with sample sizes. The ROC curves (fig 3) effectively demonstrate weak predictive power. The module scatter plots (figs 5-6) honestly display the null result. Figures are saved at 150 DPI — adequate for screen use.
Literature context: Strong. Eight references are cited in the REPORT with specific connections to findings (e.g., Rosconi et al. 2022 linked to the magnitude-over-context conclusion; Huang et al. 2022 linked to accessory genome metal resistance). The references.md file lists 12 references with full bibliographic details and PubMed search queries, enabling others to reproduce the literature search.
Suggestions
-
Add
statsmodelstorequirements.txt(Critical): NB03 importsfrom statsmodels.stats.multitest import multipletestsbutstatsmodelsis not listed inrequirements.txt. Addstatsmodels>=0.13to avoid aModuleNotFoundErroron fresh installs. -
Add the missing save step for
gene_fitness_conservation.csv(Important): The file exists on disk and is listed in the REPORT, but no notebook cell writes it. Add a cell at the end of NB03's section 6 or 7 to savegene_datatodata/gene_fitness_conservation.csv. This ensures full reproducibility vianbconvert --execute. -
Add NB01 code cell outputs or a summary data file (Moderate): NB01 code cells lack saved outputs since it requires Spark, which is expected. The markdown header has an execution summary, but saving a small summary file (e.g.,
data/enigma_summary.jsonwith table names and row counts) would let downstream notebooks or readers verify the discovery results without Spark access. -
Consider a quantitative conservation metric (Moderate): The binary core/auxiliary classification creates a high baseline (76.3% core) and a ceiling effect at the module level (median 1.0). Using the fraction of genomes carrying each gene cluster as a continuous variable would increase statistical power, as noted in the REPORT's Future Directions. This is the single change most likely to strengthen the results if the pangenome data supports it.
-
Clarify the auxiliary gene count discrepancy (Minor): NB03 cell 8 reports 645 auxiliary genes, while cell 4 shows 583 in the
gene_infotable. A brief comment explaining that the difference comes from genes without pangenome links being counted as non-core after the inner merge would prevent confusion. -
Consider showing ROC curves with CV-AUC values (Minor): NB03 cell 19 plots ROC curves using in-sample probabilities but the legend labels show the in-sample AUC values (e.g., "Field-only (AUC=0.516)"). Since 10-fold CV-AUC values are computed in cell 18, updating the legend to show CV-AUC (e.g., "Field-only (CV-AUC=0.517)") would be more rigorous, though the difference is negligible here.
-
Document the heavy-metals broad classification rationale earlier (Minor): The README and RESEARCH_PLAN classify heavy-metals as "field" without explanation. The REPORT discusses this in the Interpretation section, but adding a one-line note in NB02's markdown header or classification scheme table would make the rationale visible at the point of decision.
This review was generated by an AI system. It should be treated as advisory input, not a definitive assessment.
Visualizations

Fig Condition Importance Heatmap

Fig Conservation By Condition Class

Fig Ecological Vs Lab Modules

Fig Field Vs Lab Specificity

Fig Module Conservation Vs Activity

Fig Roc Conservation Prediction