Core Gene Paradox -- Why Are Core Genes More Burdensome?
CompletedResearch Question
Why are core genome genes MORE likely to show positive fitness effects when deleted, and what functions and conditions drive this burden paradox?
Research Plan
Hypothesis
The fitness_effects_conservation project found that core genes are 1.3x more likely to be burdens than auxiliary genes (OR=0.77 for auxiliary vs core, p=5.5e-48) and that condition-specific fitness genes are more core (OR=1.78). This contradicts the "streamlining" model where accessory genes are metabolic burdens.
Key framing: Lab conditions are an impoverished proxy for nature. A gene burdensome in LB may be essential in soil or biofilm. Genes that are simultaneously costly (lab) AND conserved (pangenome) are the strongest evidence for purifying selection -- nature maintains them despite their cost.
Approach
- Dissect the burden paradox by functional category (SEED top-level categories) to determine which functions drive the core-burden excess
- Identify trade-off genes -- genes that are important (fit < -1) in some conditions and burdensome (fit > 1) in others -- and test their enrichment in core
- Construct a selection-signature matrix crossing lab fitness cost with pangenome conservation
- Analyze burden patterns by condition type (stress, carbon source, nitrogen source, etc.)
- Case studies of specific functional categories (motility, cell wall)
Revision History
- v1 (2026-02): Migrated from README.md
Overview
This project dissects why core genes are more burdensome than accessory genes -- a finding from fitness_effects_conservation that contradicts the "streamlining" model. It reveals that the paradox is function-specific (driven by motility, RNA metabolism, protein metabolism), that trade-off genes (important in some conditions, costly in others) are enriched in core, and constructs a selection-signature matrix identifying genes under purifying selection in natural environments.
Key Findings
The Burden Paradox Is Function-Specific
Not all functional categories show the paradox. Core genes are disproportionately burdensome in Protein Metabolism (+6.2pp), Motility (+7.8pp), and RNA Metabolism (+12.9pp). But Cell Wall reverses: non-core cell wall genes are MORE burdensome (-14.1pp).
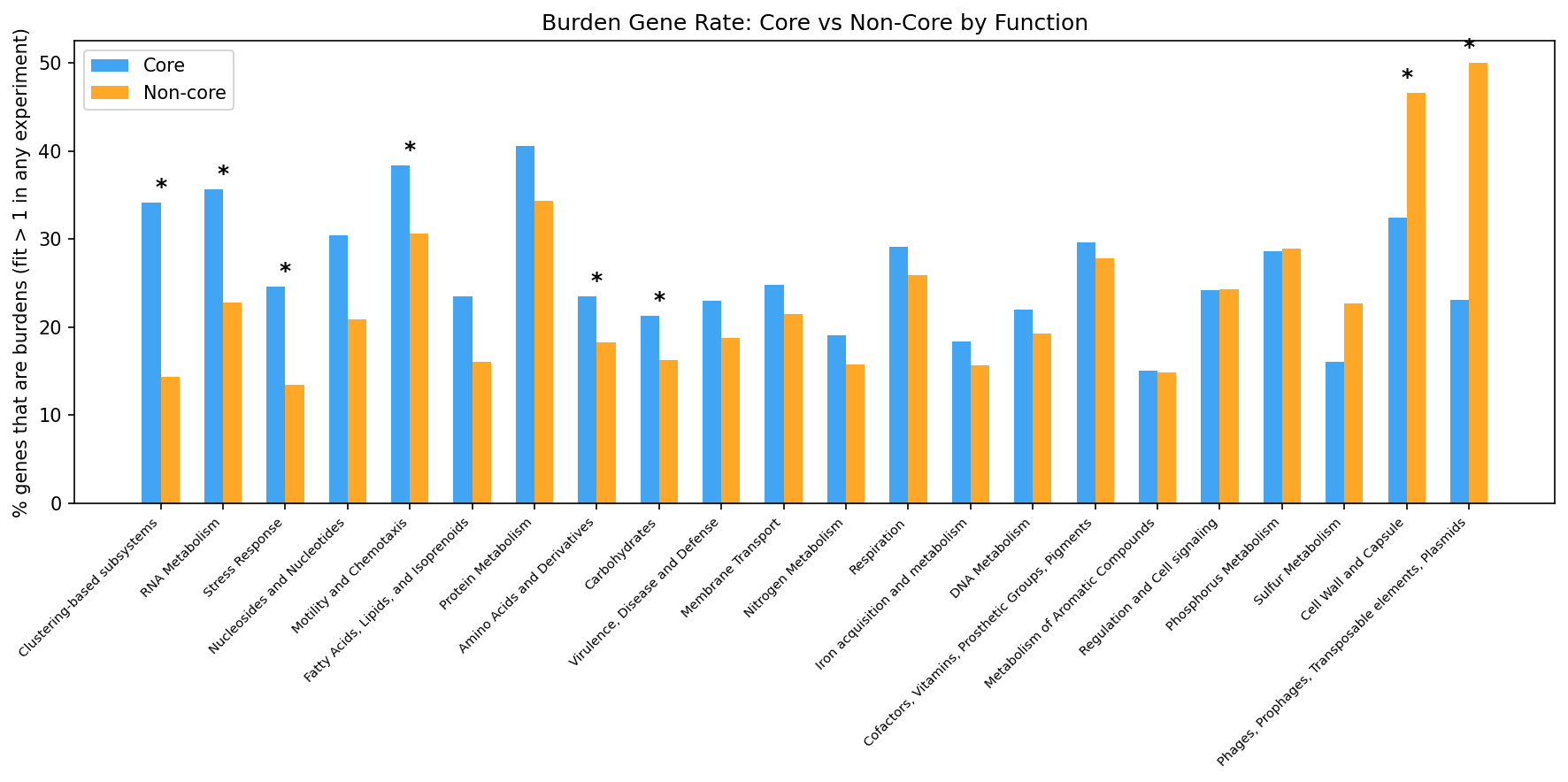
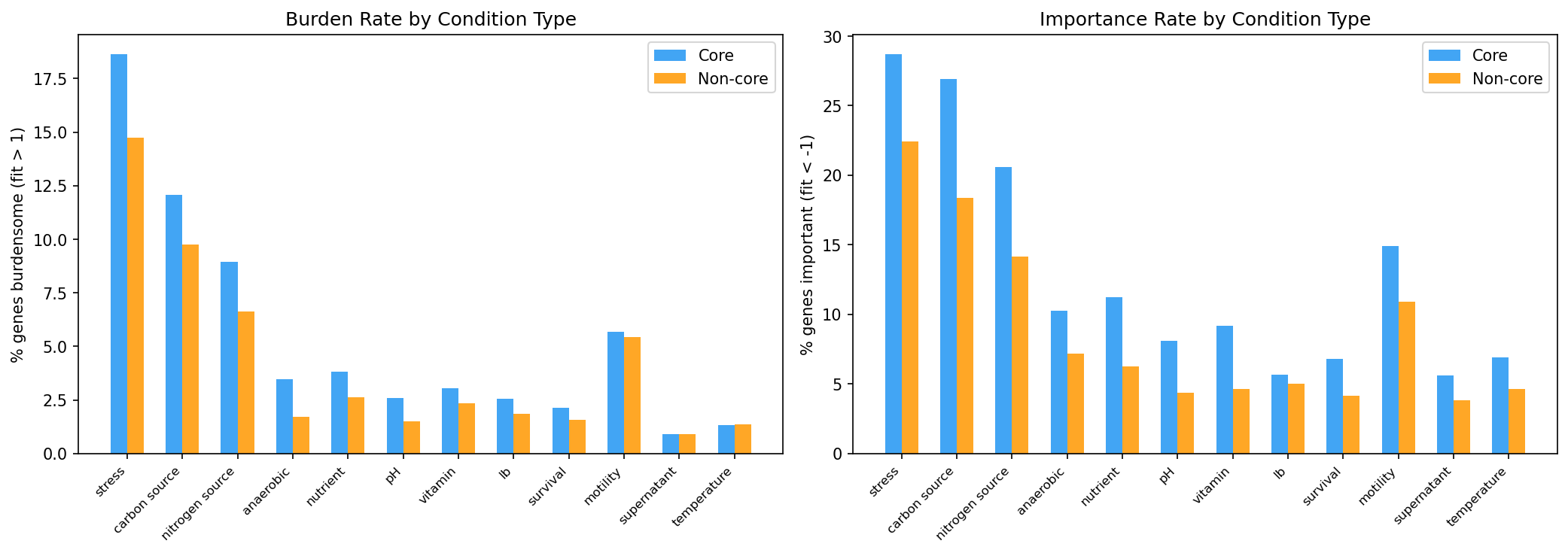
(Notebook: 01_burden_anatomy.ipynb)
Trade-Off Genes Are Enriched in Core
25,271 genes (17.8%) are true trade-off genes -- important (fit < -1) in some conditions, burdensome (fit > 1) in others. These are 1.29x more likely to be core (OR=1.29, p=1.2e-44). Core genes have more trade-offs because they participate in more pathways with condition-dependent costs and benefits.
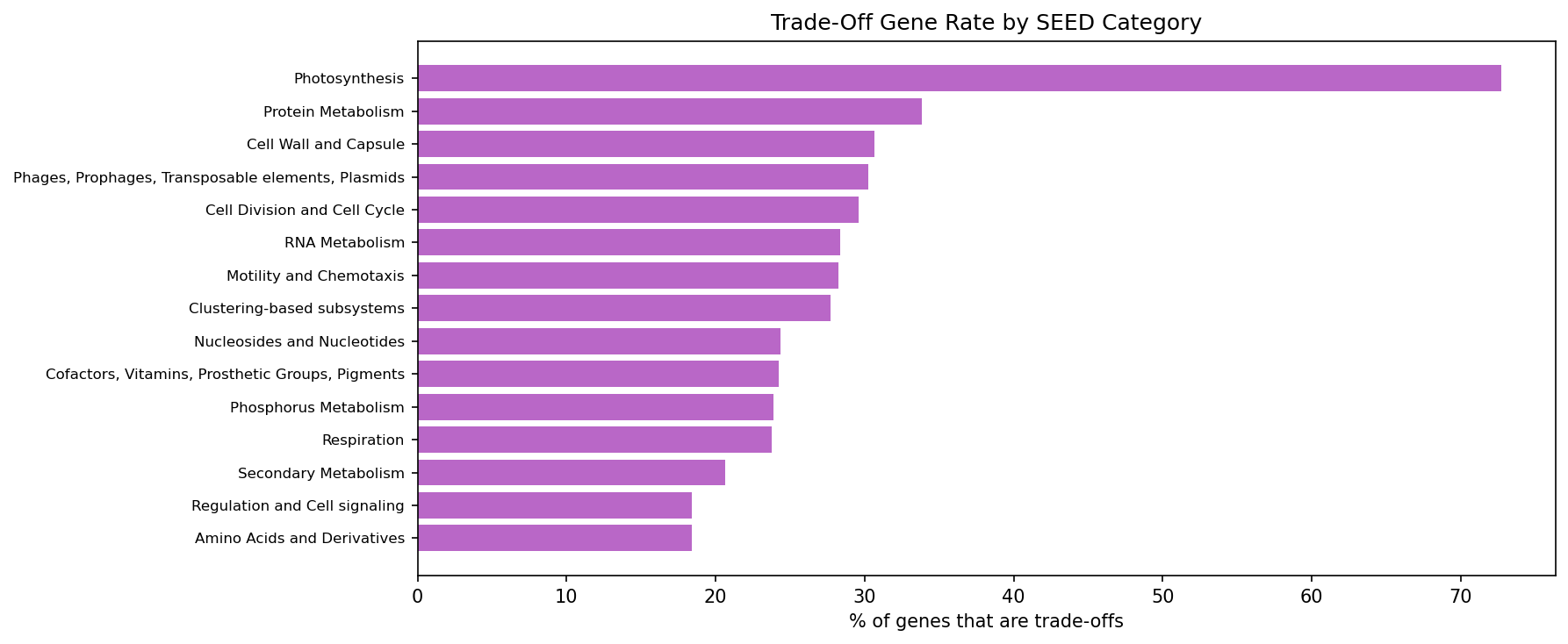
(Notebook: 01_burden_anatomy.ipynb)
The Selection Signature Matrix
| Conserved (core) | Dispensable (non-core) | |
|---|---|---|
| Costly (burden in lab) | 28,017 | 5,526 |
| Neutral (no burden) | 86,761 | 21,886 |
- Costly + Conserved (28,017 genes): Natural selection maintains them despite lab-measured cost -- they're essential in natural environments not captured by lab experiments
- Costly + Dispensable (5,526 genes): Candidates for ongoing gene loss -- burdensome AND not universally conserved
- Neutral + Conserved (86,761): Classic housekeeping genes
- Neutral + Dispensable (21,886): Niche-specific genes
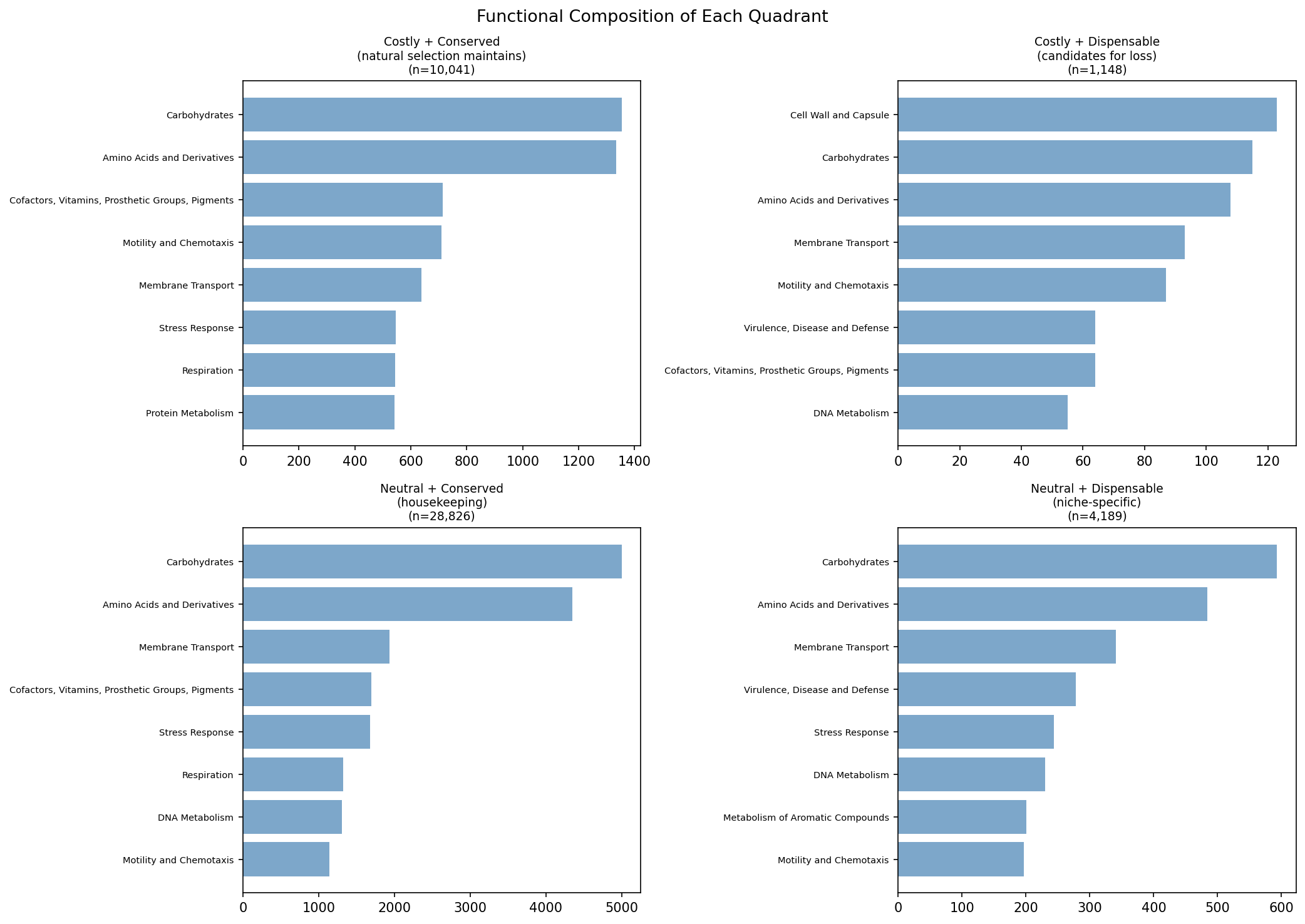
(Notebook: 01_burden_anatomy.ipynb)
Case Studies
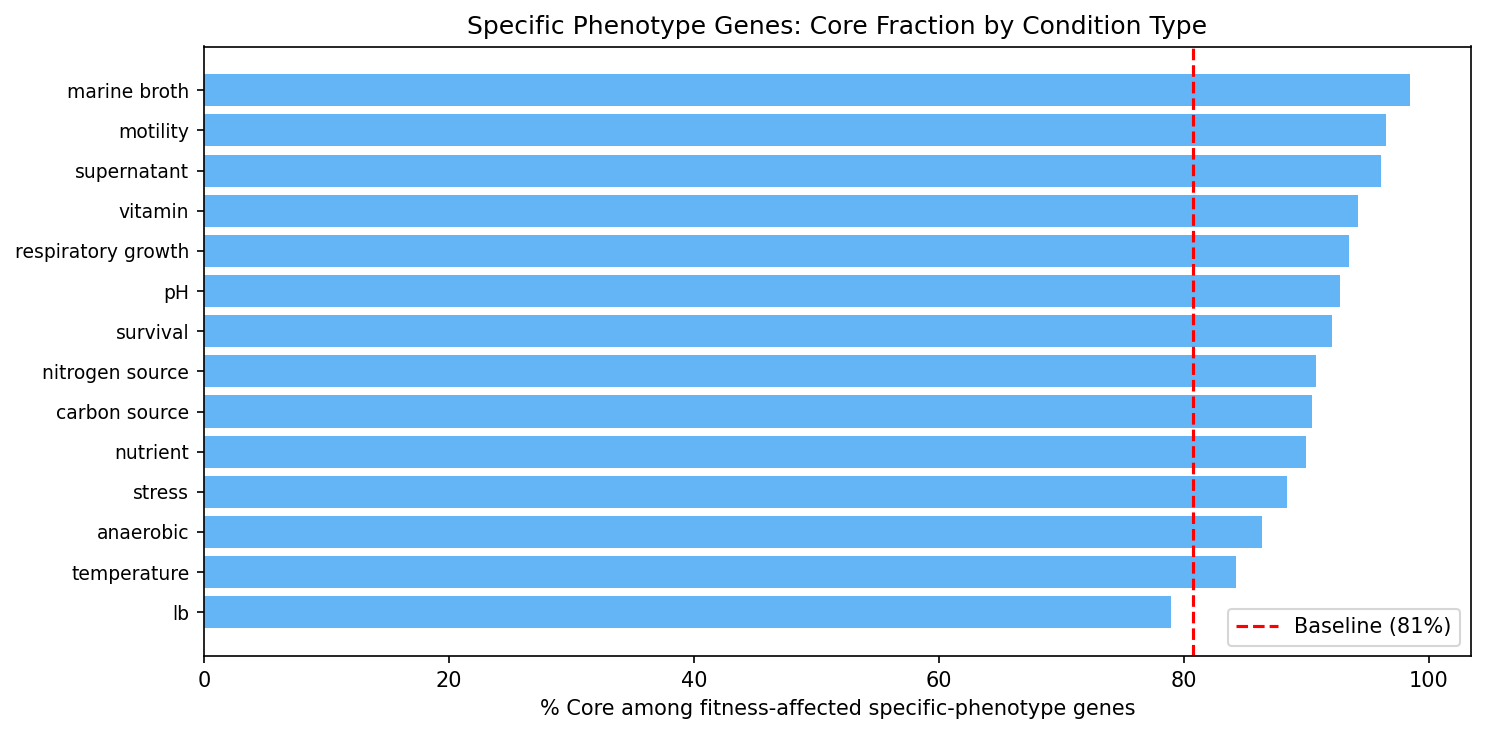
Genes with strong condition-specific effects are more likely core, reinforcing that the conserved genome is functionally active.
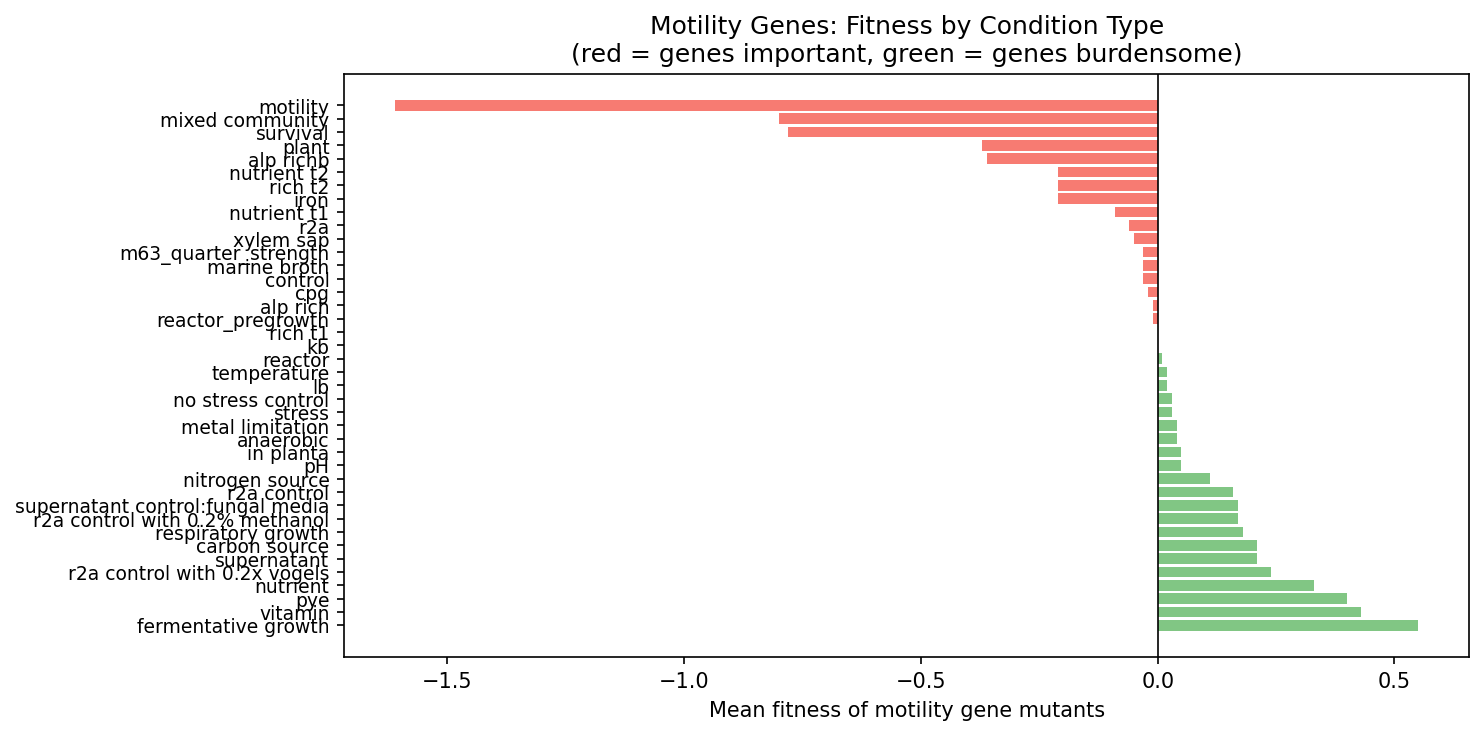
Motility genes exemplify the burden paradox: energetically expensive flagellar machinery is conserved because it is essential for chemotaxis in natural environments, despite being costly under lab conditions.
(Notebook: 01_burden_anatomy.ipynb)
Interpretation
The burden paradox resolves when we recognize that lab conditions are an impoverished proxy for nature. Core genes are more burdensome in the lab because they encode functions (motility, ribosomal components, RNA metabolism) that are energetically expensive but essential in natural environments. The 28,017 costly-but-conserved genes are the strongest evidence for purifying selection maintaining genes despite their metabolic cost. The function-specific pattern makes biological sense: flagella are expensive but essential for chemotaxis; ribosomal components are costly but required for rapid growth responses.
Literature Context
- Price et al. (2018) generated the Fitness Browser data used here, providing genome-wide mutant fitness measurements across diverse bacteria that reveal condition-dependent costs and benefits.
- Rosconi et al. (2022) demonstrated that gene essentiality is strain-dependent across S. pneumoniae pangenomes, supporting our finding that conservation reflects selection across diverse environments rather than universal essentiality.
- Koskiniemi et al. (2012) showed that dispensable genes impose fitness costs in Salmonella, consistent with our observation that core genes carry higher burden under lab conditions due to the metabolic expense of maintaining active pathways.
Limitations
- Lab conditions capture only a fraction of the environmental conditions bacteria face in nature
- "Burden" (fit > 1) may reflect trade-offs rather than true dispensability
- The 90% identity threshold for DIAMOND matching may miss rapidly evolving genes
- Condition types in the FB are biased toward what's experimentally convenient, not what's ecologically relevant
Data
Sources
| Dataset | Description | Source |
|---|---|---|
| Fitness Browser | RB-TnSeq mutant fitness data | Price et al. (2018) |
| KBase pangenome link table | Gene-to-cluster conservation mapping | conservation_vs_fitness/data/fb_pangenome_link.tsv |
| Fitness stats | Per-gene fitness summary | fitness_effects_conservation/data/fitness_stats.tsv |
| SEED annotations | Functional category assignments | conservation_vs_fitness/data/seed_annotations.tsv |
Generated Data
This project generates figures only; intermediate results are computed within the notebook from upstream data.
References
- Price MN et al. (2018). "Mutant phenotypes for thousands of bacterial genes of unknown function." Nature 557:503-509. PMID: 29769716
- Parks DH et al. (2022). "GTDB: an ongoing census of bacterial and archaeal diversity through a phylogenetically consistent, rank normalized and complete genome-based taxonomy." Nucleic Acids Res 50:D199-D207. PMID: 34520557
- Rosconi F et al. (2022). "A bacterial pan-genome makes gene essentiality strain-dependent and evolvable." Nat Microbiol 7:1580-1592.
- Koskiniemi S et al. (2012). "Selection-driven gene loss in bacteria." PLoS Genet 8:e1002787. PMID: 22761588
Discoveries
Trade-off genes are enriched in the core genome
February 202625,271 genes (17.8%) are true trade-offs — important (fit < -1) in some conditions, burdensome (fit > 1) in others. These are 1.29x more likely to be core (OR=1.29, p=1.2e-44). Core genes have more trade-offs because they participate in more pathways with condition-dependent costs and benefits. This
Read more →The burden paradox is function-specific, not universal
February 2026The core-burden paradox is driven by specific functional categories: RNA Metabolism (+12.9pp), Motility/Chemotaxis (+7.8pp), Protein Metabolism (+6.2pp) all show core genes as more burdensome. But Cell Wall reverses: non-core cell wall genes are MORE burdensome (-14.1pp). The paradox is not a unifor
Read more →Genes that are both burdensome in the lab AND core in the pangenome represent the strongest evidence for purifying selection in natural environments. They're costly to maintain, yet every strain keeps them — nature requires them in conditions not captured by the lab. By contrast, 5,526 genes are cos
Read more →Derived Data
This project builds on processed data from other projects.
Review
Summary
This is a well-motivated follow-up project that dissects a surprising finding from the upstream fitness_effects_conservation analysis: core genome genes are 1.3× more likely to show positive fitness effects (burdens) when deleted than accessory genes. The single notebook (01_burden_anatomy.ipynb) delivers a logically structured six-section analysis — function-specific burden rates, condition-type decomposition, trade-off gene enrichment, condition-specific paradox, a motility case study, and a selection-signature matrix — all supported by clear figures with saved outputs. The framing around lab conditions as an impoverished proxy for natural environments is compelling and scientifically grounded. The main areas for improvement are the absence of a data/ directory or requirements.txt, a potential performance issue in the motility analysis, and limited statistical testing in some sections.
Methodology
Research question: Clearly stated and testable — "Why are core genome genes MORE likely to show positive fitness effects when deleted?" The question builds directly on quantified results from the upstream project (OR=0.77, p=5.5e-48), making it a well-scoped follow-up.
Approach: The six-section decomposition is sound:
1. Breaking the paradox down by functional category identifies which gene classes drive the effect, rather than treating all genes uniformly.
2. The condition-type analysis tests whether specific experimental conditions create the burden signal.
3. Trade-off gene analysis (sick in some conditions, beneficial in others) provides the mechanistic explanation for why core genes appear burdensome.
4. The selection-signature 2×2 matrix (costly/neutral × conserved/dispensable) is an elegant synthesis that reframes the paradox as evidence for natural selection.
Data sources: Well-identified — the project reuses cached data from fitness_effects_conservation and conservation_vs_fitness, clearly documented in the README and notebook setup cell. The Dyella79 exclusion (known locus tag mismatch, per docs/pitfalls.md) is correctly applied (cell setup, line link = link[link['orgId'] != 'Dyella79']).
Reproducibility of reasoning: The notebook is self-contained and the logic chain from upstream findings to new analysis is traceable.
Code Quality
Notebook organization: Excellent. The notebook follows a clean setup → analysis × 6 → summary structure, with markdown headers separating each section. Each section produces both printed statistics and a saved figure.
Notebook outputs: All code cells have saved outputs (text and images confirmed across all 8 code cells). This is a significant strength — a reader can follow the entire analysis without re-execution.
Figures: All 6 figures listed in the README exist in the figures/ directory with reasonable file sizes (56–122 KB). Each corresponds to a major analysis section.
Statistical methods:
- Fisher's exact test for trade-off gene enrichment in core (OR=1.29, p=1.2e-44) is appropriate for the 2×2 contingency table.
- However, several other comparisons lack formal statistical tests. The burden rate differences by functional category (Section A) are presented as raw percentage-point differences without confidence intervals or significance tests. Given the large sample sizes, most differences would be significant, but a few categories with smaller gene counts (near the min_genes >= 50 threshold) could be noise.
Potential performance issue: In cell motility, the line by_cond.apply(lambda r: (r['orgId'], r['locusId']) in mot_loci, axis=1) iterates row-by-row over the full by_cond DataFrame using a Python lambda. Since fitness_stats_by_condition.tsv is 77 MB, this could be slow. A merge-based approach would be more efficient:
mot_keys = motility[['orgId','locusId']].drop_duplicates()
mot_cond = by_cond.merge(mot_keys, on=['orgId','locusId'], how='inner')
Pitfall awareness:
- The Dyella79 exclusion (documented in docs/pitfalls.md under "FB Locus Tag Mismatch") is correctly handled.
- Essential genes are not explicitly included in this analysis — the focus is on burden (positive fitness), so essential genes (no fitness data) are naturally excluded from burden classification. This is acceptable for the research question but could be noted.
- The project correctly uses cached local data rather than making Spark queries, avoiding API timeout issues.
Minor code issues:
- In cell burden-by-condition, the n_beneficial column from by_cond is used to define burden, but by_cond comes from fitness_stats_by_condition.tsv which has per-condition-type statistics. The column semantics should be verified — if n_beneficial counts experiments within a condition type where fit > 1, this is correct; if it's the overall count, the per-condition breakdown may be mixing levels.
Findings Assessment
Conclusions supported by data:
- The function-specific burden pattern (Section A) is well-supported, showing that the paradox is not uniform — Protein Metabolism, Motility, and RNA Metabolism drive the effect while Cell Wall reverses it.
- The trade-off gene enrichment (OR=1.29, p=1.2e-44, Section C) is robustly significant.
- The selection-signature matrix (Section F) provides a compelling synthesis: 28,017 genes that are both costly in the lab and conserved in pangenomes represent the strongest evidence for natural selection maintaining genes despite their lab-measured cost.
- The motility case study (Section E) is an effective illustration of condition-dependent trade-offs.
Limitations acknowledged: The README lists four concrete limitations including lab condition bias, the ambiguity of "burden," DIAMOND identity thresholds, and condition-type bias. These are appropriate and honest.
Completeness: The analysis is complete — no placeholder cells, no "TODO" markers, and the summary cell successfully prints all key results.
Visualization quality: Figures are properly titled and labeled. The paired bar charts (Sections A, B) effectively show core vs non-core comparisons. The 2×2 quadrant figure (Section F) is informative though somewhat dense.
Suggestions
-
Add statistical tests to Section A (burden by function): The burden rate differences by functional category are presented without significance tests. Add chi-squared tests or Fisher's exact tests for each category, with multiple-testing correction (e.g., Bonferroni or FDR). This would distinguish real functional differences from noise, especially for categories near the 50-gene minimum threshold.
-
Replace row-wise
applywith merge in motility analysis: Theby_cond.apply(lambda r: (r['orgId'], r['locusId']) in mot_loci, axis=1)pattern in cellmotilityis O(n) with a large constant. Replace with a merge join for better performance:
python mot_keys = motility[['orgId','locusId']].drop_duplicates() mot_cond = by_cond.merge(mot_keys, on=['orgId','locusId'], how='inner') -
Add a
requirements.txt: The project has no dependency specification. While the dependencies are standard (pandas, numpy, matplotlib, scipy), arequirements.txtwould complete the reproducibility story. The upstream project's README listsPython 3.10+, pandas, numpy, matplotlib, scipy— this project should do the same or provide a file. -
Document the empty
data/directory or remove it: Thedata/directory exists but is empty. Either populate it with intermediate outputs (e.g., the merged DataFramemas a TSV for downstream use) or remove it and update the project structure in the README, which currently does not list it. -
Clarify
n_beneficialcolumn semantics in condition analysis: In Section B,n_beneficialfromfitness_stats_by_condition.tsvis used to define burden within each condition type. Add a brief comment or markdown note confirming that this column counts within-condition-type experiments where the gene showed positive fitness, not the overall count across all experiments. -
Consider adding a combined summary figure: The project has 6 individual figures but no single summary visualization that ties together the main narrative (function-specific paradox → trade-offs → selection signature). A small multi-panel figure in the README or a final notebook cell could strengthen the presentation. (Nice-to-have)
-
Note essential gene treatment: The analysis focuses on burden (positive fitness when deleted), so essential genes (no fitness data) are naturally excluded from the burden definition. Adding a brief note in the notebook acknowledging that the 27,693 essential genes are not classified as burdens or non-burdens would prevent reader confusion. (Nice-to-have)
This review was generated by an AI system. It should be treated as advisory input, not a definitive assessment.
Visualizations

Burden By Condition

Burden By Function

Motility Case Study

Selection Signature Matrix

Specific Phenotype Conditions

Tradeoff Genes Conservation