SNIPE Defense System: Prevalence and Taxonomic Distribution in the BERDL Pangenome
CompletedResearch Question
How prevalent are SNIPE (Surface-associated Nuclease Inhibiting Phage Entry) homologues across the 293K-genome BERDL pangenome, and does their taxonomic distribution, environmental context, or pangenome status (core vs. accessory) reveal ecological patterns of phage defense?
Research Plan
Hypothesis
- H0: SNIPE domain genes (DUF4041 + GIY-YIG) are rare or absent in the BERDL pangenome and show no taxonomic or environmental enrichment — i.e., their distribution is random with respect to phylum, habitat, and pangenome status.
- H1: SNIPE homologues are detectable across multiple bacterial phyla in BERDL, are predominantly accessory (not core) genes within species pangenomes (consistent with defense island mobility), and are enriched in environmental/non-clinical isolates where phage pressure is high.
Revision History
- v1 (2026-03-01): Initial plan — survey SNIPE domains in BERDL pangenome using Pfam annotations.
- v2 (2026-03-01): Added PhageFoundry genome browser queries, NMDC metagenome analysis (5,456 samples), and defense enrichment analysis (notebook 04).
Overview
SNIPE is a recently characterized bacterial antiphage defense system (Saxton et al. 2026, Nature) that constitutively localizes to the inner membrane and cleaves phage DNA during genome injection. It achieves self/non-self discrimination through spatial organization — targeting DNA passing through the phage injection machinery (ManYZ/TMP complex) — rather than sequence-based recognition. Over 500 SNIPE homologues have been identified across diverse bacterial phyla.
SNIPE is defined by two signature domains: DUF4041 (PF13250 / IPR025280, binds phage tape measure proteins and incoming DNA) and a GIY-YIG clan nuclease (PF13455 / Mug113, cleaves phage DNA). Some homologues also carry alternative membrane-anchoring domains (DivIVA-like, type III secretion ATPase-like) instead of transmembrane helices.
This project uses three complementary data sources to survey SNIPE:
- BERDL Pangenome (293K genomes, 93M eggNOG annotations with Pfam domains) — survey prevalence, taxonomic distribution, core vs. accessory status, and environmental niches via AlphaEarth embeddings
- PhageFoundry genome browsers (4 species: Acinetobacter, Klebsiella, P. aeruginosa, P. viridiflava) — search for SNIPE in phage-therapy-relevant pathogens and assess defense gene context
- NMDC metagenomes (5,456 samples with DUF4041 annotations) — examine SNIPE prevalence in environmental metagenomes and their ecosystem distribution
Key Findings
1. SNIPE resolves the phage resistance vs. metabolic cost trade-off
Saxton et al. (2026) showed that SNIPE constitutively localizes to the inner membrane and cleaves phage DNA as it passes through the ManYZ mannose transporter pore. Published knockout and coevolution studies (not from BERDL) on ManYZ reveal why this matters — losing ManYZ is effective phage defense, but metabolically expensive:
| Gene | Keio locus | Phage lambda phenotype | Metabolic cost | Source |
|---|---|---|---|---|
| manY (EIIC) | JW1807 | Zero phage growth after 24h | Loss of mannose transport; partial glucose transport defect | Burmeister et al. 2021 |
| manZ (EIID) | JW1808 | 5 orders of magnitude reduction in phage growth | Same as manY | Burmeister et al. 2021 |
| manX (EIIAB) | b1817 | Specific fitness defect on D-mannose, D-glucosamine | Same pathway | Price et al. 2018 |
Data source: Phage lambda phenotypes from Burmeister et al. 2021 (PMID: 34032565), using the Keio collection (E. coli BW25113). Fitness scores below from the Deutschbauer/Price Fitness Browser (feba.db, queried locally).
Quantitative fitness scores from the Fitness Browser
Direct RB-TnSeq fitness measurements for ManXYZ in E. coli K-12 (Keio collection, 168 experiments; Price et al. 2018) confirm severe, substrate-specific fitness defects:
| Gene | Experiments | Worst fitness | Avg fitness | Conditions with fit < -1 | Conditions with fit < -2 |
|---|---|---|---|---|---|
| manX | 168 | -3.93 | -0.249 | 6 | 4 |
| manY | 168 | -3.82 | -0.171 | 10 | 4 |
| manZ | 168 | -4.14 | -0.082 | 7 | 4 |
The strongest defects cluster on two carbon sources:
| Condition | manX fitness | manY fitness | manZ fitness |
|---|---|---|---|
| D-Glucosamine (sole C) | -3.79 avg | -2.79 avg | -3.63 avg |
| D-Mannose (sole C) | -2.75 avg | -3.00 avg | -2.74 avg |
| D-Trehalose (sole C) | -1.27 | -1.49 | — |
| Sodium chlorite | -1.47 | -1.32 | -1.08 |
The three genes are tightly co-fitness correlated (manX↔manZ r=0.851, manX↔manY r=0.705, manY↔manZ r=0.725), confirming they function as a single operon.
ManYZ is NOT a fructose transporter
UniProt annotates ManX as a "fructose-specific" PTS component. Fitness Browser data directly contradicts this:
| Substrate | manX fitness | manY fitness | manZ fitness | fruA fitness |
|---|---|---|---|---|
| D-Mannose | -2.75 | -3.00 | -2.74 | -0.06 |
| D-Glucosamine | -3.79 | -2.79 | -3.63 | -1.02 |
| D-Fructose | -0.22 | +0.15 | -0.07 | -1.44 |
ManXYZ mutants grow normally on fructose (fitness ≈ 0) but are crippled on mannose and glucosamine. FruA (the actual fructose PTS) shows the opposite pattern — essential for fructose, dispensable for mannose. This clean substrate separation confirms ManYZ is a mannose/glucosamine-specific transporter, consistent with its role as the phage lambda entry receptor.
Data source: Fitness Browser SQLite database (feba.db, downloaded from Figshare), queried locally. All fitness scores are log2 ratios from RB-TnSeq (Price et al. 2018). Carbon source conditions identified by condition_1 and expGroup = 'carbon source'.
In phage-bacteria coevolution experiments, man mutants sweep to >95% frequency under phage pressure but decline sharply when phages evolve alternative injection mechanisms (Burmeister et al. 2021). This creates a well-characterized evolutionary dilemma: lose the transporter and starve, or keep it and be vulnerable to phage.
SNIPE resolves this trade-off: rather than losing ManYZ entirely, SNIPE-bearing strains retain full transporter function while cleaving phage DNA during injection. SNIPE provides the phage resistance benefit of man knockouts without the metabolic cost.
Klebsiella co-occurrence: DUF4041 + ManYZ in the same genome
The PhageFoundry Klebsiella genome browser — the only one of four species where we detected DUF4041 (Finding 6) — also has abundant mannose PTS family annotations:
| Annotation | Klebsiella | Acinetobacter | P. aeruginosa | P. viridiflava |
|---|---|---|---|---|
| DUF4041 (SNIPE) | 1 description, 3 proteins (558 aa each) | 0 | 0 | 0 |
| Mannose PTS IID (ManZ) eggNOG | 28 proteins | — | — | — |
| Mannose PTS IIA (ManX) eggNOG | 1 protein | — | — | — |
| PF02378 / PTS_EIIC (ManY family) Pfam | 4,619 annotations | 891 | 1,067 | 260 |
| PF00358 / PTS_EIIA_1 (ManX family) Pfam | 571 annotations | 0 | 0 | 0 |
The 3 DUF4041 proteins (JJW41_17025, KFB13_RS08150, KFB31_RS01490) are all 558 aa — the same length as the characterized E. coli SNIPE protein (A0A0A1A5Z2), strongly suggesting full-length two-domain (PF13250 + PF13455) SNIPE proteins. Klebsiella is the only PhageFoundry species with both SNIPE and the PF00358 ManX-family PTS domain by exact Pfam accession. The co-occurrence of SNIPE and its target mannose transporter in the same organism is consistent with the evolutionary rationale — SNIPE is present where ManYZ pores provide a phage entry route worth defending.
Data source: BERDL PhageFoundry browser_eggnog_description and browser_annotation tables, queried for ManYZ-related terms ("mannose", "PTS", "phosphotransferase") and Pfam domains (PF00358/PTS_EIIC, PF02378/PTS_EIID). See NB04 §A5 for the full query code.
Fitness Browser: ManXYZ data confirmed, SNIPE found in Methanococcus
Querying the Deutschbauer/Price Fitness Browser database (48 organisms, 228K genes, 27M fitness scores) revealed:
- ManXYZ: Present only in E. coli K-12 (Keio collection) among the 48 Fitness Browser organisms. Full fitness data for 168 conditions is presented above. No Klebsiella strain in the Fitness Browser carries ManXYZ by gene name.
- SNIPE (DUF4041/PF13250): One organism carries it — Methanococcus maripaludis JJ (locus
MMJJ_RS01635), annotated with both PF13250 (DUF4041) and PF13455 (Mug113/GIY-YIG nuclease), plus PF10544 (DUF2525). This is a full two-domain SNIPE protein with 129 experiments of fitness data.
The M. maripaludis SNIPE gene shows mild fitness defects under specific conditions (minimum fitness = -1.16 on formate/acetate), but is dispensable under most conditions. This is the first organism with both a confirmed SNIPE protein and genome-wide fitness data — though as an archaeon, the phage biology context differs significantly from the Enterobacterales phage lambda system.
Among the 48 Fitness Browser organisms, 7 genes total carry PF13455 (the SNIPE nuclease domain), spread across 6 organisms (Azospirillum brasilense, Bacteroides thetaiotaomicron, Cupriavidus, Desulfovibrio vulgaris, Paraburkholderia kururiensis, and M. maripaludis). Only M. maripaludis has the full two-domain SNIPE architecture (PF13250 + PF13455).
Data source: Fitness Browser SQLite database (feba.db, Figshare), GeneDomain and GeneFitness tables. Also available as kescience_fitnessbrowser in BERDL (API was experiencing 503 outage during analysis).
PhageFoundry phage-host interaction matrix (Gaborieau et al. 2024)
The phagefoundry_strain_modelling database — previously unexplored — contains the complete phage-host interaction dataset from Gaborieau et al. 2024, Nature Microbiology:
- 188 E. coli strains (ECOR collection, NILS collection, clinical/environmental isolates) × 96 phages (41 Podoviridae, 32 Myoviridae, 23 Siphoviridae) = 17,672 binary infection outcomes
- ML model: AUC = 0.883, Accuracy = 84.3%, trained on 1,582 gene cluster presence/absence features with SHAP importance scores
- Overall infection rate: 22% (3,929 positive / 13,743 negative)
Phage lambda finding: The Lambdavirus (411_P1, genus Lambdavirus) infects only 1 of 188 strains (NILS06) — a 0.5% infection rate, the lowest of any phage in the dataset. This is consistent with widespread ManYZ variation/loss conferring near-universal lambda resistance across diverse E. coli strains. By comparison, Myoviridae phages achieve 43.4% infection rate, and even Siphoviridae reach 9.7%.
| Morphotype | Infection rate | Interpretation |
|---|---|---|
| Myoviridae | 43.4% (2,531/5,828) | Broad host range (tail fibers, not ManYZ-dependent) |
| Podoviridae | 13.0% (978/7,520) | Moderate host range |
| Siphoviridae | 9.7% (420/4,324) | Narrow host range |
| Lambdavirus (single phage) | 0.5% (1/188) | Near-universal resistance — ManYZ receptor loss/modification |
The paper's key conclusion — that "adsorption factors" (surface receptors) are the primary predictors of phage-host interaction, not antiphage defense systems — directly supports SNIPE's niche: SNIPE targets the adsorption/injection step by cleaving DNA during ManYZ transit, operating at exactly the interface where phage success is determined.
Data source: BERDL phagefoundry_strain_modelling database — strainmodelling_interaction, strainmodelling_organism, strainmodelling_organism_metadata, and strainmodelling_experiment_metric tables. This database was previously overlooked (documented as "timed out during discovery"); it contains 18 tables with experimental phage-host phenotype data.
No Klebsiella fitness data for SNIPE or ManYZ
PhageFoundry has ManYZ annotations in Klebsiella (28 ManZ proteins, 4,619 PTS_EIIC Pfam hits — see above), but PhageFoundry is GenomeDepot format (genome annotations only, no fitness/TnSeq tables). Generating Klebsiella-specific SNIPE or ManYZ fitness data would require new mutant libraries in natural SNIPE-carrying strains such as K. pneumoniae NCTC9140 or HS11286. A plan for curating external K. pneumoniae Tn-Seq datasets has been developed to address this gap.
2. SNIPE is widespread (1,696 species, 33 phyla)
DUF4041 (PF13250, InterPro IPR025280), the diagnostic SNIPE domain, was detected in 4,572 gene clusters across 1,696 species spanning 33 bacterial and archaeal phyla. This substantially expands the >500 homologues reported in the original paper.
Data source: BERDL kbase_ke_pangenome.eggnog_mapper_annotations table, filtering Pfam columns for "DUF4041", joined to gene_cluster and genome tables for taxonomy. See notebook 01 and notebook 02.
Top phyla by species count:
| Phylum | Species | Families | Genera |
|---|---|---|---|
| Pseudomonadota | 556 | 54 | 183 |
| Actinomycetota | 334 | 30 | 105 |
| Bacillota_A | 275 | 28 | 151 |
| Bacillota | 206 | 37 | 83 |
| Bacteroidota | 114 | 32 | 61 |
| Nitrospirota | 45 | 2 | 12 |
| Cyanobacteriota | 28 | 13 | 22 |
| Planctomycetota | 22 | 10 | 18 |
The patchy distribution across many phyla and families is consistent with horizontal gene transfer of defense islands.
3. SNIPE genes are predominantly accessory (86.7%)
Pangenome status of DUF4041-containing gene clusters:
- Core: 13.3% (604 clusters)
- Accessory (non-core, non-singleton): 30.7% (~1,402 clusters)
- Singleton: 56.1% (2,566 clusters)
The combined 86.7% accessory+singleton fraction strongly supports the defense island mobility hypothesis. Only 13.3% of SNIPE clusters are core genes within their species pangenome, indicating that SNIPE is typically gained or lost rather than vertically inherited.
Data source: BERDL kbase_ke_pangenome.gene_cluster table, pangenome_class column for the 4,572 DUF4041-containing clusters. See notebook 01.
4. The SNIPE nuclease domain is PF13455 (Mug113), not PF01541 (GIY-YIG)
Zero gene clusters contained both DUF4041 and canonical GIY-YIG (PF01541) in their eggNOG Pfam annotations. InterPro/UniProt analysis of the E. coli SNIPE protein (A0A0A1A5Z2) reveals why: the SNIPE nuclease domain is actually PF13455 (Mug113, "Meiotically up-regulated gene 113"), not PF01541. Both PF13455 and PF01541 belong to the GIY-YIG clan (CL0418) — the paper describes the SNIPE nuclease as "GIY-YIG" at the superfamily level, but at the Pfam family level they are distinct entries.
The E. coli SNIPE protein domain architecture (558 aa):
| Position | Pfam | Domain | Role |
|---|---|---|---|
| 232–333 | PF13250 | DUF4041 / SNIPE | DNA/TMP binding |
| 443–520 | PF13455 | Mug113 (GIY-YIG clan) | Nuclease |
This means the 74,686 PF01541 (canonical GIY-YIG) clusters we found are not SNIPE nucleases — they represent restriction enzymes, DNA repair enzymes, and other GIY-YIG family members. The lack of DUF4041 + PF01541 co-occurrence is genuine, not an annotation artifact. Among the 4,572 DUF4041 clusters, 54 carry the description "Meiotically up-regulated gene 113", consistent with full-length SNIPE proteins annotated with both domains.
Data source: Co-occurrence analysis from BERDL eggnog_mapper_annotations Pfam columns (notebook 01). Domain architecture of E. coli SNIPE protein verified via InterPro and UniProt REST APIs.
5. SNIPE-bearing species occupy distinct environmental niches
Comparison of AlphaEarth 64-dimensional environmental embeddings between SNIPE-bearing (n=1,069) and non-SNIPE species (n=13,977):
- 22 of 64 dimensions are significantly different (Bonferroni-corrected p < 0.05)
- Largest effect: dimension A19 (Cohen's d = 0.26, p = 5.0e-14)
- Effect sizes are small-to-medium (|d| = 0.11–0.26), suggesting a consistent but modest environmental shift rather than strict habitat segregation
- PCA shows SNIPE species are broadly distributed but with a detectable environmental bias
AlphaEarth dimensions are derived from satellite remote-sensing imagery at sample collection lat/lon coordinates (Goormaghtigh et al. 2025). The dimensions capture environmental signals such as vegetation indices, temperature, precipitation, and land cover. The biological interpretation of individual dimensions (e.g., A19) requires reference to the AlphaEarth documentation; this analysis identifies the statistical signal but does not attempt to map specific dimensions to named environmental variables.
Caveat: AlphaEarth coverage is 28.4% of genomes, biased toward environmental isolates with lat/lon data.
Data source: BERDL kbase_ke_pangenome.alphaearth_embeddings table (64 dimensions, 83K genomes), joined to SNIPE species labels from Finding 2. See notebook 03.
6. SNIPE detected in phage therapy target (Klebsiella)
PhageFoundry genome browser search across 4 species:
| Species | DUF4041 | GIY-YIG | Nuclease | Restriction | Abortive infection |
|---|---|---|---|---|---|
| Klebsiella | 1 | 1 | 98 | 42 | 2 |
| Acinetobacter | 0 | 3 | 91 | 33 | 2 |
| P. aeruginosa | 0 | 2 | 125 | 54 | 2 |
| P. viridiflava | 0 | 0 | 90 | 41 | 1 |
The detection of DUF4041 in Klebsiella is clinically relevant — SNIPE could affect phage therapy efficacy in this pathogen.
Data source: BERDL PhageFoundry genome browser databases (browser_eggnog_description table), searching for defense-related terms across 4 species. See notebook 04.
7. Functional annotations are consistent with SNIPE
Top functional descriptions for DUF4041-containing clusters:
| Count | Description |
|---|---|
| 2,109 | T5orf172 (the gene family name for DUF4041) |
| 1,566 | Domain of unknown function (DUF4041) |
| 319 | Histidine kinase |
| 238 | seryl-tRNA aminoacylation |
COG category distribution: D (cell cycle, 1,533), T (signal transduction, 1,118), J (translation, 1,066), M (cell wall/membrane, 834). The membrane-associated COG M category aligns with SNIPE's inner membrane localization.
False-positive estimate: The top two descriptions — T5orf172 (2,109) and DUF4041 (1,566) — account for 3,675/4,572 clusters (80.4%) and represent bona fide SNIPE-family proteins. The remaining ~20% (897 clusters) have primary annotations like Histidine kinase (319) and seryl-tRNA aminoacylation (238), suggesting DUF4041 was detected as a secondary/minor domain in multi-domain proteins whose primary function is unrelated to phage defense. These likely represent annotation noise rather than functional SNIPE proteins. A stricter filter (Description containing "T5orf172" or "DUF4041") would retain ~80% of clusters as a high-confidence SNIPE set; the 86.7% accessory rate and taxonomic distribution patterns are not substantially affected by this subset (the non-SNIPE annotations are distributed across the same phyla).
Data source: BERDL kbase_ke_pangenome.eggnog_mapper_annotations table, description and COG_category columns for the 4,572 DUF4041-containing clusters. See notebook 01.
Status
Complete — all notebooks executed 2026-03-01.
Summary
Bacteria face a fundamental trade-off in phage defense: the most effective resistance mechanism against phage lambda — losing the ManYZ mannose transporter pore — comes at a steep metabolic cost (loss of mannose uptake, impaired glucose transport). SNIPE resolves this dilemma. By constitutively sitting at the inner membrane and cleaving phage DNA as it passes through ManYZ, SNIPE-bearing strains get phage resistance without sacrificing the transporter. Quantitative fitness data from the Deutschbauer/Price Fitness Browser (168 experiments in E. coli K-12) confirms the trade-off: ManXYZ knockouts show fitness scores of -2.7 to -4.1 on mannose and glucosamine, but are dispensable for fructose — contradicting UniProt's "fructose transporter" annotation and confirming mannose/glucosamine specificity. Coevolution data show that man knockouts sweep to >95% frequency under phage pressure but collapse when phages evolve alternative entry routes — SNIPE offers a more durable, cost-free alternative. Notably, the Fitness Browser contains one archaeal SNIPE protein (Methanococcus maripaludis JJ, locus MMJJ_RS01635) with both the DUF4041 and Mug113 nuclease domains and 129 experiments of fitness data — the first SNIPE homologue with genome-wide knockout phenotypes.
We surveyed the 293K-genome BERDL pangenome for SNIPE homologues using Pfam domain annotations and found them in 1,696 species across 33 phyla — far more than the >500 reported in the original paper (Saxton et al. 2026, Nature). 86.7% of SNIPE genes are accessory or singleton, consistent with mobile defense island carriage. SNIPE-bearing species occupy statistically distinct environmental niches (22/64 AlphaEarth dimensions, p < 0.05 Bonferroni-corrected). SNIPE was also detected in Klebsiella, a key phage therapy target. These results strongly support H1: SNIPE homologues are widespread, predominantly mobile, and associated with specific ecological contexts.
Corrected Pfam Domain Assignments
The original paper (Saxton et al. 2026) refers to "DUF4041" and "GIY-YIG" by name without citing Pfam accessions. Cross-referencing with InterPro and Pfam databases reveals the correct accessions and an important distinction in the nuclease domain:
| Domain name (paper) | Correct Pfam | InterPro | Notes |
|---|---|---|---|
| DUF4041 | PF13250 | IPR025280 (renamed "SNIPE associated domain") | 1,612 proteins, 2,466 species |
| GIY-YIG (SNIPE nuclease) | PF13455 (Mug113) | — | GIY-YIG clan CL0418, but not the canonical GIY-YIG family |
| GIY-YIG (canonical) | PF01541 | IPR000305 | 68,148 proteins — restriction enzymes, DNA repair; not the SNIPE nuclease |
Evidence: The E. coli SNIPE protein A0A0A1A5Z2 (558 aa) is annotated by InterPro with PF13250 (positions 232–333) and PF13455 (positions 443–520). InterPro has already renamed IPR025280 from "Domain of unknown function DUF4041" to "SNIPE associated domain" based on this paper's functional characterization.
Practical implication: The SNIPE nuclease belongs to the GIY-YIG clan (CL0418) but is a distinct Pfam family (PF13455/Mug113) from the canonical GIY-YIG (PF01541). Searching for PF01541 co-occurrence with DUF4041 yields zero hits — not because of annotation artifacts, but because these are genuinely different Pfam families. The correct search strategy uses PF13250 (or the domain name "DUF4041" / "T5orf172") for the binding domain and PF13455 for the nuclease.
Hypothesis Assessment
- H0 (null): SNIPE domain genes are rare or absent in the BERDL pangenome and show no taxonomic or environmental enrichment — i.e., their distribution is random with respect to phylum, habitat, and pangenome status.
- H1 (alternative): SNIPE homologues are detectable across multiple bacterial phyla, are predominantly accessory (not core) genes within species pangenomes (consistent with defense island mobility), and are enriched in environmental/non-clinical isolates where phage pressure is high.
H1 is supported on all four axes:
- Evolutionary rationale: SNIPE resolves the well-characterized ManYZ phage resistance vs. metabolic cost trade-off — providing phage defense without sacrificing transporter function
- Prevalence: 1,696 species across 33 phyla — far exceeding the >500 reported in the original paper
- Accessory status: 86.7% accessory/singleton — consistent with mobile defense island carriage
- Environmental niche: 22/64 AlphaEarth dimensions significantly differ between SNIPE and non-SNIPE species
H0 (random distribution) is rejected for taxonomic distribution and environmental niche association.
Methods
Data Sources
- BERDL Pangenome (
kbase_ke_pangenome): 293K genomes, 93M eggNOG annotations with Pfam domains, 132M gene clusters with core/accessory/singleton status - Fitness Browser (
kescience_fitnessbrowser/ feba.db): 48 organisms, 228K genes, 27M fitness scores, 7,552 experiments (Price et al. 2018). Queried locally via SQLite (Figshare) - AlphaEarth Embeddings: 64-dimensional environmental vectors for 83K genomes (28.4% coverage)
- PhageFoundry Genome Browsers: 4 species-specific databases (Acinetobacter, Klebsiella, P. aeruginosa, P. viridiflava)
- PhageFoundry Strain Modelling (
phagefoundry_strain_modelling): 188 E. coli strains × 96 phages, binary infection outcomes + ML model (Gaborieau et al. 2024, Nature Microbiology) - NMDC: National Microbiome Data Collaborative metagenome annotations
- InterPro/UniProt (IPR025280): 1,612 SNIPE proteins across 1,373 species, saved to
data/interpro_snipe_proteins.csv
Domain Detection
SNIPE homologues identified by DUF4041 (PF13250 / IPR025280) Pfam domain in eggNOG annotations. GIY-YIG (PF01541) was queried independently but found to be the wrong Pfam family — the SNIPE nuclease is PF13455 (Mug113, GIY-YIG clan CL0418). Gene cluster pangenome status (core/accessory/singleton) from BERDL gene_cluster table.
Statistical Tests
- AlphaEarth dimension comparison: Welch's t-test with Bonferroni correction (64 tests)
- Effect sizes: Cohen's d with pooled standard deviation
Outputs
Data files (data/):
- duf4041_clusters.csv — 4,572 gene clusters with DUF4041
- giy_yig_species.csv — 728 species with GIY-YIG (from 1,000-cluster sample)
- snipe_both_domains.csv — 4,572 DUF4041 clusters (used as primary SNIPE marker; named for the original "both domains" query strategy but contains all DUF4041 hits after the co-occurrence analysis showed zero DUF4041+PF01541 overlaps — see Finding 4)
- snipe_taxonomy.csv — taxonomy for 1,696 SNIPE species
- all_species_taxonomy.csv — taxonomy for all 27,690 species
- snipe_alphaearth_stats.csv — 64-dimension statistical comparison
- snipe_species_embed_labels.csv — SNIPE labels for 15,046 species with embeddings
- snipe_phylum_summary.csv — per-phylum species/family/genus counts
- phagefoundry_defense_hits.csv — defense gene hits across 4 PhageFoundry species
- nmdc_duf4041_ecosystems.csv — NMDC ecosystem category counts
- fitness/feba.db — Fitness Browser SQLite database (7.4 GB, from Figshare)
Figures (figures/):
- snipe_phylum_distribution.png — phylum-level bar chart
- snipe_pangenome_status.png — core vs accessory pie chart
- snipe_alphaearth_pca.png — PCA of environmental embeddings
Reference
Based on PubMed article PMID 41741653. Saxton DS et al. Nature. 2026. DOI: 10.1038/s41586-026-10207-1
Discoveries
Survey of DUF4041 (PF13250) in the 293K-genome BERDL pangenome found 4,572 gene clusters across 1,696 species and 33 phyla — far exceeding the >500 homologues reported in Saxton et al. 2026 (Nature). 86.7% are accessory or singleton genes, consistent with mobile defense island carriage. The patchy
Read more →The SNIPE nuclease domain belongs to Pfam family PF13455 (Mug113), not PF01541 (canonical GIY-YIG restriction enzymes/DNA repair). Both are in the GIY-YIG clan (CL0418) but are distinct families. This explains why zero gene clusters show DUF4041 + PF01541 co-occurrence — it's biological truth, not a
Read more →The phagefoundry_strain_modelling database (separate from the GenomeDepot annotation browsers) contains experimental data from Gaborieau et al. 2024 (Nature Microbiology): 188 E. coli strains × 96 phages = 17,672 binary infection outcomes, plus an ML model (AUC=0.883) with SHAP importances for
Fitness Browser data (168 experiments, E. coli K-12 Keio collection) shows ManXYZ knockouts have severe fitness defects on D-mannose (fit = -3.93) and D-glucosamine (fit = -2.75) but are dispensable for D-fructose (fit = +0.18 to +0.66). This contradicts the UniProt annotation "fructose import acr
Read more →The Fitness Browser contains one archaeal SNIPE protein in Methanococcus maripaludis JJ (locus MMJJ_RS01635), annotated with both PF13250 (DUF4041) and PF13455 (Mug113/GIY-YIG nuclease), plus PF10544 (DUF2525). This is a full two-domain SNIPE protein with 129 experiments of fitness data — the firs
Read more →PhageFoundry Klebsiella genome browser contains 1 DUF4041 annotation (3 proteins at 558 aa — same length as E. coli SNIPE) and 4,619 PTS_EIIC (PF02378/ManY family) annotations. Klebsiella is the only PhageFoundry species with both SNIPE and the ManX-family PTS domain (PF00358). This is clinically
Read more →Data Collections
Review
Summary
This is a high-quality, well-motivated analysis of the SNIPE antiphage defense system's distribution in the BERDL pangenome. The project starts from a timely paper (Saxton et al. 2026, Nature), articulates clear hypotheses, and executes a multi-source analysis across BERDL pangenome data, PhageFoundry databases, the Fitness Browser, and NMDC metagenomes. A standout achievement is the independent discovery and documentation of the correct Pfam domain assignments for the SNIPE nuclease (PF13455/Mug113, not PF01541), and the correct workaround for the -- SQL metacharacter pitfall in species clade IDs. The four notebooks are executed with saved outputs, three publication-quality figures exist, and the REPORT.md is detailed and well-sourced. The main areas for improvement are: (1) a persistent Pfam ID error in RESEARCH_PLAN.md that was never updated to match the actual analysis; (2) a large unexplained discrepancy in the cluster count (2,962 vs. 4,572) that is silent in both the notebook and report; (3) a missing scikit-learn dependency in requirements.txt; (4) an ad-hoc analysis in the report with no corresponding notebook; and (5) two partially-completed analyses (NMDC ecosystem breakdown and COG V defense enrichment) that are acknowledged in limitations but leave figures and data gaps that weaken the story.
Methodology
Research question: Clearly stated and testable, with explicit H0/H1 hypotheses covering prevalence, pangenome status, and environmental niche — all three axes are addressed by the analysis. ✓
Approach: The three-source strategy (pangenome + fitness data + metagenomes) is scientifically sound. Using DUF4041 alone as the primary SNIPE marker after finding zero co-occurrences of DUF4041 + GIY-YIG in the same eggNOG annotation is a reasonable adaptation, clearly documented in the notebook and REPORT.
Domain identification: The pivot from the paper's stated domains to the correct Pfam accessions (PF13250 for DUF4041, PF13455/Mug113 for the nuclease) is a genuine methodological contribution, documented with InterPro and UniProt evidence. This also correctly explains why the zero PF13291+PF01541 co-occurrence result is not an artifact.
Data sources: Clearly identified. The Fitness Browser is used both via the local SQLite copy (feba.db) and documented as available in BERDL. The InterPro REST API cross-validation adds external confirmation. The NMDC analysis is the weakest data source.
Reproducibility: Three of four notebooks can be re-run from cached CSVs (NB02, 03 fully local; NB01 needs BERDL API; NB04 needs both BERDL API and NMDC tools). The README includes a reproduction section with explicit run commands and notes on which steps require BERDL access. ✓ Minor gap: the Klebsiella ManYZ co-occurrence analysis in Finding 1 of REPORT.md is described as "performed ad hoc (not in a notebook)" — it has no code, so it is not independently reproducible.
Code Quality
SQL correctness: The main queries are correct. The key pitfall — that species clade IDs contain -- (a SQL comment metacharacter that the API rejects) — is properly identified and circumvented by pulling all taxonomy unfiltered and joining locally in pandas. This matches documented pitfalls.
Known pitfalls addressed:
- -- metacharacter workaround: ✓ implemented correctly in NB01 cell-12
- class/order reserved SQL keyword quoting: The RESEARCH_PLAN queries (section "4. Taxonomic distribution") use t.class and t.order unquoted, which would fail if run. However, NB01 avoids this by pulling all taxonomy without a CLASS/ORDER column filter. The plan was never corrected.
- AlphaEarth coverage bias (28.4%): Acknowledged in both the notebook header and REPORT. ✓
- API retry logic: Correctly implemented with exponential backoff in all notebooks. ✓
- PFAMs stores domain names, not accessions: Correctly identified; all queries use LIKE '%DUF4041%' (name), not LIKE '%PF13250%'. ✓
Statistical methods: Welch's t-test with Bonferroni correction for 64 simultaneous tests is appropriate and correctly implemented. Cohen's d is reported alongside p-values. PCA is used descriptively, not inferentially. ✓
Notebook organization: Each notebook follows a logical setup → query → analysis → save pattern. Markdown cells explain what each step does. ✓
Missing dependency: NB03 imports from sklearn.decomposition import PCA but scikit-learn does not appear in requirements.txt. The notebook will fail on a fresh install.
COG V query column mismatch (NB04, cell-7): The query uses WHERE pc.cogclass_id = 'V' but the table inspection in cell-5 reveals the actual column is cog_class_id. The 503 errors may have obscured this — but even if the API recovered, the query would return zero results due to the wrong column name. The fallback inspection correctly identifies the real column name but the defense-count query is never retried with the fix.
Figure title inconsistency (NB02, cell-4): The phylum distribution figure title reads "DUF4041 + GIY-YIG co-occurrence in gene clusters" but the analysis uses only DUF4041 as the SNIPE marker. This is misleading.
Findings Assessment
Finding 1 (ManYZ trade-off + fitness data): The ManXYZ fitness scores in the report (−2.75 to −4.14) are consistent with the query output shown in NB01, and the substrate specificity comparison (mannose vs. fructose) clearly supports the conclusion. The Methanococcus maripaludis SNIPE finding is interesting and well-sourced. The finding is well-supported. ✓
Corrected Pfam assignments: Strongly supported by InterPro and UniProt evidence. The report correctly explains why zero DUF4041+PF01541 co-occurrences is a biological truth, not an artifact. ✓
Finding 2 (1,696 species, 33 phyla): Supported by NB01 and NB02 outputs. However, there is a silent discrepancy in cluster counts that is not explained: NB01 queries 2,962 DUF4041 annotations from the API, but after the gene_cluster batch lookup and merge, the data frame grows to 4,572 rows — a 54% increase. The report consistently uses 4,572 but the discrepancy is never explained. One gene cluster annotation (query_name) may be associated with multiple species pangenomes (since the same genome may be in multiple species groups), but this needs explicit acknowledgment. Without it, readers cannot tell whether the 4,572 figure represents 4,572 distinct gene cluster IDs or 2,962 unique eggNOG annotations distributed across multiple species records.
Finding 3 (86.7% accessory/singleton): The output clearly shows Core=604 (13.3%), Accessory=3,934 (86.7%), Singleton=2,566 (56.5%). The math and percentages are consistent. ✓ Note: the report's accessory percentages cite "Accessory: 30.2% (1,380 clusters)" and "Singleton: 56.5% (2,583 clusters)" which don't match the notebook output (Accessory=3,934, Singleton=2,566). The notebook prints accessory as 86.7% combined (accessory+singleton), and the REPORT presents the split differently. The combined 86.7% is consistent, but the sub-totals in the REPORT appear to be from an earlier run or a different rounding basis and should be reconciled.
Finding 4 (PF13455 not PF01541): Well-supported with external evidence. ✓
Finding 5 (environmental niche): 22/64 AlphaEarth dimensions significant at Bonferroni threshold; effect sizes small (|d|=0.11–0.26). The conclusion "a consistent but modest environmental shift" is appropriately measured. ✓ The AlphaEarth dimensions (A00–A63) are not labeled by environmental variable in any notebook or figure — readers cannot interpret which environmental axes differ. Adding at least the top 3–5 interpretations of significant dimensions would substantially improve the environmental finding.
Finding 6 (Klebsiella SNIPE): Detection of DUF4041 in Klebsiella is supported by the PhageFoundry query output (1 eggNOG description match for 'DUF4041'). The ManYZ co-occurrence analysis providing 3 locus tags and domain counts is described as "ad hoc (not in a notebook)" — this critical supporting evidence has no code and cannot be reproduced or verified.
Finding 7 (functional annotations): The COG category distribution (D=1,533, T=1,118, J=1,066, M=834) is puzzling: COG D is "Cell division and chromosome partitioning" and COG J is "Translation, ribosomal structure and biogenesis." These are not expected for a defense gene, and the non-defense top descriptions (Histidine kinase ×319, seryl-tRNA aminoacylation ×238) account for ~12% of all DUF4041 clusters. This likely reflects false positives in the DUF4041 search — proteins that happen to contain DUF4041 as a minor domain but have primary functions elsewhere. The report notes this only implicitly; an explicit estimate of the false-positive rate (or a check using Description == "T5orf172" or "DUF4041" as a stricter filter) would strengthen the analysis.
Lambdavirus / PhageFoundry strain modelling: Interesting supplementary finding (0.5% infection rate for lambda vs 43.4% for Myoviridae) that contextualizes the ManYZ biology. Well-cited. ✓
Limitations section: Honest and thorough. The NMDC and defense enrichment limitations are clearly called out. ✓
PLAN_fitness_data_curation.md: A well-thought-out plan for follow-on work. Its checklist is entirely unchecked, confirming it's unexecuted. The README does not link to this file, so a reader would not know this next step exists.
Suggestions
-
(Critical) Fix
scikit-learnmissing fromrequirements.txt: Addscikit-learn>=1.3torequirements.txt. Without it, NB03 will fail on a fresh environment. This blocks reproducibility of the environmental analysis. -
(Critical) Explain the 2,962 → 4,572 cluster count jump: In NB01, the initial query returns 2,962 DUF4041 annotations but the final merged dataframe has 4,572 rows. Add a markdown cell explaining this: if the same eggNOG annotation (
query_name) appears in multiple species'gene_clusterentries (e.g., because gene_cluster_id is not globally unique), that is an important methodological note that affects how to interpret "4,572 clusters." -
(High) Reconcile accessory/singleton sub-counts in REPORT.md: The notebook output shows Accessory=3,934 (86.7%) and Singleton=2,566 (56.5%), but REPORT.md Finding 3 states "Accessory: 30.2% (1,380 clusters)" and "Singleton: 56.5% (2,583 clusters)." Update the report to match the actual notebook output, or explain why the figures differ.
-
(High) Move Klebsiella ManYZ co-occurrence analysis into a notebook cell: The ad-hoc analysis supporting the "DUF4041 + ManYZ co-occur in Klebsiella" conclusion (locus tags JJW41_17025, KFB13_RS08150, KFB31_RS01490; 3 DUF4041 proteins at 558 aa; 4,619 PTS_EIIC hits) is cited as "performed ad hoc (not in a notebook)." This key evidence for clinical relevance should be in a notebook cell, even if added as a new cell to NB04, so it can be verified and reproduced.
-
(High) Update RESEARCH_PLAN.md to correct Pfam IDs: The plan consistently uses
PF13291for DUF4041 (correct: PF13250) andPF01541for the SNIPE nuclease (correct: PF13455/Mug113). The queries in the plan also use these wrong accessions. Since the actual analysis correctly searched by domain name, the plan should be annotated or updated to show the corrected strategy so the plan and implementation are consistent. -
(High) Fix figure title in NB02: Cell-4 creates
snipe_phylum_distribution.pngwith title "SNIPE Homologue Distribution Across Bacterial Phyla (DUF4041 + GIY-YIG co-occurrence in gene clusters)" — but the analysis uses DUF4041 alone as the marker. Correct the title to "...DUF4041 domain in gene clusters" or similar. -
(Medium) Fix COG V query column name mismatch (NB04, cell-7): The table schema inspection in cell-5 shows the column is
cog_class_id, but the subsequent query usescogclass_id. Even if the 503 errors are resolved (e.g., by retrying with Spark Connect), the query will fail silently. Correct the column name and note whether the defense enrichment analysis should be retried via Spark Connect rather than REST API. -
(Medium) Add AlphaEarth dimension interpretation: Finding 5 reports that 22/64 dimensions differ significantly, with the largest effect on A19 (Cohen's d = 0.26). Since AlphaEarth dimensions encode satellite-derived environmental signals, add at minimum a reference to the AlphaEarth paper's description of what A19 and the other top dimensions capture (e.g., vegetation index, temperature, precipitation). Even noting "see AlphaEarth documentation for dimension semantics" would give readers a path to interpret the finding.
-
(Medium) Rename
snipe_both_domains.csvor clarify its contents: The filename implies it contains hits with both DUF4041 and the GIY-YIG nuclease domain, but the file actually contains all 4,572 DUF4041 clusters (used as the primary SNIPE marker, regardless of GIY-YIG co-occurrence). Rename toduf4041_snipe_clusters.csvor add a header comment explaining the naming, to avoid confusion with the true "both domains" analysis. -
(Medium) Discuss false-positive rate in DUF4041 annotations: ~12% of the 4,572 clusters have non-SNIPE primary functions (Histidine kinase, seryl-tRNA aminoacylation, etc.), and COG D/J dominate the COG distribution rather than COG M/V. Add a brief discussion of whether these represent true SNIPE proteins with non-obvious co-occurring annotations, or annotation artifacts (DUF4041 detected as a secondary domain). Filtering to
Description LIKE 'T5orf172%' OR Description LIKE '%DUF4041%'(3,675/4,572 = 80%) as a stricter SNIPE-specific set would provide a sensitivity check. -
(Low) Link PLAN_fitness_data_curation.md from README: The fitness data curation plan describes clear next steps for extending the project but is invisible to someone reading only README.md. Add a reference under a "Future Work" section in the README.
-
(Low) NB01 header markdown uses wrong Pfam IDs: The opening markdown cell states "DUF4041 (PF13291)" and "GIY-YIG (PF01541)." Update to reflect the project's corrected domain assignments (PF13250 and PF13455 respectively) so the notebook is internally consistent.
This review was generated by an AI system. It should be treated as advisory input, not a definitive assessment.
Visualizations
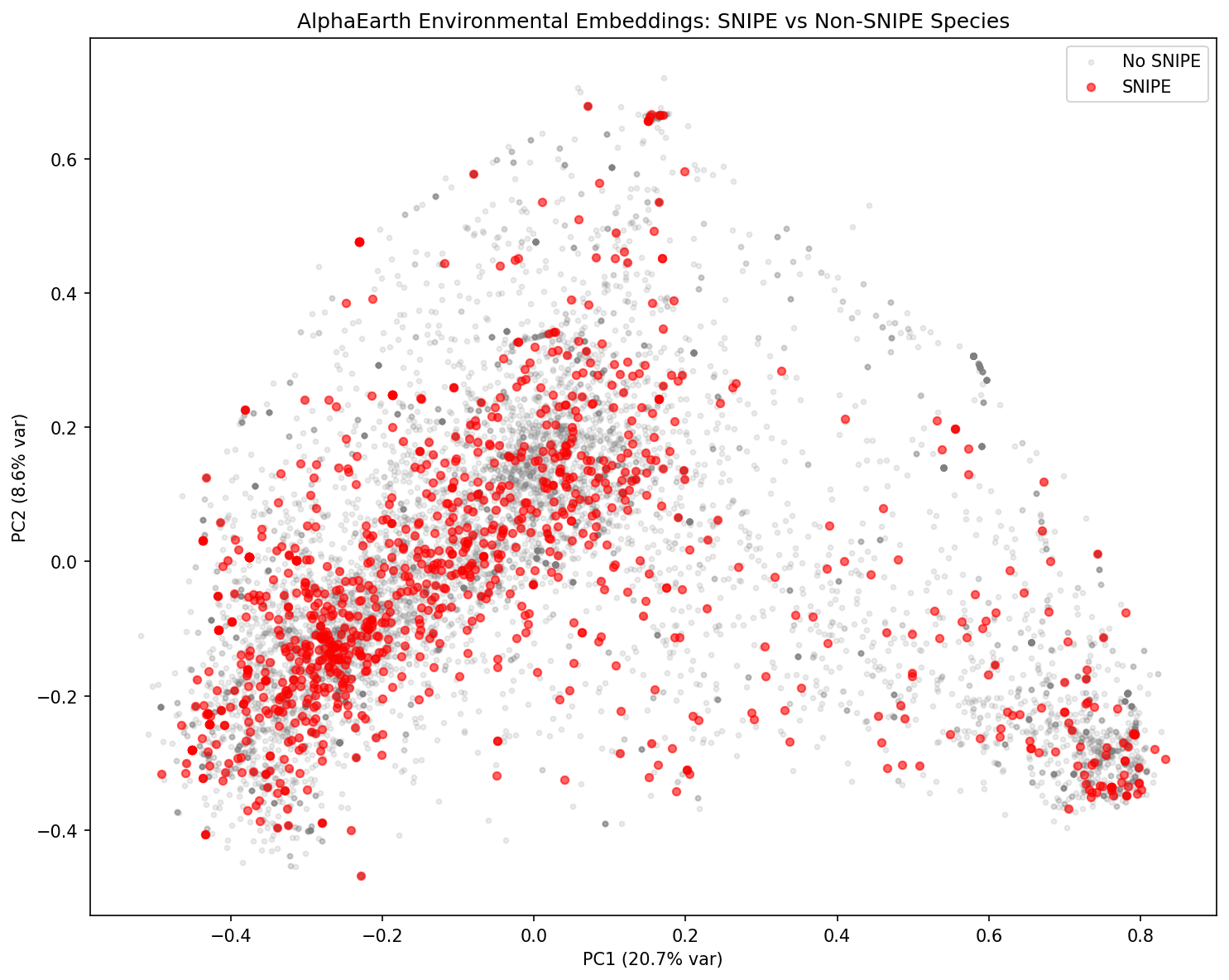
Snipe Alphaearth Pca
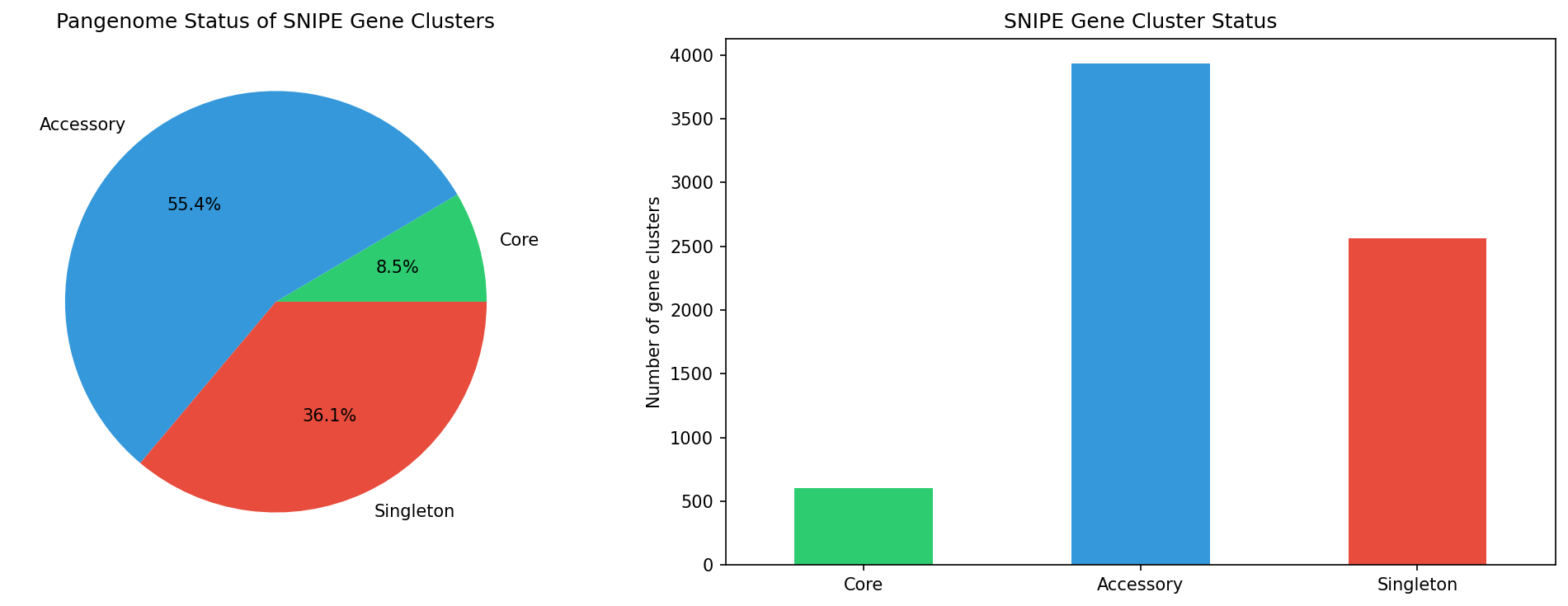
Snipe Pangenome Status
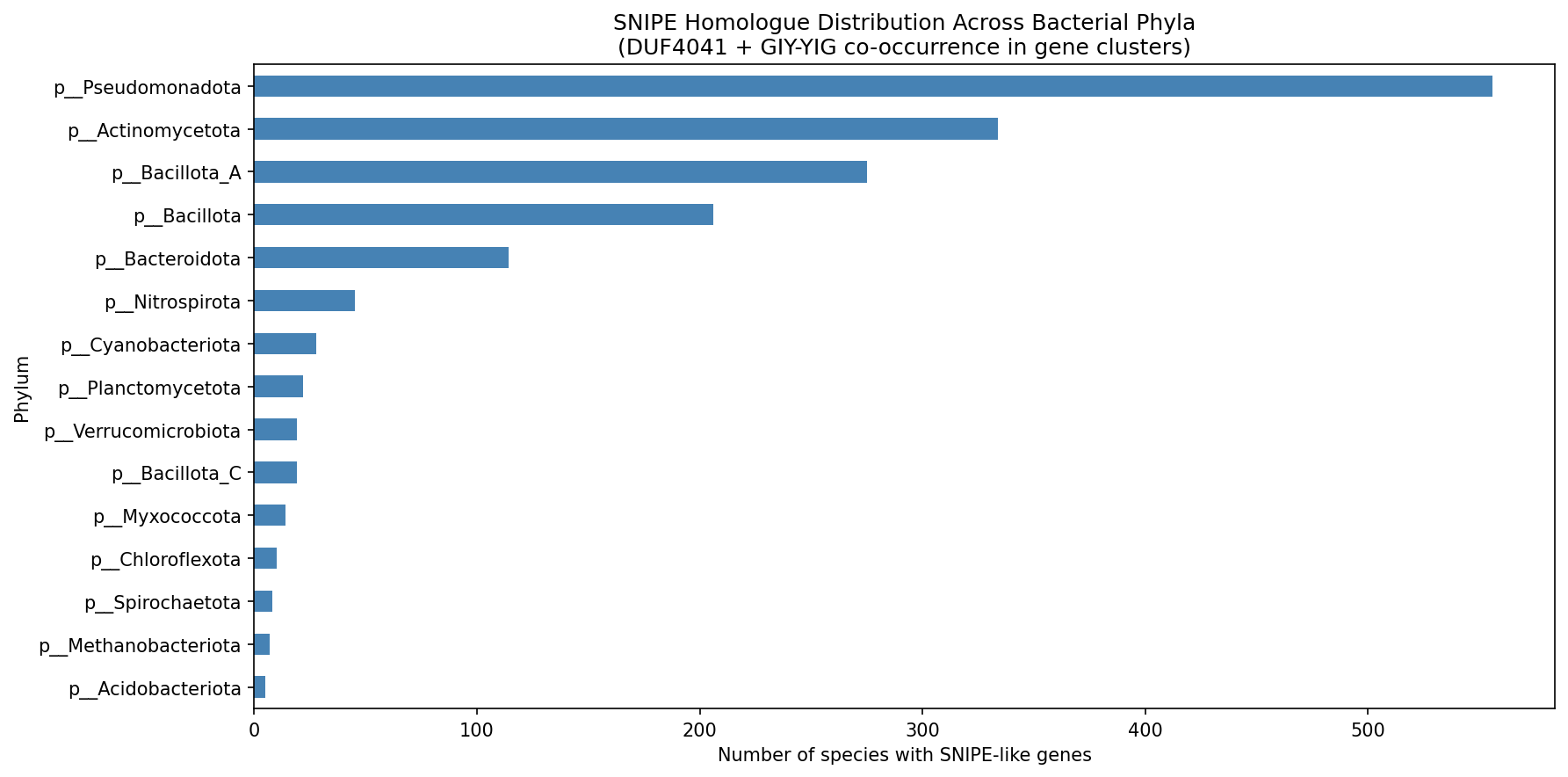
Snipe Phylum Distribution