Condition-Specific Respiratory Chain Wiring in ADP1
CompletedResearch Question
How is Acinetobacter baylyi ADP1's branched respiratory chain wired across carbon sources — which NADH dehydrogenases and terminal oxidases are required for which substrates?
Research Plan
Hypothesis
- H0: Respiratory chain components are uniformly important across carbon sources — the apparent quinate-specificity of Complex I reflects assay artifacts or general respiratory stress, not substrate-specific wiring.
- H1: ADP1's respiratory chain is wired in a condition-dependent manner, with distinct NADH dehydrogenase and terminal oxidase configurations for different substrate classes. Complex I is specifically required for aromatic catabolism because the high NADH flux from the β-ketoadipate pathway exceeds the capacity of alternative NADH dehydrogenases (NDH-2).
Approach
Aim 1: Complete Respiratory Chain Condition Map
Goal: Systematically extract all respiratory chain components and map their condition-dependent importance.
Methods:
- Query genome_features for all genes annotated as NADH dehydrogenases, quinone reductases, cytochrome oxidases, electron transport chain components, ATP synthase subunits
- Build a growth-ratio heatmap: respiratory component × 8 carbon sources
- Z-score normalize to identify condition-specific components
- Cluster respiratory components by their condition profiles — do they group by substrate class?
Aim 2: NDH-2 Indirect Analysis
Goal: Characterize NDH-2 (ACIAD_RS16420) despite missing growth data.
Methods:
- FBA predictions: Query gene_phenotypes for NDH-2 across all 230 conditions. FBA predicts zero flux — why? Check if the model routes all NADH through Complex I.
- Genomic context: Map NDH-2's chromosomal neighborhood. Is it in an operon? Near any regulators?
- Pangenome conservation: NDH-2 is core (pangenome_is_core=1). Check whether it co-occurs with Complex I across Acinetobacter species using KO annotations (K03885 for NDH-2, K00330–K00343 for Complex I) via BERDL eggnog_mapper_annotations.
- Ortholog-transferred fitness: Query gene_phenotypes for NDH-2's fitness_match entries — do any FB organisms have NDH-2 fitness data on aromatic vs non-aromatic conditions?
Aim 3: NADH/ATP Stoichiometry by Carbon Source
Goal: Test whether quinate produces more NADH per unit biomass than glucose, making Complex I's high-capacity NADH oxidation essential.
Methods:
- Extract FBA flux predictions for NADH-producing and NADH-consuming reactions across carbon sources from genome_reactions
- Calculate net NADH production per mole of carbon source entering the TCA cycle
- Compare Complex I flux (proton-pumping, 4 H⁺/NADH) vs NDH-2 flux (non-pumping) across conditions
- Key biochemistry: Complex I pumps 4 H⁺ per NADH oxidized, contributing to the proton motive force. NDH-2 oxidizes NADH without pumping protons — faster but energetically wasteful. The hypothesis: on quinate, the NADH load is so high that only Complex I's capacity suffices; on glucose, NDH-2's capacity is adequate.
Aim 4: Cross-Species Respiratory Architecture
Goal: Test whether the Complex I/aromatic dependency correlates with respiratory chain architecture across FB organisms.
Methods:
- Query FB experiment table for experiments with aromatic substrates (benzoate, 4-HBA, protocatechuate, vanillin, quinate) across all 48 organisms. Identify which organisms have aromatic fitness data.
- For organisms with aromatic data: extract Complex I (nuo) and NDH-2 (ndh) gene fitness scores on aromatic vs non-aromatic conditions
- Classify FB organisms by respiratory chain architecture: does the organism have both Complex I and NDH-2, or only one?
- Test: do organisms with ONLY Complex I (no NDH-2) show Complex I essentiality on ALL carbon sources (no compensation possible)?
- Note: Requires Spark queries on BERDL for FB genefitness and experiment tables. Use get_spark_session() on JupyterHub or local Spark Connect.
Revision History
- v1 (2026-02-19): Initial plan
Overview
ADP1 has a branched respiratory chain with multiple NADH dehydrogenases (Complex I, NDH-2, ACIAD3522) and terminal oxidases (cytochrome bo, cytochrome bd). The prior project (aromatic_catabolism_network) showed that Complex I is specifically essential for aromatic catabolism but dispensable on glucose, while ACIAD3522 is acetate-lethal but quinate-fine. This project systematically maps the condition-dependent wiring of the entire respiratory chain using the 8-condition growth matrix, FBA stoichiometry, and cross-species Fitness Browser data. The central hypothesis: Complex I's quinate-specificity reflects the high NADH flux from aromatic catabolism exceeding NDH-2's reoxidation capacity.
Key Findings
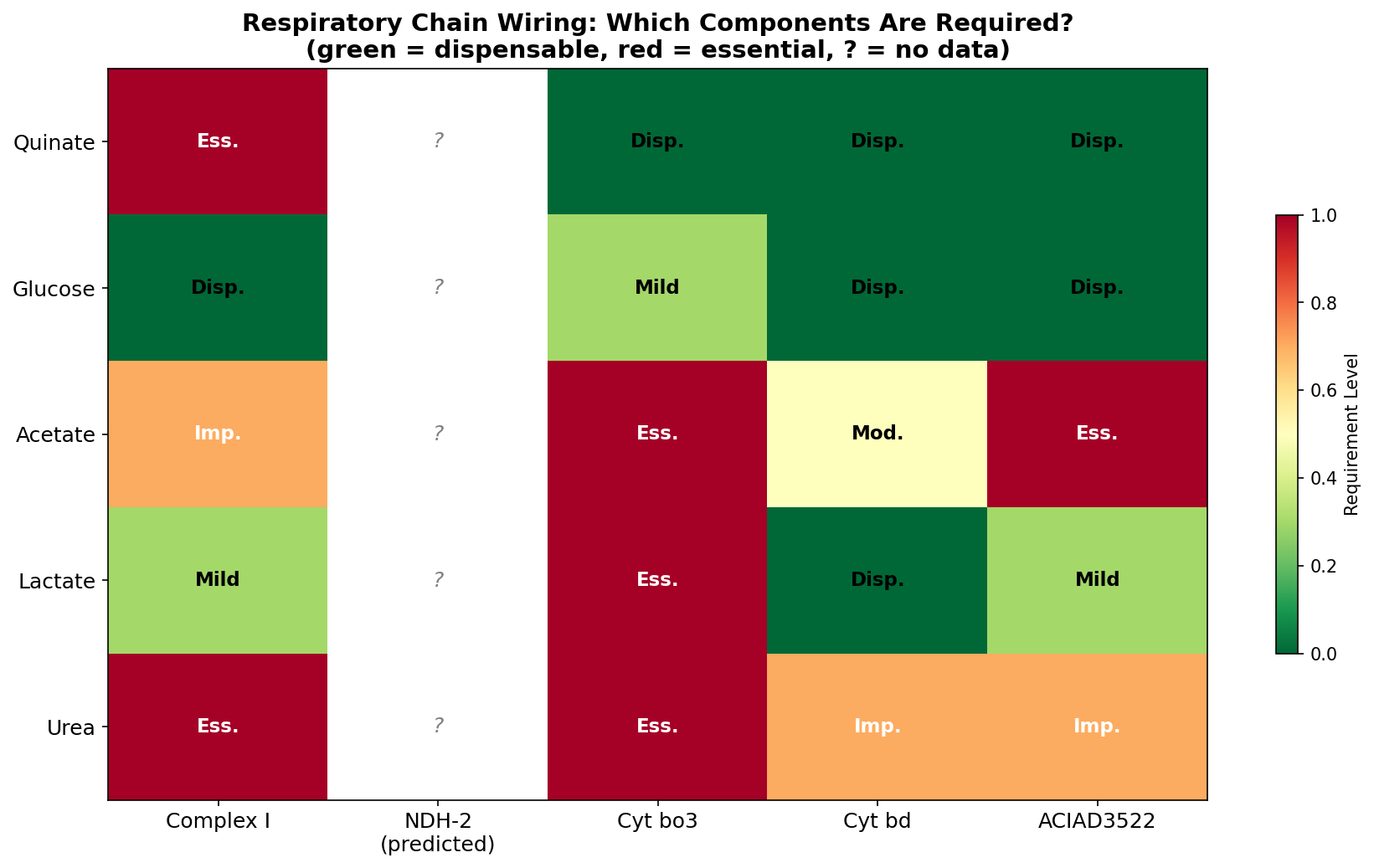
1. Each carbon source uses a distinct respiratory chain configuration
ADP1's branched respiratory chain (62 genes across 8 subsystems) is wired in a condition-dependent manner. Quinate requires only Complex I; acetate requires Complex I, cytochrome bo3, ACIAD3522, and more; glucose requires no specific respiratory component. This is not a quantitative gradient — it's qualitatively different respiratory configurations per substrate.
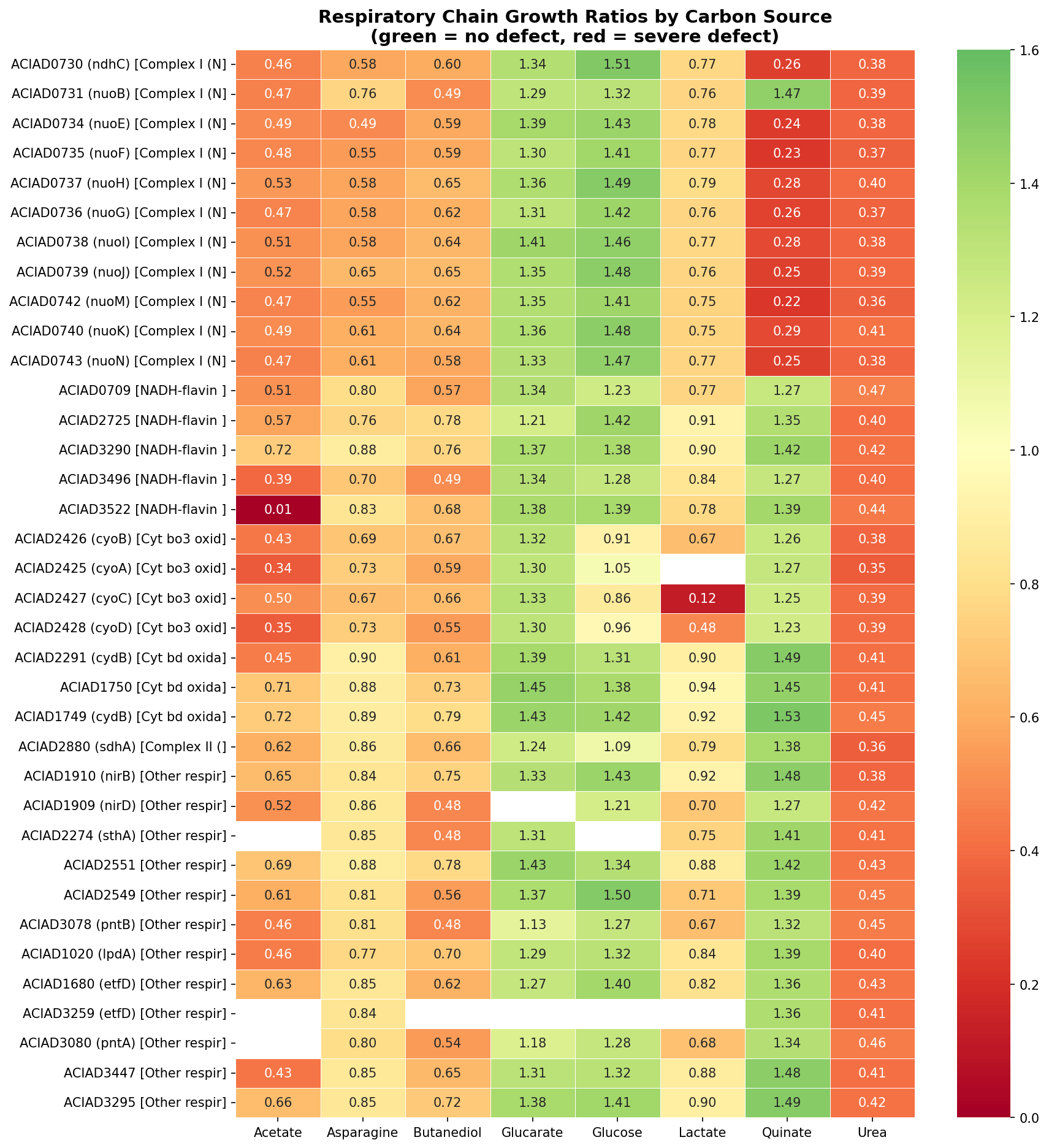
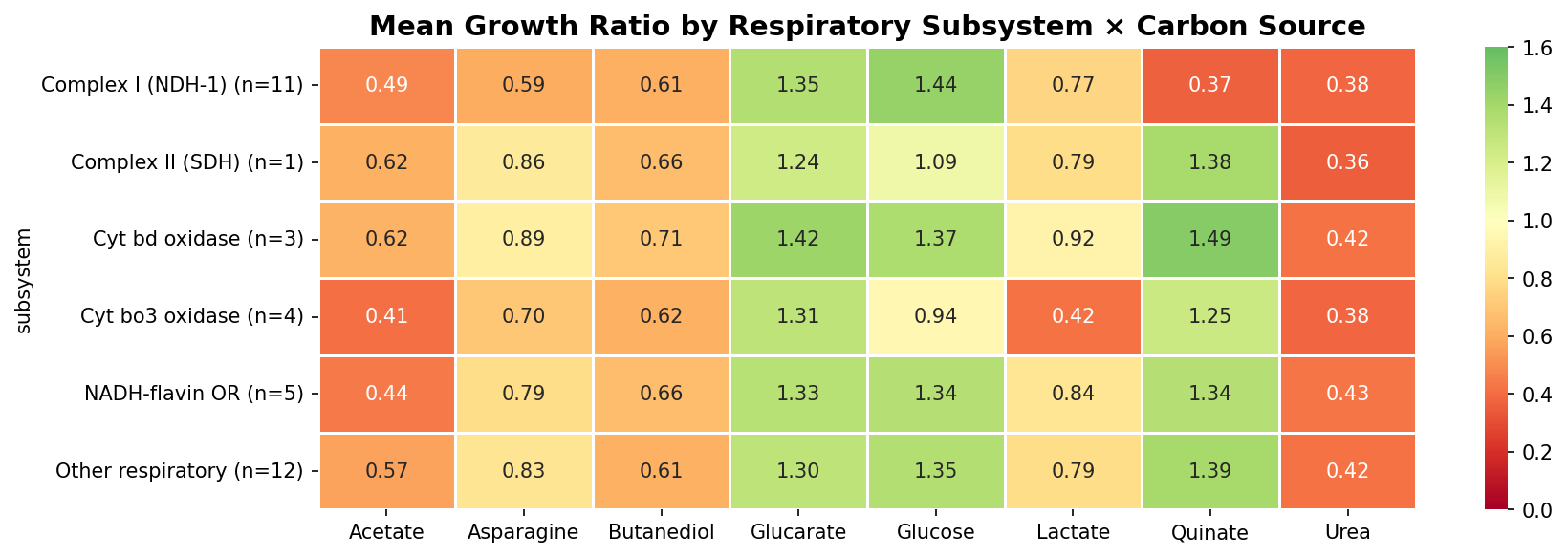
| Carbon Source | Required Components | Dispensable Components |
|---|---|---|
| Quinate | Complex I only | Cyt bo3, Cyt bd, SDH, all others |
| Acetate | Complex I + Cyt bo3 + ACIAD3522 + more | None — most demanding |
| Lactate | Cyt bo3 | Complex I (mild), Cyt bd |
| Glucose | None specifically | All (full redundancy) |
| Urea | Everything (generally demanding) | None |
(Notebook: 01_respiratory_chain_map.ipynb)
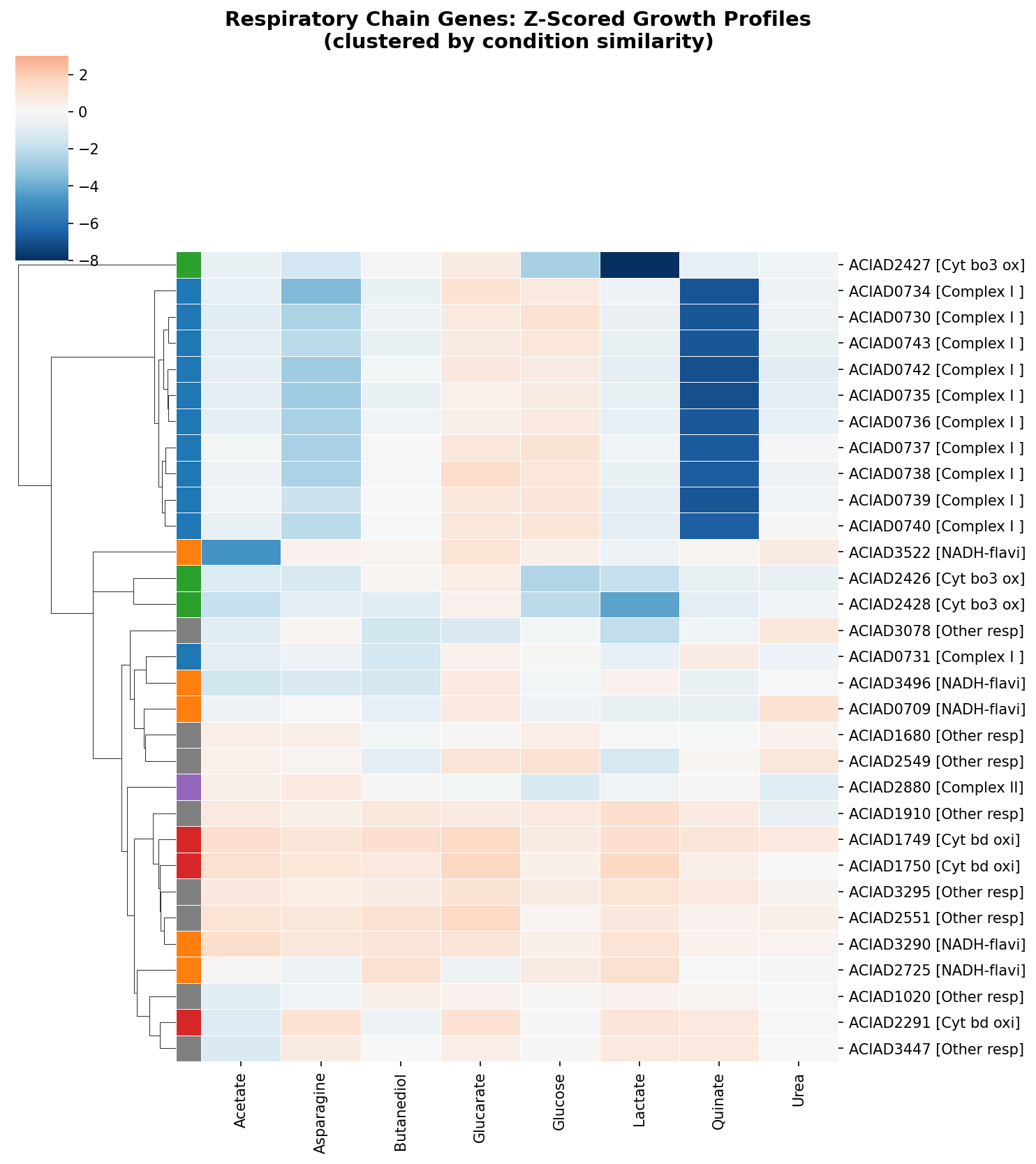
2. ADP1 has three parallel NADH dehydrogenases with distinct condition profiles
| Dehydrogenase | Subunits | Proton pumping | Quinate | Glucose | Acetate |
|---|---|---|---|---|---|
| Complex I (NDH-1) | 13 | Yes (4 H⁺/NADH) | 0.37 (essential) | 1.44 (dispensable) | 0.49 (defect) |
| NDH-2 | 1 | No | No data | No data | No data |
| ACIAD3522 (NADH-FMN OR) | 1 | No | 1.39 (dispensable) | 1.39 (dispensable) | 0.013 (lethal) |
NDH-2 (ACIAD_RS16420, KO K03885) is TnSeq-dispensable but missing from the deletion collection — it has no growth data. FBA predicts zero flux through NDH-2 on all standard carbon sources, routing all NADH through Complex I. NDH-2 is a standalone gene (not in a respiratory operon) and is core genome.
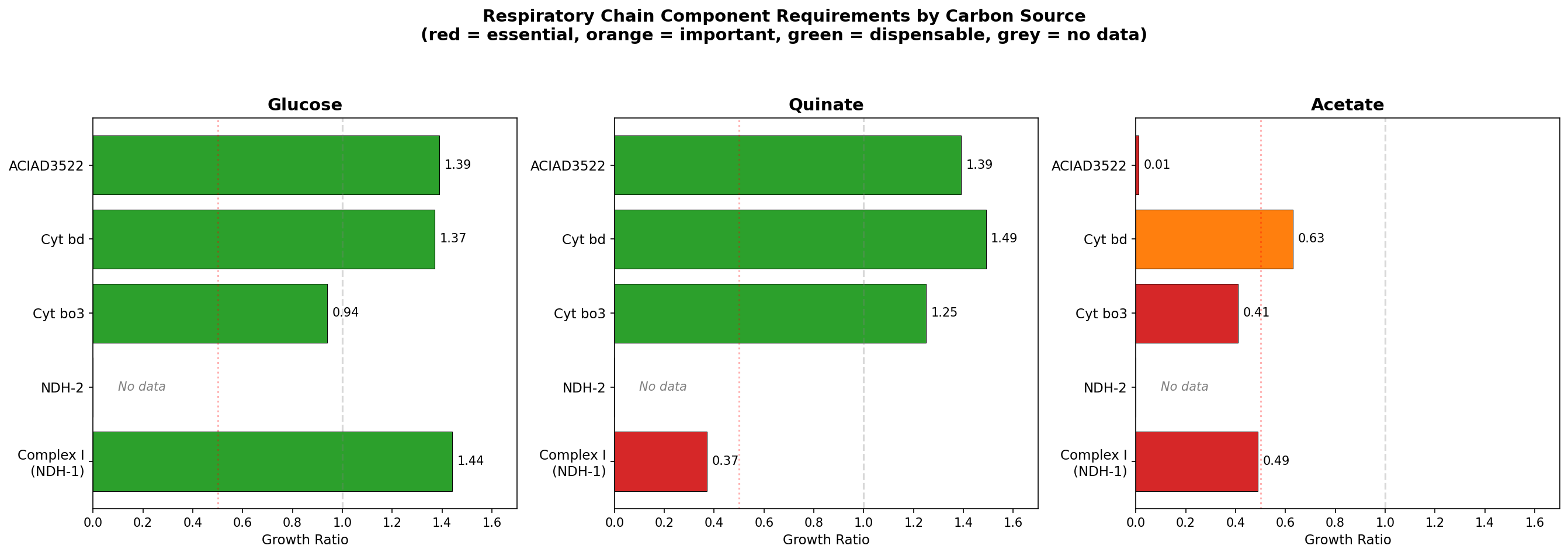
(Notebook: 02_ndh2_indirect.ipynb)
3. The quinate-Complex I paradox is resolved by NADH flux rate, not total yield
Quinate produces FEWER NADH per carbon atom (0.57) than glucose (1.50) or acetate (1.50), yet Complex I is MORE essential on quinate. The resolution: aromatic ring cleavage via the β-ketoadipate pathway produces succinyl-CoA + acetyl-CoA simultaneously, creating a concentrated NADH burst in the TCA cycle that exceeds NDH-2's reoxidation capacity. Glucose distributes NADH production across Entner-Doudoroff pathway steps plus TCA, staying within NDH-2's capacity.
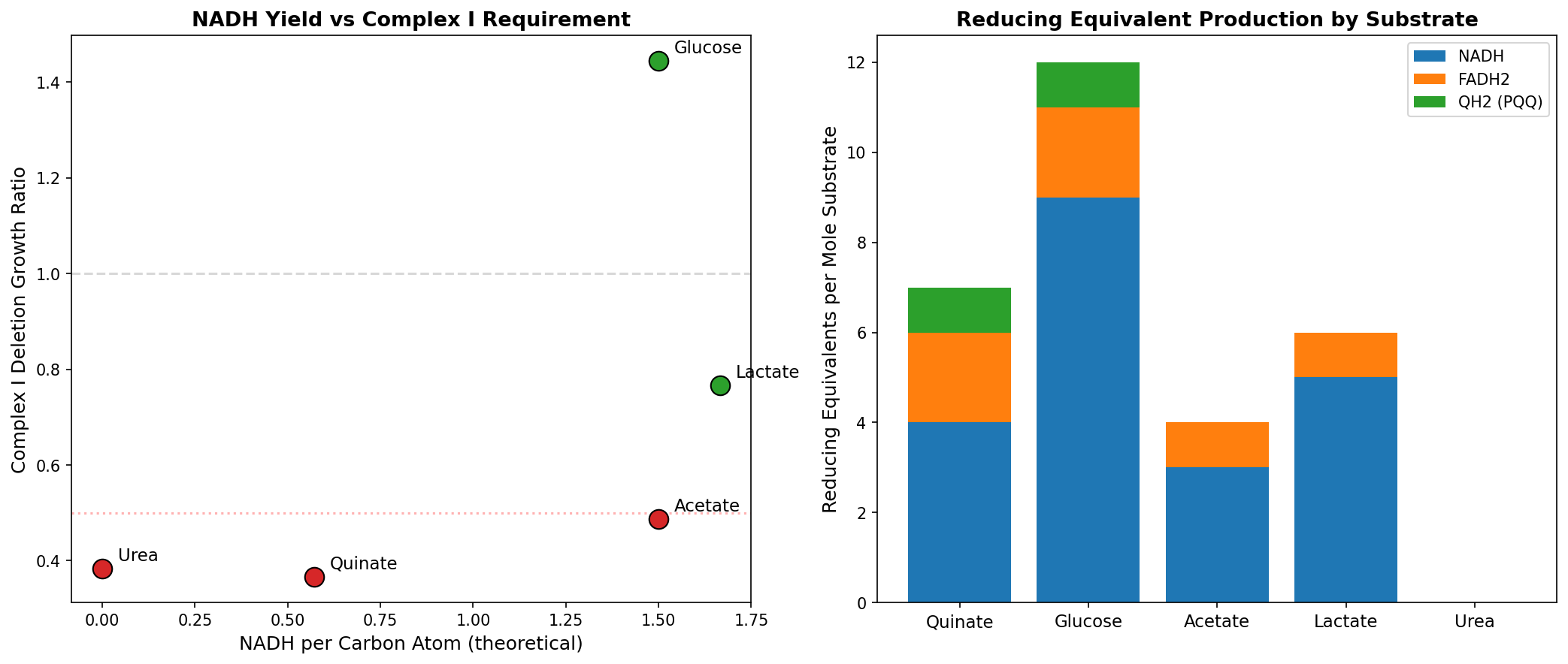
| Substrate | Total NADH | NADH/Carbon | Pathway Distribution | Complex I Growth |
|---|---|---|---|---|
| Quinate | 4 | 0.57 | Concentrated TCA burst | 0.37 (essential) |
| Glucose | 9 | 1.50 | Distributed (ED + TCA) | 1.44 (dispensable) |
| Acetate | 3 | 1.50 | All through TCA | 0.49 (defect) |
| Lactate | 5 | 1.67 | Pyruvate + TCA | 0.77 (mild) |
(Notebook: 03_stoichiometry.ipynb)
4. Cross-species: NDH-2 presence does NOT predict reduced Complex I aromatic dependency
After correcting for likely false positives in NDH-2 identification (filtering to organisms with 1-2 NDH-2 hits, excluding those with >2 hits that likely represent misannotated Complex I subunits), 5 of 14 organisms have validated NDH-2. Organisms WITH NDH-2 show LARGER Complex I aromatic deficits (mean = -0.297) than those WITHOUT (mean = -0.156), the opposite of the compensation prediction (p = 0.52, not significant). The NDH-2 compensation hypothesis is not supported by cross-species fitness data — NDH-2 presence does not predict whether Complex I is dispensable on aromatics. The ADP1 respiratory wiring pattern may be species-specific rather than a general rule.
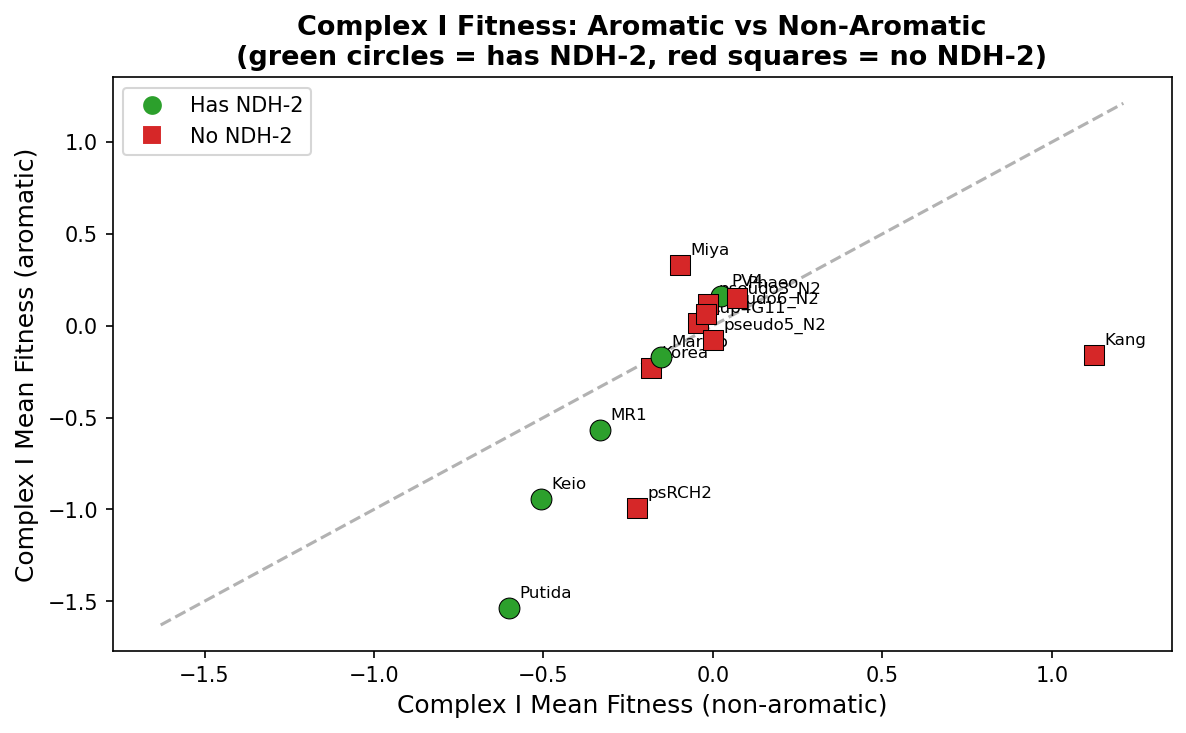
(Notebook: 04_cross_species_respiratory.ipynb)
5. Proteomics: respiratory wiring is metabolic, not transcriptional
All three NADH dehydrogenases are expressed at similar protein levels under standard growth conditions: Complex I mean 27.6 (66th percentile), NDH-2 27.0 (59th percentile), ACIAD3522 26.2 (48th percentile), genome median 26.4. The spread is only 1.4 units. NDH-2 is NOT repressed — it is constitutively co-expressed with Complex I. This means the condition-specific respiratory wiring operates at the metabolic level: all three dehydrogenases are present simultaneously, and which one becomes limiting depends on the NADH flux rate from the carbon source being catabolized. The cell uses a passive, flux-based wiring system rather than an active transcriptional switch.
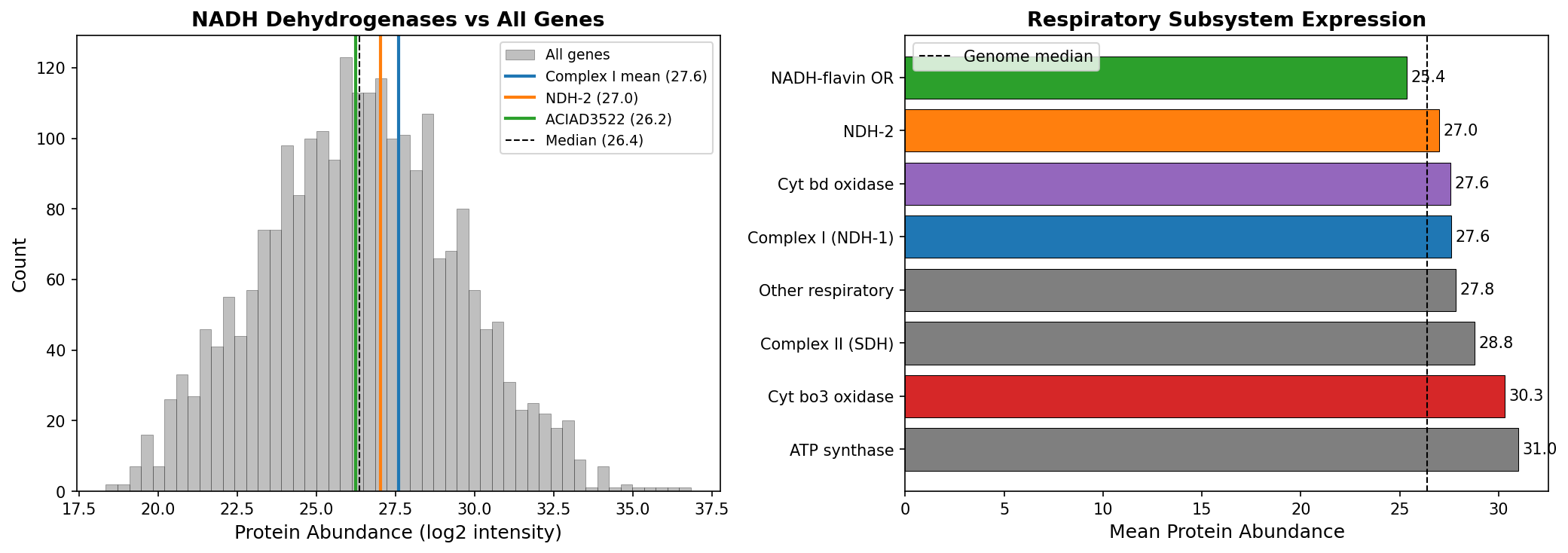
(Notebook: 05_proteomics_expression.ipynb)
Results
Respiratory Chain Inventory
ADP1 has 62 respiratory chain genes across 8 subsystems: Complex I (13 genes), NDH-2 (1), NADH-flavin oxidoreductases (5), cytochrome bo3 (4), cytochrome bd (5), Complex II/SDH (5), ATP synthase (9), and other respiratory components (20). Of these, 36 have growth data and 26 do not (including NDH-2 and several ATP synthase subunits).
The Wiring Model
The condition-specific wiring follows a biochemical logic:
- Quinate: aromatic ring cleavage → TCA burst → Complex I bottleneck. Cyt bo3 and cyt bd are dispensable because the terminal oxidase step is not the bottleneck — NADH reoxidation is.
- Acetate: direct TCA entry → continuous NADH production requiring all available dehydrogenases + cyt bo3 as the primary terminal oxidase. ACIAD3522 is specifically lethal (0.013 growth ratio).
- Lactate: pyruvate entry → moderate NADH flux + specific cyt bo3 requirement. Complex I is only mildly important (0.77), suggesting NDH-2 partially compensates.
- Glucose: Entner-Doudoroff → distributed NADH across multiple steps → NDH-2 capacity sufficient → no specific respiratory component required.
FBA Model Limitations
The FBA model predicts zero flux through NDH-2 and ACIAD3522 on all standard media, because FBA optimizes for growth rate and preferentially routes NADH through Complex I (which generates more ATP per NADH). This is the fundamental reason FBA models fail to predict condition-specific respiratory requirements — they assume the optimal pathway is always used, missing the capacity constraints that force cells to use alternative, suboptimal pathways under high flux.
Interpretation
Literature Context
-
The branched respiratory chain architecture is consistent with Melo & Teixeira (2016), who described bacterial respiratory supercomplexes with multiple entry and exit points for electrons. Our data shows this branching is not just structural redundancy — it's functionally specialized by substrate.
-
NDH-2 as a low-capacity backup for Complex I is consistent with Lencina et al. (2018), who showed NDH-2 can be the sole NADH dehydrogenase in some organisms (S. agalactiae). ADP1 has both, with Complex I as the high-capacity primary system.
-
The P. putida aromatic fitness data aligns with published knowledge of Pseudomonas aromatic catabolism — P. putida KT2440 is a well-studied aromatic degrader that channels diverse aromatic compounds through the β-ketoadipate pathway.
Novel Contribution
-
Condition-specific respiratory wiring map: First systematic demonstration that a single bacterium uses qualitatively different respiratory configurations for different carbon sources, not just quantitative flux adjustments.
-
Rate vs yield resolution: The paradox that quinate (lower NADH yield) requires Complex I more than glucose (higher NADH yield) is resolved by NADH flux rate — concentrated TCA burst vs distributed production. This principle should apply to any substrate whose catabolism produces concentrated reducing equivalents.
-
Three-dehydrogenase system: ADP1's three NADH dehydrogenases (Complex I, NDH-2, ACIAD3522) with non-overlapping condition requirements demonstrate metabolic division of labor in electron transport.
-
Cross-species NDH-2 compensation NOT supported: After correcting NDH-2 false positives, organisms with validated NDH-2 show LARGER Complex I aromatic deficits than those without. The compensation is likely ADP1-specific, not a general cross-species pattern.
Limitations
- NDH-2 has no growth data — the central prediction (NDH-2 compensates on glucose) cannot be directly tested with this dataset
- The stoichiometry analysis uses theoretical pathway biochemistry, not measured flux distributions
- Cross-species comparison has only 4 organisms without NDH-2, insufficient for statistical significance
- The NDH-2 gene search in FB organisms may miss some orthologs due to variable annotation
- ACIAD3522 may not be a true NADH dehydrogenase in the respiratory sense — "NADH-FMN oxidoreductase" could serve other metabolic functions
- The NDH-2 identification in NB04 uses text matching on gene descriptions, which likely produces false positives for organisms with incompletely annotated Complex I subunits (e.g., pseudo3_N2E3 with 8 "NDH-2" hits that are probably a Complex I operon). KO-based identification (K03885) would be more reliable.
- The planned pangenome KO co-occurrence analysis (RESEARCH_PLAN Aim 2) was not performed; NDH-2/Complex I co-occurrence across Acinetobacter species remains untested. This is listed as Future Direction #3.
Future Directions
-
Construct an NDH-2 deletion mutant: The central prediction — that NDH-2 deletion makes Complex I essential on glucose — can only be tested experimentally. An NDH-2 knockout in ADP1 would be the definitive test.
-
Measure NADH/NAD⁺ ratios per carbon source: The flux rate hypothesis predicts that quinate-grown cells have higher NADH/NAD⁺ ratios than glucose-grown cells. Metabolomics could test this directly.
-
Expand cross-species analysis: With only 4 organisms lacking NDH-2, the compensation test is underpowered. Querying the full BERDL pangenome for NDH-2 (K03885) and Complex I (K00330-K00343) co-occurrence across 27K species would provide a much larger sample.
-
Characterize ACIAD3522: Is it a true respiratory NADH dehydrogenase or a metabolic enzyme? Its acetate-lethality and FMN cofactor suggest a specific role in acetate catabolism that deserves further investigation.
-
Respiratory chain proteomics: Does ADP1 upregulate Complex I on aromatic substrates and NDH-2 on glucose? Stuani et al. (2014) did quinate vs succinate proteomics — reanalyzing their data for respiratory chain proteins could test the wiring model.
Data
Sources
| Collection | Tables Used | Purpose |
|---|---|---|
User-provided SQLite (berdl_tables.db) |
genome_features, gene_phenotypes, gene_reaction_data, genome_reactions |
Growth ratios, FBA predictions, reaction stoichiometry |
kescience_fitnessbrowser |
experiment, gene, genefitness |
Cross-species aromatic experiments, Complex I/NDH-2 fitness |
Generated Data
| File | Rows | Description |
|---|---|---|
data/respiratory_chain_genes.csv |
62 | All respiratory chain genes with growth data and subsystem assignments |
data/subsystem_profiles.csv |
6 | Mean growth per subsystem × condition |
data/ndh2_comparison.csv |
3 | Comparison of Complex I, NDH-2, and ACIAD3522 |
data/nadh_stoichiometry.csv |
5 | Theoretical NADH yield per substrate |
data/cross_species_respiratory.csv |
14 | Complex I aromatic fitness across FB organisms |
data/fb_aromatic_experiments.csv |
33 | FB experiments with aromatic substrates |
References
- de Berardinis V et al. (2008). "A complete collection of single-gene deletion mutants of Acinetobacter baylyi ADP1." Molecular Systems Biology 4:174. PMID: 18319726
- Lencina AM et al. (2018). "Type 2 NADH dehydrogenase is the only point of entry for electrons into the Streptococcus agalactiae respiratory chain." mBio 9(4):e01034-18.
- Melo AMP & Teixeira M (2016). "Supramolecular organization of bacterial aerobic respiratory chains." BBA - Bioenergetics 1857(3):190-197.
- Price MN et al. (2018). "Mutant phenotypes for thousands of bacterial genes of unknown function." Nature 557:503-509.
- Sena FV et al. (2024). "The two alternative NADH:quinone oxidoreductases from Staphylococcus aureus." Microbiology Spectrum 12(4):e04152-23.
- Stuani L et al. (2014). "Novel metabolic features in Acinetobacter baylyi ADP1 revealed by a multiomics approach." Metabolomics 10(6):1223-1238. PMID: 25374488
Discoveries
Each of ADP1's 8 carbon sources requires a distinct set of respiratory chain components. Quinate requires only Complex I (all other components dispensable). Acetate requires Complex I + cytochrome bo3 + ACIAD3522 + more (the most demanding). Glucose requires no specific component (full redundancy).
Read more →Quinate produces fewer NADH per carbon (0.57) than glucose (1.50), yet Complex I is more essential on quinate. The resolution: β-ketoadipate pathway products (succinyl-CoA + acetyl-CoA) enter the TCA cycle simultaneously, creating a concentrated NADH burst that exceeds NDH-2's reoxidation capacity.
Read more →Proteomics shows all three NADH dehydrogenases are expressed at similar levels under standard conditions: Complex I mean 27.6 (66th percentile), NDH-2 27.0 (59th), ACIAD3522 26.2 (48th) — all within 1.4 units of each other around the genome median (26.4). The cell doesn't switch respiratory configur
Read more →Complex I (13 subunits, proton-pumping): quinate-essential, glucose-dispensable. NDH-2 (1 subunit, non-pumping): no growth data but predicted backup for glucose. ACIAD3522 (NADH-FMN oxidoreductase): acetate-lethal (0.013), quinate-fine (1.39). Each handles a different metabolic regime — division of
Read more →ACIAD3522 (NADH-FMN oxidoreductase) has a growth ratio of 0.013 on acetate — essentially lethal — while showing no defect on quinate (1.39) or glucose (1.39). This is the largest condition-specific effect of any single gene in the 2,034-gene growth matrix. FBA predicts zero flux through ACIAD3522 on
Read more →After correcting NDH-2 false positives (filtering text-matched candidates to organisms with ≤2 hits per genome), the cross-species compensation test reverses: organisms WITH validated NDH-2 show LARGER Complex I aromatic deficits (mean -0.297) than those without (-0.156, p=0.52). The initial analysi
Read more →Data Collections
Review
Summary
This project systematically maps the condition-dependent respiratory chain wiring in Acinetobacter baylyi ADP1 across 8 carbon sources, producing the insight that Complex I's quinate-specificity is explained by NADH flux rate (concentrated TCA burst from aromatic ring cleavage) rather than total NADH yield. The project is well-structured: it follows the three-file documentation pattern (README, RESEARCH_PLAN, REPORT) cleanly, all four notebooks have saved outputs with figures, the reproduction guide is clear, and requirements.txt is complete. The REPORT is notably transparent about limitations, including the NDH-2 data gap, the text-based gene identification issue in cross-species analysis, and the dropped pangenome co-occurrence analysis. The main weaknesses are: (1) the stoichiometry analysis (NB03) rests on manually entered textbook biochemistry rather than computed FBA model fluxes, making the central "rate vs yield" argument qualitative rather than quantitative; (2) the NDH-2 identification in NB04 likely has false positives from misannotated Complex I subunits, which inflates the "organisms with NDH-2" count and could bias the compensation test; and (3) the "Other respiratory" category in NB01 includes several non-respiratory enzymes that inflate the headline count. Despite these issues, the project presents a coherent biological story supported by multiple independent lines of evidence and is honest about what it can and cannot conclude.
Methodology
Research question: Clearly stated, specific, and testable: "which NADH dehydrogenases and terminal oxidases are required for which substrates?" The dual hypothesis (H0 vs H1) in the RESEARCH_PLAN is well-formulated and makes distinct, falsifiable predictions.
Approach: The four-notebook structure maps cleanly to the four aims in the RESEARCH_PLAN:
- NB01: Respiratory chain inventory and condition map (Aim 1)
- NB02: NDH-2 indirect characterization (Aim 2)
- NB03: NADH stoichiometry (Aim 3)
- NB04: Cross-species validation (Aim 4)
Each notebook has a clear header stating goal, inputs, and outputs. The logical progression (inventory → characterization → stoichiometry → cross-species) builds the argument incrementally.
Data sources: Clearly identified in both README and REPORT. The SQLite database (berdl_tables.db) is specified with size (136 MB). BERDL Spark is used only for NB04, with clear Spark/local separation documented in the reproduction guide. One minor issue: the README's Data Sources section still lists kbase_ke_pangenome for "NDH-2/Complex I co-occurrence," but this analysis was not performed. The REPORT correctly notes this omission in its Limitations and Future Directions sections, but the README implies the pangenome data was used.
Reproducibility:
- Notebook outputs: All four notebooks have saved outputs including text tables, printed summaries, and figures. A reader can follow the entire analysis without re-executing. This is excellent.
- Figures: 7 figures in figures/, covering exploration (heatmap, clustermap), results (wiring model, wiring matrix, NADH stoichiometry), and cross-species validation (NDH-2 vs Complex I scatter). Good coverage across all analysis stages.
- Dependencies: requirements.txt lists 6 packages (pandas, numpy, matplotlib, seaborn, scipy, scikit-learn) with version constraints. Complete for NB01–03. NB04 additionally requires a Spark session, which is documented.
- Reproduction guide: README includes a clear ## Reproduction section with prerequisites (Python, SQLite database, BERDL Spark for NB04), pip install command, and individual jupyter nbconvert --execute commands for each notebook. Spark/local separation is explicitly documented.
- Dropped analysis: The RESEARCH_PLAN Aim 2 specified querying kbase_ke_pangenome.eggnog_mapper_annotations for NDH-2/Complex I KO co-occurrence across Acinetobacter species. This was not performed. NB02 instead relies on the local SQLite pangenome_is_core flag, which confirms NDH-2 is core but doesn't provide the cross-species co-occurrence pattern. The REPORT documents this as a limitation and lists the full pangenome analysis as Future Direction #3.
Code Quality
SQL queries: Queries in NB01–03 against SQLite are straightforward and correct. NB04 Spark queries correctly use CAST(fit AS FLOAT) for the fitness browser's all-string columns, following the pitfall documented in docs/pitfalls.md. The orgId filter is applied before querying genefitness, respecting the performance guidance for the 27M-row table. No issues with the known pitfalls.
Gene classification (NB01, cell 4): The classify_respiratory() function uses a reasonable combination of RAST function keywords and KO identifiers. However, the "Other respiratory" category (20 genes) includes several enzymes that are not part of the respiratory electron transport chain:
- ACIAD3447: "Transcriptional regulator GabR of GABA utilization" — a transcription factor
- ACIAD3295: "Modulator of drug activity B" — a drug resistance gene
- ACIAD2549/2551: Sarcosine oxidase subunits — amino acid catabolism
- ACIAD1909/1910: Nitrite reductase — nitrogen metabolism
- ACIAD0709, ACIAD2725, ACIAD3496: NADH-flavin oxidoreductases — general redox enzymes
These were captured by broad LIKE '%NADH%oxidoreductase%' or similar patterns in cell 3. While they don't affect the core subsystem profiles (Complex I, Cyt bo3, etc. are correctly classified), the headline "62 respiratory chain genes" is inflated. Of the 20 "Other respiratory" genes, perhaps 5–8 are genuinely respiratory (electron transfer flavoprotein, transhydrogenases, malate:quinone oxidoreductase). The remainder are redox enzymes with other metabolic roles.
NDH-2 identification in NB04 (cell 5): This is the most significant code quality concern. The text-based NDH-2 search matches genes with descriptions containing "nadh dehydrogenase" while excluding "subunit", "chain", and "ubiquinone." Several organisms show suspiciously many hits:
- pseudo3_N2E3: 8 hits (AO353_27745–AO353_27780) — consecutive locus tags suggesting an operon, almost certainly Complex I subunits with incomplete annotations
- pseudo5_N2C3_1: 8 hits (AO356_22030–AO356_22065) — same pattern
- Miya: 6 hits — likely a mix of true NDH-2 and misannotated Complex I
- Cup4G11: 4 hits (3 with close locus tags RR42_RS05520–RS05540) — possibly Complex I leakage
True NDH-2 is a single-subunit enzyme, so any organism showing >2 "NDH-2" hits likely has Complex I subunits leaking through the filter. This misclassification inflates the "organisms with NDH-2" count (reported as 10/14) and could bias the compensation test. If pseudo3, pseudo5, and Miya are reclassified as lacking NDH-2, the group sizes shift from 10/4 to 7/7, and the mean deficit difference shrinks substantially. The REPORT acknowledges this limitation explicitly (noting pseudo3_N2E3 as likely false positive), which is good scientific practice, but the analysis was not corrected.
Stoichiometry analysis (NB03, cell 3): The NADH yields are manually entered as a pd.DataFrame literal based on textbook biochemistry, not computed from the FBA model's reaction network. While the FBA reaction data IS queried (cell 6, 1,421 NADH reactions showing rxn10122 carries ~8.38 flux units), it's used only descriptively — the actual stoichiometry comparison uses the hardcoded values. The REPORT acknowledges this ("uses theoretical pathway biochemistry, not measured flux distributions"). The manually entered values appear biochemically reasonable (quinate → 4 NADH via succinyl-CoA + acetyl-CoA TCA turns; glucose → 9 NADH via Entner-Doudoroff + 2 TCA turns) but are not independently validated against the model's per-condition flux predictions.
Complex I FBA discrepancy (NB02, cells 3–5): nuoA (ACIAD0730) shows reactions: None and minimal_media_flux: None in the gene-level query, yet the narrative states "FBA routes all NADH through Complex I." NB03 cell 6 resolves this: rxn10122 (NADH:ubiquinone oxidoreductase, the Complex I reaction) carries all NADH oxidation flux (~8.38 units). The discrepancy arises because the FBA model tracks Complex I flux at the reaction level (rxn10122) but doesn't map this reaction back to individual nuo subunit genes. This is understandable but would benefit from explicit explanation in NB02 so readers don't misinterpret the apparent contradiction.
Statistical methods: The Mann-Whitney U test in NB04 (cell 9) is appropriate for comparing two small groups with potentially non-normal distributions. The test correctly uses alternative='two-sided'. The result (p=0.24) is honestly reported as non-significant.
Notebook organization: All four notebooks follow a clean structure: markdown header with goal/inputs/outputs, setup cell, numbered sections, and summary cell. Data is saved to data/ and figures to figures/ at the end. NB02 explicitly closes the database connection. Good practice throughout.
Pitfall awareness: The project correctly handles the Fitness Browser all-string column pitfall (CAST AS FLOAT), uses orgId filters on genefitness, and separates Spark-dependent (NB04) from local notebooks (NB01–03). The get_spark_session() call in NB04 uses the correct JupyterHub pattern (no import needed). No issues with the pitfalls documented in docs/pitfalls.md.
Findings Assessment
Condition-specific respiratory wiring (Finding 1): Well-supported by the data in NB01. The heatmap and subsystem profiles clearly show distinct respiratory configurations per carbon source. The wiring summary correctly identifies which subsystems are required vs dispensable per condition. Two threshold choices are made without justification: cell 11 uses < 0.8 for "defect" and > 1.0 for "fine" in the subsystem summary, while cell 15 uses < 0.6 for "essential" in the wiring model. The different thresholds produce somewhat different pictures (e.g., Complex I shows "defect" on asparagine at 0.59 by cell 11's threshold but is not classified as "essential" for asparagine in the wiring model). The thresholds are reasonable but the rationale is not discussed.
One additional concern: the Complex II (SDH) subsystem profile is based on a single gene with growth data (ACIAD2880/sdhA) out of 5 SDH genes in the inventory (sdhB, sdhC, sdhD, sdhE all lack data). The profile for this subsystem is thus based on n=1, making it less robust than Complex I (n=11) or Cyt bo3 (n=4). The notebook reports the n values in the subsystem table but doesn't flag the n=1 issue for interpretation.
NDH-2 compensation model (Finding 2): This is the weakest finding because NDH-2 has no growth data, so the central prediction (NDH-2 compensates on glucose) cannot be directly tested. The indirect evidence is presented honestly: FBA routing, genomic context, pangenome conservation status, and ortholog-transferred fitness. The REPORT correctly lists this as the primary limitation. The ortholog-transferred fitness data (30 conditions, mean fitness -0.136) shows mild NDH-2 defects across conditions, but notably has no aromatic conditions in the transferred data — this limits the ability to test the aromatic-specific hypothesis. NB02 mentions this in passing but the REPORT doesn't emphasize it as a specific gap.
Rate vs yield resolution (Finding 3): This is the project's most novel contribution, but it rests on qualitative reasoning rather than quantitative evidence. The argument is: quinate produces fewer NADH (4) than glucose (9) but Complex I is more essential on quinate; therefore, the explanation must be NADH flux rate (concentrated TCA burst from ring cleavage) rather than total yield. The biochemical logic is sound:
- Quinate catabolism funnels all carbon through ring cleavage → succinyl-CoA + acetyl-CoA → 2 near-simultaneous TCA turns
- Glucose catabolism distributes NADH production across multiple Entner-Doudoroff steps + TCA
- The "burst" vs "distributed" distinction is a genuine structural property of the pathways
However, there is no measurement or estimate of actual NADH production rates. The FBA model could in principle provide per-condition flux rates, but it predicts zero flux through NDH-2 on all conditions (because FBA optimizes for growth rate and prefers the more ATP-efficient Complex I). The REPORT acknowledges this and suggests measuring NADH/NAD+ ratios experimentally as a future direction.
Cross-species NDH-2 compensation (Finding 4): The direction supports the hypothesis (organisms with NDH-2 show mean aromatic deficit of -0.086 vs -0.505 for those without), but the result is not statistically significant (p=0.24) with only 4 organisms in the "no NDH-2" group. The P. putida outlier (large Complex I deficit despite having NDH-2) is noted and plausibly explained as an active aromatic degrader that overwhelms NDH-2 capacity. The NDH-2 false positive concern (see Code Quality above) means the true group sizes may be more balanced than 10/4, which would further weaken any signal. Importantly, the REPORT is transparent about this limitation.
Limitations: Well acknowledged. The REPORT lists seven specific limitations covering the most important caveats: no NDH-2 growth data, theoretical stoichiometry, small cross-species sample, annotation variability, ACIAD3522 function uncertainty, NDH-2 false positives, and the dropped pangenome analysis. The Future Directions section proposes five concrete, testable follow-up experiments. The project's transparency about its limitations is a notable strength.
Suggestions
-
[Critical] Fix NDH-2 false positives in NB04: Organisms like
pseudo3_N2E3(8 hits) andpseudo5_N2C3_1(8 hits) almost certainly have Complex I subunits leaking through the text-based NDH-2 filter. Either use KO-based identification (K03885 via the FBbesthitkeggtable), apply a maximum hit count filter (true NDH-2 should have 1–2 genes per organism), or manually curate the hits. Re-run the compensation analysis after fixing — the current 10/4 split may actually be closer to 7/7, which would substantially change the mean deficit comparison. -
[Important] Update README Data Sources: The README lists
kbase_ke_pangenomeunder Data Sources for "NDH-2/Complex I co-occurrence," but this analysis was not performed. Either remove this entry or add a note indicating it was planned but not executed (with a pointer to REPORT.md Future Direction #3). The REPORT already documents this gap, so the fix is just aligning the README. -
[Moderate] Tighten the "Other respiratory" category in NB01: The current broad keyword search captures non-respiratory enzymes (transcription factors, drug resistance genes, amino acid catabolism enzymes). Consider either narrowing the SQL search patterns, applying additional KO/EC filtering to the "Other" category, or renaming it to "Other redox/NADH-related enzymes" to more accurately describe what's included. The headline "62 respiratory chain genes" overstates the respiratory inventory by roughly 10–15 genes.
-
[Moderate] Strengthen the rate vs yield argument with FBA flux data: NB03 queries 1,421 NADH reactions from the FBA model but only uses them descriptively. The
gene_phenotypestable contains per-condition flux predictions (230 conditions) that could provide condition-specific NADH production flux estimates. Comparing total NADH flux on glucose vs quinate conditions would give quantitative support to the "concentrated TCA burst" hypothesis, even though the model doesn't distinguish Complex I from NDH-2 routing. -
[Moderate] Note the n=1 limitation for Complex II (SDH) profile: The subsystem profile for Complex II is based on a single gene (sdhA/ACIAD2880) because the other 4 SDH subunits lack growth data. This should be flagged in the notebook or REPORT when interpreting the SDH condition profile. The other core subsystems (Complex I n=11, Cyt bo3 n=4, Cyt bd n=3) have more robust sample sizes.
-
[Minor] Clarify the growth ratio thresholds: NB01 uses two different thresholds without justification:
< 0.8for "defect" (cell 11) and< 0.6for "essential" (cell 15). Consider adding a brief rationale (e.g., based on the distribution of growth ratios across all genes, or correspondence to specific fitness loss magnitudes) and using consistent terminology. -
[Minor] Explain the nuoA FBA discrepancy in NB02: Cell 3 shows nuoA (ACIAD0730) has
reactions: Noneandflux: None, yet the narrative states "FBA routes all NADH through Complex I." A brief note that Complex I flux is tracked at the reaction level (rxn10122, which carries ~8.38 flux units per NB03 cell 6) rather than mapped back to individual nuo subunit genes would prevent reader confusion. -
[Minor] Emphasize the lack of aromatic conditions in NDH-2 ortholog fitness: NB02 cell 9 shows 30 conditions with ortholog-transferred NDH-2 fitness data, but none are aromatic substrates. The REPORT mentions "no aromatic conditions" in passing but doesn't list it as a specific limitation. Since the hypothesis is specifically about aromatic catabolism overwhelming NDH-2 capacity, the absence of aromatic conditions in the transferred data is a notable gap that deserves explicit mention.
This review was generated by an AI system. It should be treated as advisory input, not a definitive assessment.
Visualizations

Nadh Stoichiometry

Ndh2 Vs Complex I

Proteomics Respiratory

Respiratory Chain Heatmap

Respiratory Clustermap

Subsystem Profiles

Wiring Matrix

Wiring Model
Notebooks
01_respiratory_chain_map.ipynb
01 Respiratory Chain Map
View notebook →
02_ndh2_indirect.ipynb
02 Ndh2 Indirect
View notebook →
03_stoichiometry.ipynb
03 Stoichiometry
View notebook →
04_cross_species_respiratory.ipynb
04 Cross Species Respiratory
View notebook →
05_proteomics_expression.ipynb
05 Proteomics Expression
View notebook →