Pangenome Openness Analysis
CompletedResearch Question
Do open pangenomes show different patterns of environmental vs phylogenetic effects compared to closed pangenomes?
Research Plan
Hypothesis
Open pangenomes should show stronger environment effects because:
- High gene turnover suggests ecological adaptation pressure
- Species with more HGT may acquire niche-specific genes from the environment
- Closed pangenomes may be more constrained by phylogenetic inertia
Approach
- Calculate openness metrics using pre-computed pangenome statistics:
- Core fraction:
no_core / no_gene_clusters - Singleton fraction:
no_singleton_gene_clusters / no_gene_clusters -
Openness = 1 - core_fraction (higher = more open)
-
Correlate with ecotype effects from the ecotype_analysis project:
- Environment partial correlation (controlling for phylogeny)
-
Phylogeny partial correlation (controlling for environment)
-
Statistical test: Spearman correlation between openness and effect sizes
Revision History
- v1 (2026-02): Migrated from README.md
Overview
This project tests whether pangenome openness (the ratio of variable to total genes) predicts whether environmental similarity or phylogenetic relatedness drives gene content variation. It builds on the ecotype_analysis project's partial correlation results, correlating them with pre-computed pangenome statistics from the BERDL pangenome table. The null result suggests pangenome structure is independent of eco-phylo dynamics.
Key Findings
No Correlation Found
Analysis of pangenome openness vs environment/phylogeny effects revealed no significant relationship:
| Metric | Spearman rho | p-value |
|---|---|---|
| Openness vs Environment effect | -0.05 | 0.54 |
| Openness vs Phylogeny effect | 0.03 | 0.73 |
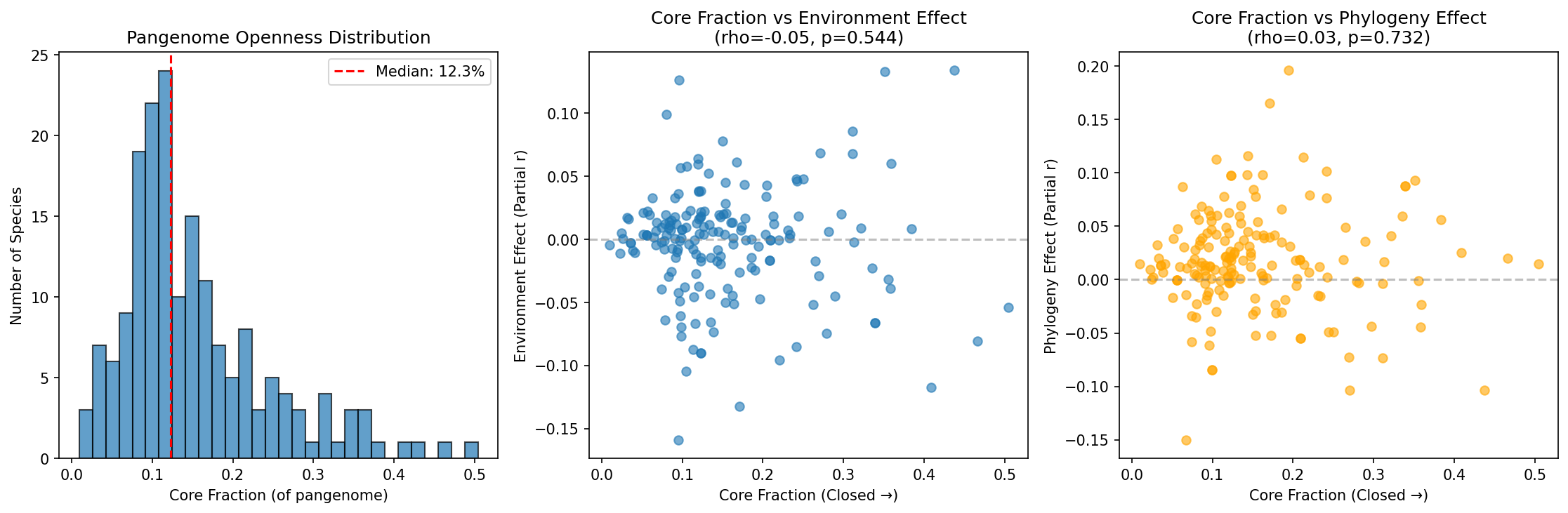
(Notebook: 01_explore_gene_data.ipynb)
Interpretation
Whether a species has an open or closed pangenome does not predict whether environment or phylogeny dominates its gene content. This suggests:
- Pangenome structure is independent of eco-phylo dynamics: Open pangenomes don't necessarily mean environment-driven adaptation
- HGT may not track environmental similarity: Gene acquisition might be opportunistic rather than niche-specific
- Core/accessory classification may not capture functional adaptation: The genes that vary may not be the ones responding to environment
Literature Context
- Tettelin et al. (2005) introduced the open/closed pangenome framework for bacterial genomes. Our null result suggests that the open/closed distinction, while useful for describing gene content variability, does not predict the ecological vs phylogenetic drivers of that variability.
- McInerney et al. (2017) argued that pangenome structure reflects a balance of selection, drift, and HGT. Our finding that openness is uncorrelated with environment effects is consistent with HGT being opportunistic rather than ecologically directed.
- Parks et al. (2022) provided the GTDB taxonomy and pangenome framework used here, enabling consistent cross-species comparisons of pangenome structure.
Limitations
- Sample size is limited to species with both pangenome statistics and ecotype analysis results
- Openness is a single summary metric that may not capture the full complexity of pangenome structure
- Environment and phylogeny effects are derived from partial correlations that may not fully disentangle confounded variables
- The ecotype analysis upstream may have limited statistical power for some species with few genomes
Future Directions
- Stratify by gene function: Do open pangenomes show environment effects specifically in L (mobile) or V (defense) categories?
- Test alternative openness metrics: Try auxiliary fraction, Heap's law alpha, or pangenome fluidity
- Lifestyle interaction: Test openness x lifestyle interaction (e.g., open pathogens vs open environmental species)
Data
Sources
| Dataset | Description | Source |
|---|---|---|
| KBase pangenome statistics | Pre-computed openness metrics per species | pangenome table via Spark |
| Ecotype analysis results | Environment and phylogeny effect sizes | ecotype_analysis project |
Generated Data
| File | Description |
|---|---|
data/pangenome_stats.csv |
Pre-computed pangenome statistics for target species |
data/pangenome_ecotype_merged.csv |
Merged pangenome stats with ecotype analysis results |
data/species_selection_stats.csv |
Species selection statistics |
data/target_genomes_expanded.csv |
Expanded target genome information |
References
- Parks DH et al. (2022). "GTDB: an ongoing census of bacterial and archaeal diversity through a phylogenetically consistent, rank normalized and complete genome-based taxonomy." Nucleic Acids Res 50:D199-D207. PMID: 34520557
- Tettelin H et al. (2005). "Genome analysis of multiple pathogenic isolates of Streptococcus agalactiae: implications for the microbial 'pan-genome'." Proc Natl Acad Sci USA 102:13950-13955. PMID: 16172379
- McInerney JO et al. (2017). "Why prokaryotes have pangenomes." Nat Microbiol 2:17040. PMID: 28350002
Discoveries
Pangenome table has pre-computed stats
January 2026The pangenome table already contains no_core, no_aux_genome, no_singleton_gene_clusters, no_gene_clusters - no need to compute from gene_cluster table.
Tested whether open pangenomes (low core fraction) show different patterns. Results:
- rho=-0.05, p=0.54 for environment effect
- rho=0.03, p=0.73 for phylogeny effect
- No significant relationship found
Data Collections
Visualizations

Pangenome Vs Effects