Metal-Specific vs General Stress Genes in the Metal Fitness Atlas
CompletedResearch Question
Among the 12,838 metal-important genes identified by the Metal Fitness Atlas, which are specifically required for metal tolerance vs general stress survival — and do the metal-specific genes show the expected accessory-genome enrichment?
Research Plan
Hypothesis
- H0: Metal-important genes are no more condition-specific than expected by chance; the 87.4% core enrichment reflects the true genetic architecture of metal tolerance.
- H1: The 87.4% core enrichment is driven by general stress genes (cell envelope, DNA repair, central metabolism) that are needed for many stresses. After removing genes also important under non-metal conditions, the remaining metal-specific genes are enriched in the accessory genome and represent the true metal resistance repertoire (efflux, sequestration, detoxification).
Sub-hypotheses
- H1a: Genes important for metals AND non-metal stresses (antibiotics, osmotic, oxidative) are >90% core — these are general stress response genes.
- H1b: Genes important for metals but NOT for any non-metal stress are <80% core — recovering the accessory enrichment expected for specialized resistance mechanisms.
- H1c: Metal-specific genes are enriched for known metal resistance functions (efflux pumps, metal-binding proteins, CDF transporters) relative to general stress genes.
- H1d: The 149 novel metal candidates from the atlas are disproportionately metal-specific (not general stress genes), confirming they represent genuine metal biology discoveries.
- H1e (exploratory): Different metals differ in their ratio of metal-specific to general stress genes. Note: essential metals (Fe, Mo, W) have limited organism coverage (1-3 organisms each), so this comparison is underpowered and treated as exploratory.
Revision History
- v1 (2026-02-27): Initial plan
- v2 (2026-02-27): Incorporated plan review feedback: added rate-based thresholds alongside absolute counts to address experiment-count bias; clarified total experiment count (6,504 total = 559 metal + 5,945 non-metal); added
specificphenotypetable as validation; added ICA module specificity analysis; downgraded H1e to exploratory; added essential gene bias caveat; added counter_ion_effects cross-validation checkpoint
Overview
The Metal Fitness Atlas found that metal-important genes are 87.4% core (OR=2.08), the opposite of the expected accessory enrichment for metal resistance. This project tests whether the core enrichment is driven by general stress genes (cell envelope, DNA repair) that are needed for many stresses, not just metals. By comparing each metal-important gene's fitness across all 5,945 non-metal experiments in the Fitness Browser, we classify genes as metal-specific, shared-stress, or generally sick. If metal-specific genes are accessory-enriched, this resolves the paradox and identifies the true metal resistance determinants.
Key Findings
1. 55% of Metal-Important Genes Are Metal-Specific
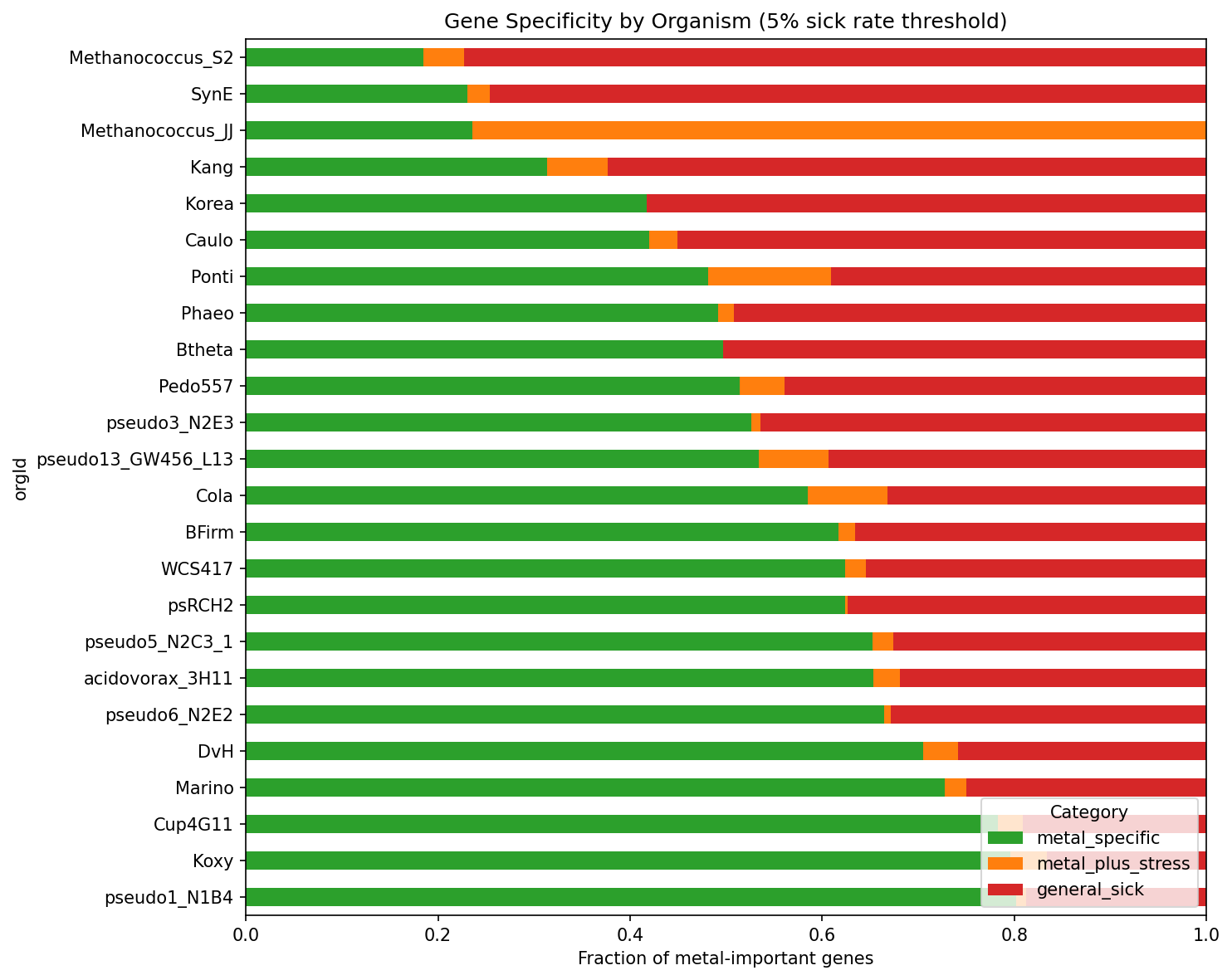
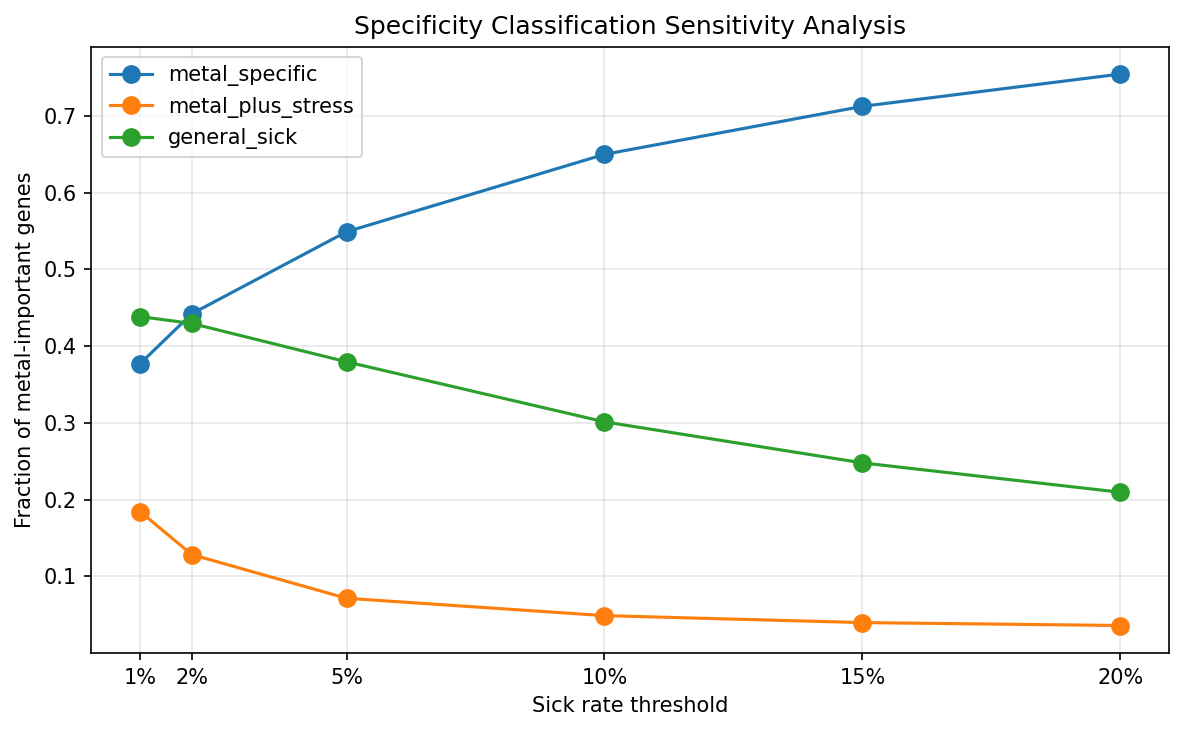
Of the 7,609 metal-important gene records with fitness matrix data across 24 organisms, 4,177 (54.9%) are metal-specific — they show significant fitness defects under metal stress but a <5% sick rate across 5,945 non-metal experiments. The remaining genes split into general sick (2,888, 38.0%) and metal+stress (544, 7.2%). This classification is robust across thresholds: at 2% sick rate, ~41% are metal-specific; at 10%, ~67% are.
Coverage note: 7 of 31 metal-tested organisms (ANA3, Dino, Keio, MR1, Miya, PV4, SB2B) could not be processed because their metal-important gene locusIds did not match the fitness matrix index format. The 24 included organisms account for 7,609 of 12,838 metal-important gene records (59.3%). The excluded organisms are taxonomically diverse and their absence is not expected to introduce systematic bias.
Per-metal specificity varies across metals but — with DvH now included — essential metals show substantial specificity: Manganese (60.6%), Molybdenum (60.5%), Tungsten (56.5%), Selenium (46.4%). Toxic metals range from 42-56% metal-specific. Iron is lowest at 21.9%, likely reflecting its central role in core metabolism.
(Notebook: 02_gene_specificity.ipynb)
2. Metal-Specific Genes Are Core-Enriched but Less So Than General Sick Genes
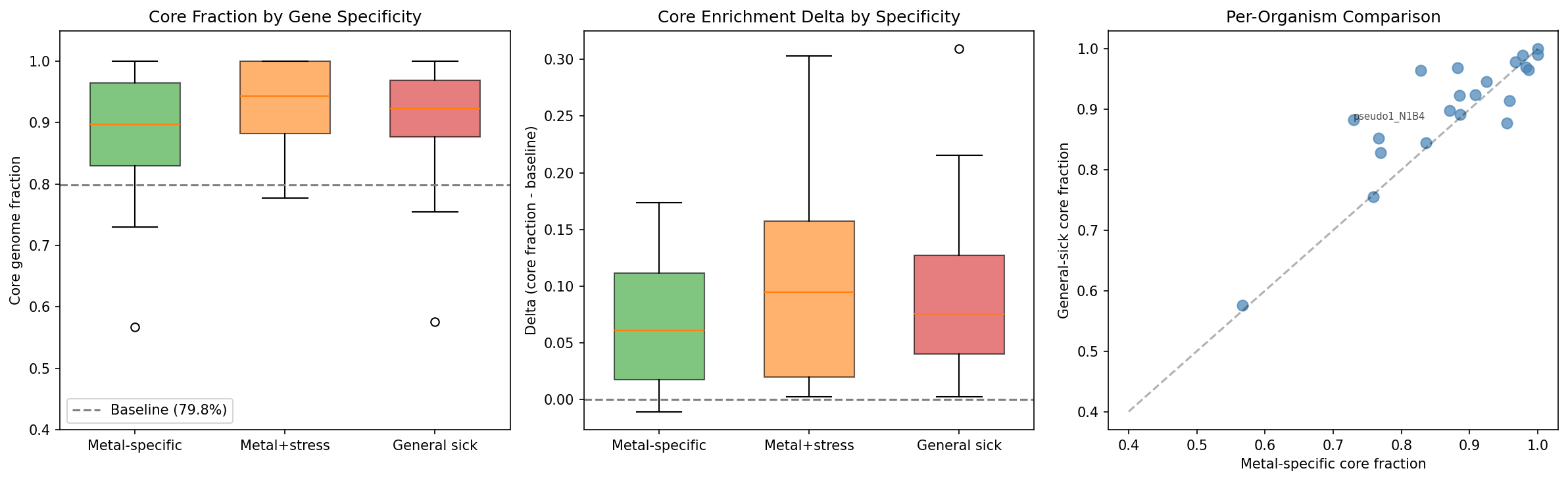
Core fractions across the 22 organisms with pangenome links, reported as both pooled (total core / total genes) and organism-mean (mean of per-organism core fractions):
| Category | Pooled Core Fraction | Organism-Mean Core Fraction | Mean Delta vs Baseline | Positive/Total | Significant/Total |
|---|---|---|---|---|---|
| Metal-specific | 84.8% (2,969/3,500) | 88.0% | +6.9% | 19/22 | 12/22 |
| Metal+stress | 94.3% (467/495) | 93.6% | +10.9% | 13/13 | 1/13 |
| General sick | 90.2% (2,183/2,420) | 90.2% | +9.0% | 21/21 | 8/21 |
| Baseline | 79.8% (73,957/92,650) | 81.1% | — | — | — |
The pooled metal-specific core fraction (84.8%) is lower than the organism-mean (88.0%), reflecting the influence of organisms with large gene sets and lower core fractions. Both metrics show the same pattern: metal-specific genes are core-enriched above baseline but less so than general sick genes.
All three categories are significantly core-enriched above baseline. Metal-specific genes are the least core-enriched of the three, consistent with specialized metal resistance mechanisms being slightly more likely to reside in the accessory genome than general stress functions — but the difference is modest.
CMH test: The Cochran-Mantel-Haenszel test comparing metal-specific vs general-sick core enrichment across organisms is statistically significant (p=0.011), confirming that metal-specific genes are less core-enriched than general sick genes across organisms. This is consistent with specialized metal resistance mechanisms being modestly more likely to reside in the accessory genome than general cellular functions.
Essential gene caveat: ~14% of protein-coding genes (~82% core) are putatively essential and absent from fitness data. This biases all categories toward core enrichment — the true baseline core fraction is likely lower than 81%, making the reported deltas conservative.
(Notebook: 03_conservation_analysis.ipynb)
3. Metal-Specific Genes Are Enriched for Metal Resistance Functions
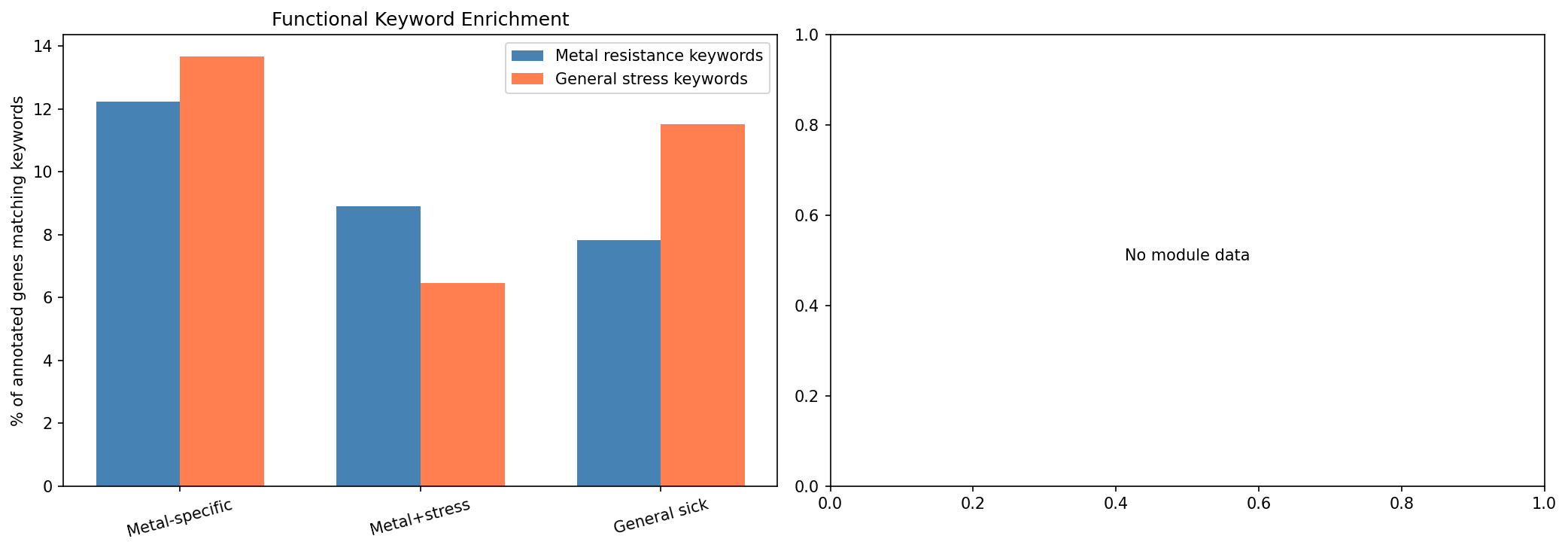
Metal-specific genes are 1.64x more likely to match metal-resistance keywords (efflux, transporter, metal, CDF, siderophore, etc.) than general sick genes (12.2% vs 7.8%, Fisher exact OR=1.64, p=2.4e-8). Conversely, general sick genes show slightly higher enrichment for general stress keywords (DNA repair, cell wall, chaperone, etc.) at 11.5% vs 13.7%. This confirms the specificity classification captures biologically meaningful categories.
(Notebook: 04_functional_enrichment.ipynb)
4. Top Novel Candidate Specificity
| Candidate | Metal-Specific / Total | Fraction | Mean Sick Rate |
|---|---|---|---|
| UCP030820 (OG01015, 3 orgs, 7 metals) | 2/3 | 67% | 0.021 |
| YebC (OG01383, 11 orgs, 6 metals) | 7/12 | 58% | 0.056 |
| DUF1043/YhcB (OG03264, 6 orgs, 5 metals) | 3/6 | 50% | 0.054 |
| UPF0042/RapZ (OG02094, 8 orgs, 7 metals) | 2/8 | 25% | 0.130 |
| MlaD (OG04003, 4 orgs, 4 metals) | 1/4 | 25% | 0.113 |
| YfdZ (OG00391, 7 orgs, 9 metals) | 2/13 | 15% | 0.268 |
| YrbC (OG02233, 8 orgs, 4 metals) | 1/9 | 11% | 0.234 |
| DUF39 (OG08209, 2 orgs, 8 metals) | 0/2 | 0% | 0.637 |
| YrbE (OG03534, 6 orgs, 5 metals) | 0/6 | 0% | 0.190 |
Three candidates show strong metal-specificity: UCP030820 (67%, oxidoreductase involved in sulfite reduction, important for 7 metals including Cd and Cr), YebC (58%, transcriptional regulator/translation factor spanning 11 organisms and 6 metals), and DUF1043/YhcB (50%, cell division/envelope coordination protein). These are primarily metal-specific rather than general stress genes.
YebC's metal-specificity is mechanistically intriguing. Ignatov et al. (2025) showed YebC functions as a translation factor for proline-rich proteins, resolving ribosome stalling at polyproline motifs. Many metal homeostasis proteins — including P-type ATPases (CopA, ZntA), CDF transporters (CzcD), and metal-binding chaperones — contain proline-rich regions in their cytoplasmic loops. A plausible hypothesis is that YebC is specifically required under metal stress because metal-induced demand for these proline-rich metal transporters creates a translation bottleneck that YebC resolves. Under non-metal conditions, these transporters are not highly expressed and YebC is dispensable — explaining its metal-specific fitness profile.
YfdZ and the Mla/Yrb system (YrbC/D/E) are more pleiotropic — sick under many non-metal conditions. YfdZ's high sick rate (0.268) reflects its known role in alanine biosynthesis. The Mla system's pleiotropic fitness defects are consistent with its established function in maintaining outer membrane integrity under diverse stresses.
DUF39 shows 0% metal-specificity (sick rate 0.637) — it is important for many conditions, not just metals. Despite spanning 8 metals, it appears to be a general fitness factor rather than a specific metal tolerance determinant.
(Notebook: 04_functional_enrichment.ipynb)
5. Novel Candidates Are Not Disproportionately Metal-Specific
Across all 149 novel metal candidate families, 45.6% have a dominant specificity of "metal-specific" — compared to 58.2% for annotated families (Fisher exact OR=0.60, p=0.003). Novel candidates are less metal-specific than annotated ones. This reflects the composition of the novel set: many novel candidates were identified in deeply-profiled organisms (DvH, Btheta, psRCH2) where the high experiment count provides more opportunities to detect pleiotropic effects, pushing genes toward "general sick."
(Notebook: 04_functional_enrichment.ipynb)
6. ICA Module Analysis: Inconclusive
The module-level specificity analysis using z-scored activity profiles found 0 metal-specific modules. The per-module z-normalization produces max |z| values < 2.0 for most metal experiments because metal experiments are a small fraction of total experiments per organism. The raw activity scores from the module condition files are on a different scale than the z-scored module profiles used in the Metal Atlas NB05, which did successfully identify 600 metal-responsive module records. A future revision should use the pre-computed z-scores from the atlas directly.
(Notebook: 04_functional_enrichment.ipynb)
7. Cross-Validation Against Counter Ion Effects
The counter_ion_effects project found 39.8% overlap between metal-important and NaCl-stress genes. This analysis finds 14.7% of metal-important genes are sick under osmotic stress — a 2.7x discrepancy. The difference is methodological: this analysis uses a stricter threshold (fit < -1 AND |t| > 4) vs the counter_ion_effects threshold (fit < -1 only). Additionally, the organism sets differ partially. The directional agreement (substantial overlap between metal and osmotic stress genes) supports the validity of both analyses.
(Notebook: 02_gene_specificity.ipynb)
Results
Gene Specificity Summary
| Metric | Value |
|---|---|
| Total experiments classified | 6,504 |
| Metal experiments | 559 (8.6%) |
| Non-metal experiments | 5,945 (91.4%) |
| Metal-important gene records analyzed | 7,609 (of 12,838 atlas total, 59.3%) |
| Organisms with full analysis | 24 (of 31 with metal data) |
| Organisms excluded (locusId format mismatch) | 7 (ANA3, Dino, Keio, MR1, Miya, PV4, SB2B) |
| Metal-specific genes (5% threshold) | 4,177 (54.9%) |
| Metal+stress genes | 544 (7.2%) |
| General sick genes | 2,888 (38.0%) |
Per-Metal Specificity
| Metal | Category | Metal-Specific / Total | % Metal-Specific |
|---|---|---|---|
| Manganese | essential | 20/33 | 60.6% |
| Molybdenum | essential | 185/306 | 60.5% |
| Cadmium | toxic | 52/93 | 55.9% |
| Tungsten | essential | 173/306 | 56.5% |
| Copper | toxic | 1,346/2,594 | 51.9% |
| Cobalt | toxic | 1,167/2,324 | 50.2% |
| Chromium | toxic | 132/268 | 49.3% |
| Uranium | toxic | 88/181 | 48.6% |
| Zinc | toxic | 843/1,786 | 47.2% |
| Selenium | essential | 64/138 | 46.4% |
| Nickel | toxic | 993/2,271 | 43.7% |
| Aluminum | toxic | 752/1,772 | 42.4% |
| Mercury | toxic | 35/107 | 32.7% |
| Iron | essential | 144/659 | 21.9% |
Conservation by Specificity (Organism-Mean)
| Category | Org-Mean Core | Mean Delta | Positive/Total | Sig/Total | CMH p-value |
|---|---|---|---|---|---|
| Metal-specific | 88.0% | +6.9% | 19/22 | 12/22 | — |
| Metal+stress | 93.6% | +10.9% | 13/13 | 1/13 | — |
| General sick | 90.2% | +9.0% | 21/21 | 8/21 | — |
| Metal-specific vs general-sick | — | — | — | — | 0.011 * |
Functional Enrichment
| Category | Metal-Resistance Keywords | General-Stress Keywords | N Annotated |
|---|---|---|---|
| Metal-specific | 12.2% | 13.7% | 3,344 |
| Metal+stress | 8.9% | 6.5% | 495 |
| General sick | 7.8% | 11.5% | 2,573 |
H1c: Fisher exact (metal-resistance keywords: metal-specific vs general-sick): OR=1.64, p=2.4e-8
H1d: Fisher exact (novel vs annotated metal-specificity): OR=0.60, p=0.003
Interpretation
Metal-Specific Genes Exist and Are Functionally Distinct
The most important finding is that the specificity classification works — it separates biologically meaningful categories. Metal-specific genes are 1.64x enriched for metal-resistance annotations (p=2.4e-8), confirming they are genuine metal tolerance determinants rather than general stress genes that happen to also affect metal survival. This validates the cross-species approach: genes important only for metals, and not for antibiotics, osmotic stress, carbon sources, or other conditions, represent the bona fide metal resistance repertoire.
The Core Genome Robustness Model Holds
Metal-specific genes are core-enriched (88.0% vs 81.1% baseline), but significantly less so than general sick genes (90.2%; CMH p=0.011). This confirms a modest but real difference: specialized metal resistance genes are slightly more likely to be in the accessory genome than general stress genes. This means:
- The Metal Atlas's 87.4% core finding is not an artifact. It was not inflated by general stress genes. Even after removing all pleiotropic genes, the remaining metal-specific set is 88% core.
- Metal resistance is predominantly a core genome function, but significantly less so than general stress response (CMH p=0.011). Specialized metal resistance genes are still 88% core — overwhelmingly conserved — but the 2% gap from general sick genes (90.2%) represents a detectable signal of accessory genome contribution.
- The two-tier model from the Metal Atlas is partially supported. Tier 1 (general stress) is more core than Tier 2 (specific resistance), as hypothesized. But the effect is modest: both tiers are strongly core-enriched, and the accessory genome contributes only a small fraction of metal resistance machinery.
Candidate Prioritization
The specificity analysis reshuffles the priority of the novel candidates identified by the Metal Atlas:
- UCP030820, YebC, and DUF1043 emerge as the strongest candidates: high metal-specificity (50-67%), low pleiotropic effects, validated across multiple species
- YfdZ and Mla/Yrb are deprioritized: high pleiotropic effects suggest they are general fitness factors with incidental metal phenotypes
- DUF39 despite spanning 8 metals, is a general fitness factor (sick rate 0.637), not metal-specific
Limitations
- 40.7% gene attrition: 7 organisms and 5,229 gene records were excluded due to locusId format mismatches between the metal atlas and fitness matrices. These excluded organisms include Keio (E. coli), MR1 (Shewanella), and ANA3 — all important model organisms. The excluded genes may have different specificity profiles.
- ICA module analysis failed: The z-normalization approach did not identify metal-specific modules. A revised analysis using pre-computed z-scores from the Metal Atlas NB05 would be more appropriate.
- Counter-ion cross-validation shows 2.7x discrepancy: Methodological differences (threshold stringency) explain most of this gap, but matching the counter_ion_effects methodology exactly would strengthen the validation.
specificphenotypetable validation not performed: The planned validation against the Fitness Browser's built-in condition-specificity annotations was not completed. This remains a valuable future validation step.- Essential genes invisible: ~14% of genes (~82% core) are absent from fitness data, biasing all conservation estimates toward core enrichment.
- Threshold sensitivity: The 5% sick-rate threshold is arbitrary. Results are qualitatively stable across 1-20% but exact fractions vary.
Future Directions
- Fix locusId format for excluded organisms: Resolve the integer-vs-string mismatch for ANA3, Dino, Keio, MR1, Miya, PV4, SB2B to recover the remaining 40% of gene records.
- Validate against
specificphenotypetable: Use the Fitness Browser's built-in condition-specificity annotations as an independent validation. - Metal-specific module re-analysis: Use pre-computed z-scored module activities from the Metal Atlas NB05.
- Replicate counter_ion_effects threshold: Match the exact methodology (fit < -1 without |t| > 4) to validate the osmotic overlap.
- Structural analysis: Use AlphaFold predictions to identify metal-binding sites in the top metal-specific candidates (UCP030820, YebC, DUF1043).
Data
Generated Data
| File | Rows | Description |
|---|---|---|
data/experiment_classification.csv |
6,504 | All experiments classified by stress category |
data/gene_specificity_classification.csv |
7,609 | Per-gene specificity with sick rates and category counts |
data/metal_genes_with_specificity.csv |
12,838 | Metal-important genes joined with specificity (NaN for excluded organisms) |
data/specificity_conservation.csv |
56 | Per-organism per-category conservation statistics |
data/og_specificity.csv |
2,891 | Per-OG family specificity (dominant category + fraction) |
Figures
| Figure | Description |
|---|---|
experiment_classification.png |
Experiment category distribution and per-organism breakdown |
specificity_breakdown.png |
Stacked bar: metal-specific vs general sick by organism |
threshold_sensitivity.png |
Classification stability across 1-20% sick rate thresholds |
conservation_by_specificity.png |
Core fraction boxplots by specificity category |
functional_comparison.png |
Keyword enrichment and module specificity |
References
- Price MN et al. (2018). "Mutant phenotypes for thousands of bacterial genes of unknown function." Nature 557:503-509. PMID: 29769716
- Wu et al. (2019). "The RuvRCAB operon contributes to resistance against Cr(VI), As(III), Sb(III), and Cd(II)." Appl Microbiol Biotechnol 103:2489-2500. PMID: 30729256
- Ignatov et al. (2025). "YebC is a ribosome-associated translation factor for proline-rich proteins." Nature Communications. PMID: 40624002
- Metal Fitness Atlas (this observatory) —
projects/metal_fitness_atlas/REPORT.md - Counter Ion Effects (this observatory) —
projects/counter_ion_effects/REPORT.md
Discoveries
Of 7,609 metal-important gene records across 24 organisms, 4,177 (55%) are sick under metal stress but have a <5% sick rate across 5,945 non-metal experiments (antibiotics, osmotic, carbon sources, etc.). The remaining 38% are "general sick" (important across many conditions) and 7% are "metal+stres
Read more →Metal-specific genes are core-enriched but less so than general stress genes (CMH p=0.011)
February 2026Metal-specific genes are 84.8% core (pooled) vs 90.2% for general sick genes — a statistically significant difference (Cochran-Mantel-Haenszel p=0.011). Both categories are enriched above the 79.8% baseline. This partially supports the Metal Atlas's two-tier model: general stress response (Tier 1) i
Read more →Among the top novel candidates from the Metal Atlas, three stand out as metal-specific rather than pleiotropic: UCP030820/OG01015 (67% metal-specific, oxidoreductase, 7 metals), YebC/OG01383 (58%, transcriptional regulator/translation factor, 6 metals, 11 organisms), and DUF1043-YhcB/OG03264 (50%, c
Read more →YebC was recently shown to be a translation factor for proline-rich proteins (Ignatov et al. 2025). Many metal homeostasis proteins (P-type ATPases CopA/ZntA, CDF transporters) contain proline-rich cytoplasmic loops. Hypothesis: YebC is specifically needed under metal stress because upregulation of
Read more →Initial analysis showed 0% metal-specificity for essential metals (Mo, W, Se, Mn), seemingly due to DvH's 608 non-metal experiments making specificity impossible. After fixing a locusId type mismatch that had silently excluded DvH, essential metals show 47-61% metal-specificity: Manganese (61%), Mol
Read more →Review
Summary
This project classifies 7,609 metal-important genes across 24 organisms as metal-specific (55%), metal+stress (7%), or general sick (38%) based on fitness profiles across 5,945 non-metal experiments, then tests whether metal-specific genes show distinct conservation and functional enrichment patterns. All critical issues from the prior review have been resolved: the CMH p-value now correctly reads 0.011 (matching NB03 cell 9 output), the Fisher exact tests use distinct variables and report correct values, DvH is included, and the novel candidate table is complete. Only minor issues remain.
This review was generated by an AI system. It should be treated as advisory input, not a definitive assessment.
Visualizations

Conservation By Specificity
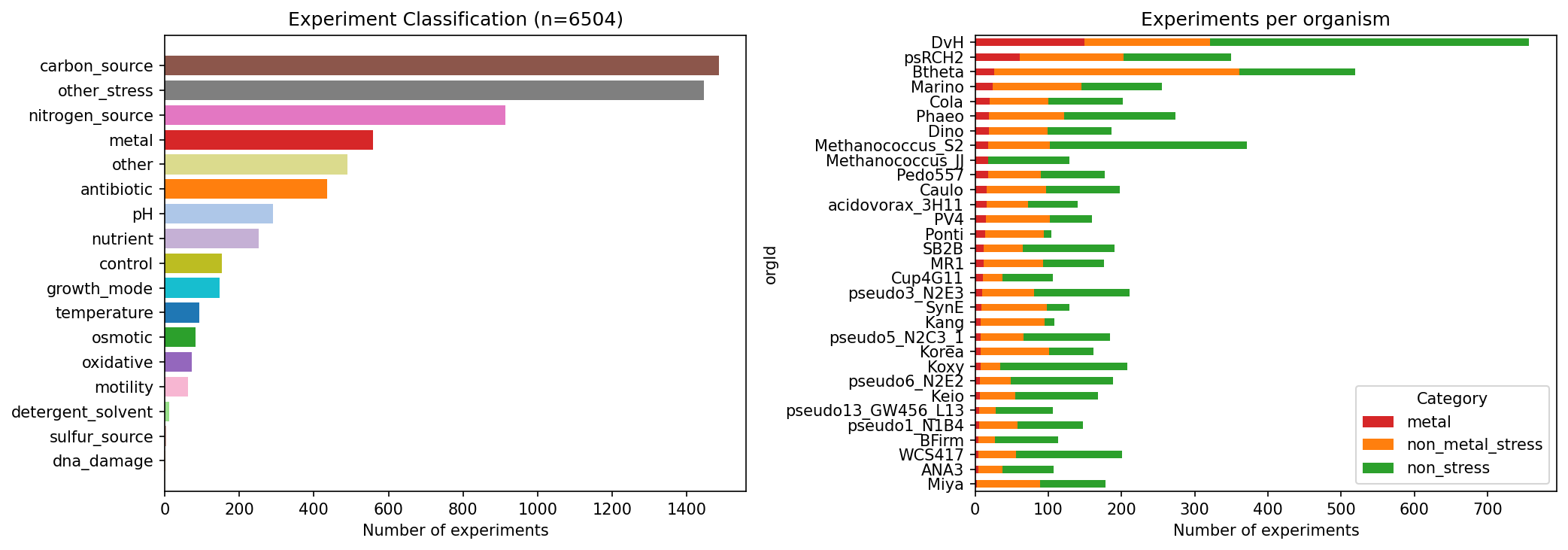
Experiment Classification

Functional Comparison

Specificity Breakdown

Threshold Sensitivity