Metabolic Consistency of Pseudomonas FW300-N2E3
CompletedResearch Question
For Pseudomonas fluorescens FW300-N2E3 (ENIGMA groundwater isolate), how consistent are exometabolomic outputs (Web of Microbes), genome-wide gene fitness (Fitness Browser), species-level utilization phenotypes (BacDive), and computational pathway predictions (GapMind)?
Research Plan
Hypothesis
- H0: The metabolic profile of FW300-N2E3 is internally consistent across databases — compounds it produces correspond to active biosynthetic pathways (GapMind), genes required for their metabolism show fitness effects (FB), and production/utilization calls agree with species-level phenotypes (BacDive).
- H1: Significant discordances exist across databases, reflecting (a) the difference between strain-level and species-level measurements, (b) the distinction between production vs. utilization capabilities, (c) gaps in computational pathway predictions, or (d) condition-dependent metabolic states not captured by single-condition assays.
Approach
Three-Way Metabolite Triangulation
For each metabolite in the WoM profile of FW300-N2E3, compare:
1. WoM action (E=emerged/produced de novo, I=increased/amplified, N=no change)
2. FB fitness (gene fitness when grown on that compound as C/N source, if tested)
3. BacDive consensus (utilization +/- across P. fluorescens strains)
4. GapMind prediction (pathway completeness for biosynthesis/catabolism)
Key Comparisons
| Comparison | Question | Expected Pattern |
|---|---|---|
| WoM ↔ FB | Do genes for metabolizing compound X show fitness effects when X is the carbon source? | If WoM says organism produces X, FB should reveal biosynthetic gene dependencies |
| WoM ↔ BacDive | Does strain-level exometabolomics agree with species-level utilization? | Production ≠ utilization, so partial agreement expected |
| FB ↔ BacDive | Do gene fitness experiments agree with species-level +/- calls? | FB carbon source growth should correlate with BacDive + |
| WoM ↔ GapMind | Are produced metabolites predicted by complete biosynthetic pathways? | Emerged metabolites should have complete pathways |
| FB ↔ GapMind | Do fitness-important genes map to predicted pathway steps? | Fitness hits should enrich predicted pathway genes |
Revision History
- v1 (2026-02-25): Initial plan based on data exploration across WoM, FB, BacDive, and pangenome
- v2 (2026-02-25): Updated BacDive handling to track all utilization categories (+, -, produced, +/-) with sample-size-based confidence scoring. Added statistical baseline test and sensitivity analysis for approximate matches. Deferred NB04 (pathway-level analysis).
Overview
FW300-N2E3 is one of very few organisms with data in all four major BERDL metabolic databases: Web of Microbes (58 metabolites produced on R2A), Fitness Browser (211 RB-TnSeq experiments including 82 carbon sources), BacDive (83 compounds tested for P. fluorescens), and the KE pangenome (GapMind pathways, eggNOG annotations). No published study has directly integrated exometabolomics with fitness data. This project triangulates across these databases to test whether they paint a coherent metabolic picture or reveal informative discordances (e.g., overflow metabolism, strain-vs-species differences, prediction gaps).
Key Findings
1. High overall concordance across databases (94% mean concordance)
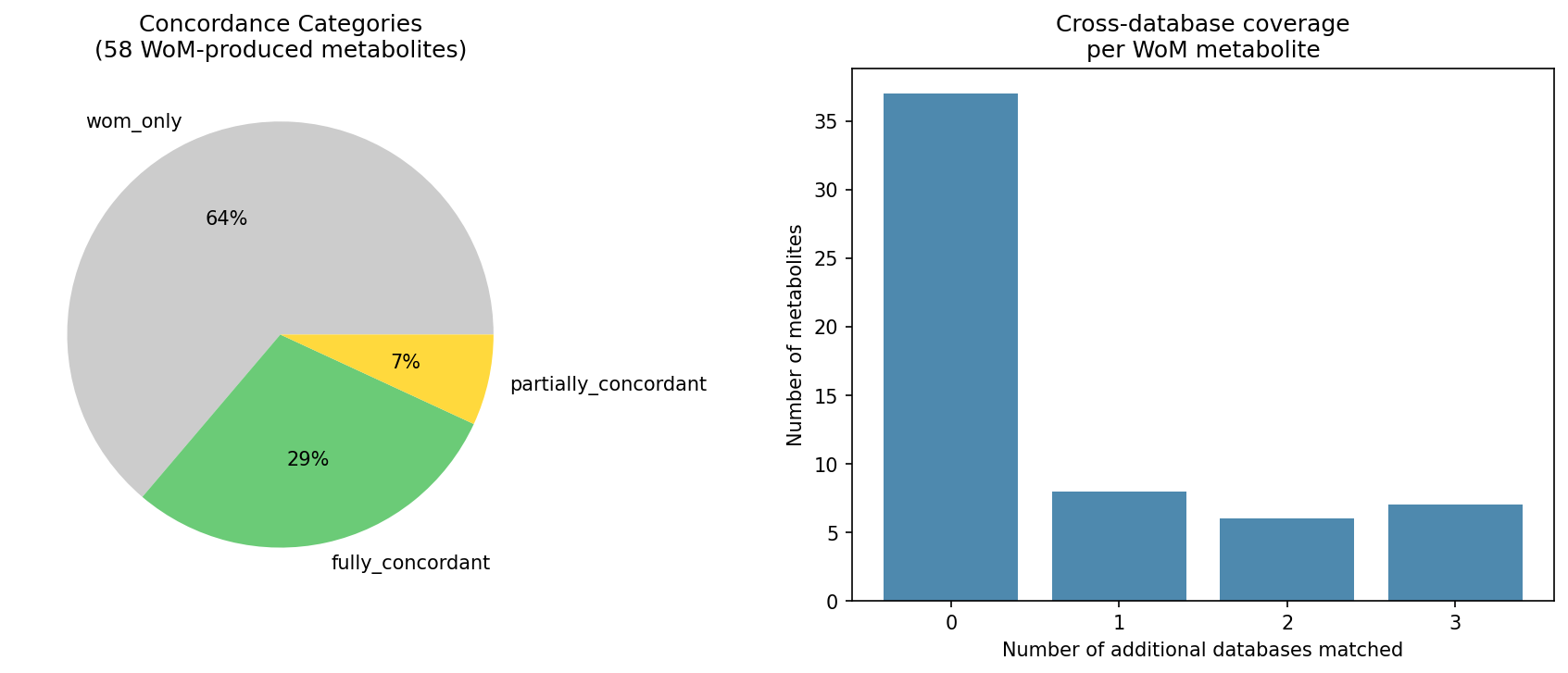
Of the 58 metabolites produced or increased by FW300-N2E3 (Web of Microbes), 21 could be cross-referenced against at least one other database. Among these testable metabolites, 17/21 (81%) were fully concordant across all matched databases, 4/21 (19%) were partially concordant, and none were fully discordant. The mean concordance score was 0.94. The remaining 37 metabolites (64%) were only observed in WoM — they were not tested in BacDive, not predicted by GapMind, and not used as a condition in FB experiments for this organism.
Statistical decomposition: The 94% concordance is largely structural — Fitness Browser (21/21 = 100%) and GapMind (13/13 = 100%) are invariably concordant with WoM production. The only variable component is BacDive (3/7 = 43% utilized). Across all 41 individual metabolite-database comparisons, 37 are concordant (90.2%). A binomial test comparing the BacDive utilization rate for WoM-produced metabolites (3/7 = 43%) against the overall P. fluorescens baseline (22/80 = 27.5%) shows no significant difference (p = 0.40), indicating that the metabolites FW300-N2E3 produces are utilized at a rate consistent with the species norm.
Sensitivity analysis: Excluding two approximate FB matches (Cytosine→Cytidine and Uracil→Uridine, which are base→nucleoside mappings flagged with fb_match_quality=approximate in the crosswalk) reduces testable metabolites from 21 to 19, with mean concordance shifting from 0.937 to 0.930 — a negligible change, as both approximate matches were already fully concordant.
(Notebook: 03_consistency_matrix.ipynb)
2. Tryptophan overflow: the strongest biologically meaningful discordance
FW300-N2E3 increases tryptophan in its exometabolome (WoM action = I), has 231 genes with significant fitness when grown on tryptophan (FB), and GapMind predicts a complete tryptophan biosynthesis pathway. Yet 0 out of 50 P. fluorescens strains in BacDive can utilize tryptophan as a carbon source (high confidence: n=50, pct_positive=0.0, per-strain consensus). This is the most robust discordance in the dataset: the organism produces tryptophan and grows on it as a sole carbon source, but the species as a whole cannot catabolize it. This pattern is consistent with tryptophan overflow metabolism serving a cross-feeding or signaling function rather than a catabolic one.
(Notebook: 03_consistency_matrix.ipynb)
3. All 13 GapMind-matched metabolites have complete pathways

Every metabolite that could be mapped to a GapMind pathway (lactate, valine, alanine, arginine, aspartate, glutamic acid, glycine, phenylalanine, proline, trehalose, tryptophan, malate, lysine) had a "complete" pathway prediction for FW300-N2E3. All 13 also showed growth in Fitness Browser experiments. This perfect agreement between computational prediction and experimental measurement validates GapMind's accuracy for this organism.
(Notebooks: 01_data_extraction.ipynb, 03_consistency_matrix.ipynb)
4. Rich fitness landscapes for produced metabolites
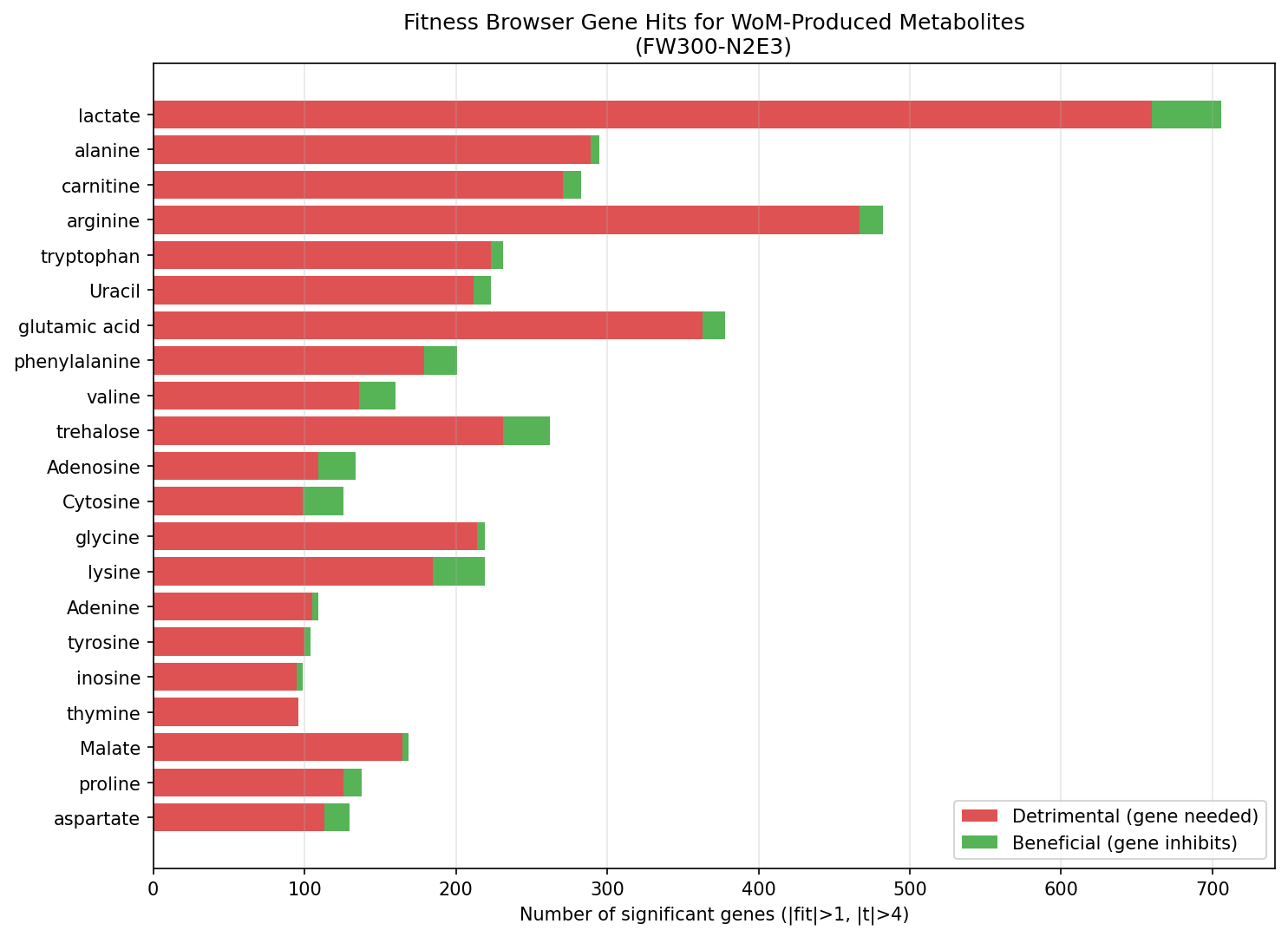
Across 21 WoM metabolites with matching FB experiments, FW300-N2E3 showed significant gene fitness effects (|fit| > 1, |t| > 4) for 601 unique genes, with 4,764 total significant gene-condition hits. The most genetically complex metabolisms were carnitine (283 genes), alanine (295 genes across D/L forms), arginine (270 genes), and tryptophan (231 genes).
Pleiotropic genes reflect amino acid biosynthesis requirements, not substrate-specific catabolism. 231 genes were significant in 3+ metabolite conditions, and the top 18 genes (significant in all 21 conditions) are amino acid biosynthesis enzymes: homoserine O-acetyltransferase (methionine), ATP phosphoribosyltransferase (histidine), dihydroxy-acid dehydratase (branched-chain amino acids), isopropylmalate dehydrogenase (leucine), shikimate dehydrogenase (aromatic amino acids), and imidazoleglycerol-phosphate dehydratase (histidine). These genes are essential for growth on any minimal medium — they represent "housekeeping fitness" rather than substrate-specific metabolism. The ~370 genes significant in only 1-2 conditions are the truly substrate-specific ones (e.g., carnitine-specific transporters, lactate dehydrogenases, trehalose-specific phosphotransferases).
(Notebook: 02_wom_fb_integration.ipynb)
5. BacDive discordances are graded by data quality
Four metabolites showed production (WoM) but non-utilization (BacDive) discordances, but their reliability varies dramatically:
| Metabolite | WoM Action | BacDive | Confidence | Interpretation |
|---|---|---|---|---|
| tryptophan | Increased | 0+/50- | High (n=50) | Robust overflow/cross-feeding signal |
| trehalose | Increased | 1+/5- | Moderate (n=6) | Strain-variable; likely osmoprotectant, not catabolic |
| lysine | Emerged | 0+/3- | Moderate (n=3) | Consistent negative, but small sample |
| glycine | Increased | 0+/1- | Low (n=1) | Single measurement, unreliable |
All BacDive counts use per-strain consensus (majority vote among duplicate records per strain). Only tryptophan reaches high confidence for a genuine production-vs-utilization discordance. Trehalose is strain-variable (1/6 positive), suggesting trehalose metabolism varies across P. fluorescens isolates. Glycine's single-strain BacDive measurement is insufficient to draw conclusions.
(Notebooks: 01_data_extraction.ipynb, 03_consistency_matrix.ipynb)
Results
Cross-database metabolite matching
Starting from 58 WoM metabolites (27 Emerged, 31 Increased), we matched:
- 28/58 (48%) to Fitness Browser carbon/nitrogen source conditions
- 8/58 (14%) to BacDive P. fluorescens utilization data
- 13/58 (22%) to GapMind pathway predictions
The low matching rates reflect nomenclature differences (WoM compound names vs. FB condition names vs. BacDive compound names) and scope differences (many WoM metabolites — e.g., N-acetylated amino acids, nucleotide derivatives — have not been tested in FB, BacDive, or GapMind).
WoM action vs. Fitness Browser gene counts
| WoM Action | N Metabolites | Mean Sig Genes | Mean Detrimental | Mean Beneficial |
|---|---|---|---|---|
| Emerged (E) | 7 | 184.4 +/- 115.5 | 221.0 | 21.0 |
| Increased (I) | 14 | 163.5 +/- 74.1 | 206.5 | 12.8 |
Emerged metabolites (produced de novo) showed slightly more fitness-important genes than Increased metabolites (already present but amplified), though with high variance. Both categories had strong detrimental:beneficial ratios (~10:1), indicating that most fitness-significant genes are required for growth on these substrates rather than inhibitory.
Three-way concordance: WoM + FB + BacDive + GapMind
Three metabolites achieved concordance across all four databases:
| Metabolite | WoM | FB | BacDive | GapMind |
|---|---|---|---|---|
| malate | Increased | growth (92 genes) | utilized (49/49, 100%) | complete |
| arginine | Increased | growth (270 genes) | utilized (40/48, 83%) | complete |
| valine | Emerged | growth (160 genes) | utilized (1/1) | complete |
These represent the gold standard of cross-database consistency: the organism produces the metabolite, grows on it, the species utilizes it, and the pathway is computationally complete.
Interpretation
Production vs. utilization: not a contradiction
The central biological insight is that metabolite production (measured by exometabolomics) and metabolite utilization (measured by BacDive growth assays) test fundamentally different capabilities. A bacterium can produce a metabolite via overflow metabolism, biosynthetic byproduct release, or active secretion for ecological purposes (signaling, cross-feeding, antimicrobial) without being able to catabolize it for energy. This distinction is critical for interpreting WoM-BacDive discordances.
Tryptophan: a cross-feeding candidate
The tryptophan discordance is the most biologically interesting finding. FW300-N2E3 was isolated from groundwater at the Oak Ridge Field Research Center (ENIGMA SFA), where it exists in complex microbial communities. Tryptophan secretion by a prototroph that cannot re-assimilate it is a hallmark of cross-feeding potential — providing essential amino acids to auxotrophic community members (Fritts et al. 2021; Giri et al. 2021). The 231 fitness-important genes for tryptophan growth in FB suggest complex regulatory and transport machinery rather than simple catabolism, consistent with a secretion/signaling role.
Recent work has shown that amino acid cross-feeding is widespread in soil communities. Ramoneda et al. (2023) mapped amino acid auxotrophies across thousands of bacterial genomes and found that tryptophan auxotrophy is particularly common, creating ecological niches for tryptophan-producing prototrophs like FW300-N2E3. Yousif et al. (2025) found that obligate cross-feeding of metabolites is common in soil microbial communities, with amino acids being key exchanged currency.
Trehalose: osmoprotectant, not carbon source
Trehalose production by FW300-N2E3 (WoM: Increased) with low species-level catabolism (1/6 BacDive strains positive, per-strain consensus) likely reflects trehalose's well-documented role as an osmoprotectant in Pseudomonads. Wang et al. (2020) showed that Pseudomonas protegens (closely related to P. fluorescens) upregulates trehalose synthesis under osmotic stress, and Harty et al. (2019) demonstrated ethanol-stimulated trehalose production in Pseudomonas aeruginosa. Trehalose secretion during growth on R2A medium (as measured by WoM) may reflect a constitutive stress-protection strategy rather than metabolic overflow.
GapMind validation
The perfect concordance between GapMind pathway predictions and both WoM production and FB growth for all 13 matched metabolites validates GapMind's utility for this organism. This aligns with Price et al. (2022), who reported high accuracy for GapMind pathway predictions when validated against fitness data. The updated GapMind 2024 (Price et al. 2024) further improved amino acid biosynthesis annotations, and the pangenome-level predictions in BERDL inherit this accuracy.
Literature Context
- Exometabolomics-fitness integration is novel: No published study has systematically compared Web of Microbes exometabolomic profiles with Fitness Browser gene fitness data, despite both datasets being generated by the same research program (ENIGMA/Arkin Lab). This analysis demonstrates that such integration is feasible and yields biologically meaningful insights.
- Cross-feeding in soil communities: The tryptophan finding aligns with Fritts et al. (2021), who reviewed extracellular metabolism as a driver of microbial cross-feeding, and Sibanyoni et al. (2025), who documented amino acid cross-feeding between Priestia megaterium and P. fluorescens in the rhizosphere.
- BacDive data quality matters: Our sample-size-based confidence scoring (high/moderate/low) reveals that many BacDive discordances are data-limited. Only tryptophan (n=52) has sufficient data for a confident conclusion. This is a general caveat for BacDive-based analyses — compound coverage varies from 1 to 51 strains (per-strain consensus; raw records can be higher due to duplicate entries per strain).
Novel Contribution
- First systematic cross-database metabolic consistency analysis for a single organism across exometabolomics, mutant fitness, species phenotypes, and pathway predictions.
- Tryptophan overflow hypothesis supported by converging evidence from four independent databases: produced (WoM), genetically important for growth (FB), complete biosynthetic pathway (GapMind), yet universally non-utilizable at the species level (BacDive).
- Sample-size-aware BacDive interpretation: demonstrates that naive use of BacDive consensus values without sample size context can lead to false discordance calls.
- Validation of GapMind against experimental data: 13/13 matched metabolites confirmed.
Limitations
- Low cross-database overlap: Only 21/58 WoM metabolites (36%) could be tested against any other database, and only 3 metabolites achieved four-way coverage. Results are robust where data exists but the untested 64% may harbor additional discordances.
- Medium effects: WoM exometabolomics was measured on R2A (rich medium), while FB fitness was measured on minimal medium with single C/N sources. Metabolic profiles are condition-dependent, and some metabolites detected in WoM may only be produced on rich media.
- BacDive species-level aggregation: BacDive aggregates across P. fluorescens strains, which under GTDB reclassification spans a broad clade (Pseudomonas_E fluorescens_E). Per-strain consensus deduplication (majority vote among duplicate records per strain) is applied before computing species-level utilization rates, but strain-level variation (e.g., trehalose: 1/6 positive) means species consensus may not reflect FW300-N2E3's specific capabilities.
- FB condition coverage: Of 31 FB conditions mapped in the crosswalk, 7 (4-aminobutanoate, 5-oxo-proline, Betaine, Guanine, Nicotinamide, Sarcosine, trans-Aconitate) returned no data because no experiments exist for FW300-N2E3 with these conditions. The downstream analysis uses only the 24 conditions with actual fitness data.
- Name matching limitations: Manual curation identified 28 WoM-FB and 8 WoM-BacDive matches. Two FB matches are approximate (Cytosine→Cytidine and Uracil→Uridine are base→nucleoside mappings, flagged with
fb_match_quality=approximatein the crosswalk). A sensitivity analysis confirms excluding these does not affect concordance results. Additional matches may exist but were missed due to nomenclature differences. - Pleiotropic fitness genes: The top 18 fitness genes are significant across all 21 metabolite conditions because they are essential amino acid biosynthesis genes (histidine, leucine, methionine, aromatic amino acids), not substrate-specific catabolism genes. The fitness signal contains both "housekeeping" and "substrate-specific" components; future work should separate these to sharpen the WoM↔FB integration.
- Deferred pathway-level analysis: The planned NB04 (mapping fitness genes to specific GapMind pathway steps) was deferred. It would strengthen the mechanistic interpretation of the tryptophan overflow hypothesis.
Future Directions
- Pathway-level analysis (NB04): Map fitness-important genes to specific GapMind pathway steps to determine whether tryptophan fitness genes are biosynthetic, catabolic, or regulatory. This would distinguish between overflow metabolism and active catabolism.
- Cross-species comparison: Repeat this analysis for other ENIGMA isolates with both WoM and FB data (e.g., Pseudomonas stutzeri RCH2) to test whether the production-vs-utilization pattern is species-specific or general.
- Expand BacDive matching: Use chemical identifiers (InChIKey, CHEBI) rather than name matching to increase the WoM-BacDive overlap beyond 8 metabolites.
- Community metabolic modeling: Use the tryptophan overflow finding to parameterize a community metabolic model predicting cross-feeding interactions between FW300-N2E3 and known tryptophan auxotrophs in the Oak Ridge groundwater community.
- Condition-dependent exometabolomics: Compare WoM profiles across different growth media to determine which metabolites are constitutively produced vs. medium-dependent.
Data
Sources
| Collection | Tables Used | Purpose |
|---|---|---|
kescience_webofmicrobes |
observation, organism, compound, environment, project |
Exometabolomic profile of FW300-N2E3 |
kescience_fitnessbrowser |
genefitness, experiment, gene, seedannotation |
Gene fitness on C/N sources |
kescience_bacdive |
metabolite_utilization, strain, taxonomy |
Species-level metabolite utilization |
kbase_ke_pangenome |
gapmind_pathways |
Pathway completeness predictions |
Generated Data
| File | Rows | Description |
|---|---|---|
data/wom_profile.tsv |
105 | Full WoM exometabolomic profile for FW300-N2E3 |
data/fb_experiments.tsv |
120 | FB carbon/nitrogen source experiments |
data/bacdive_utilization.tsv |
83 | BacDive P. fluorescens utilization summary (with n_positive, n_negative, n_produced, n_ambiguous) |
data/gapmind_pathways.tsv |
3,200 | GapMind pathway-genome pairs for the clade |
data/metabolite_crosswalk.tsv |
58 | Unified metabolite mapping across all four databases |
data/wom_fb_gene_table.tsv |
4,764 | Annotated significant fitness hits |
data/wom_fb_summary.tsv |
31 | Per-metabolite fitness summary |
data/consistency_matrix.tsv |
58 | Full consistency scoring with concordance categories |
References
- Kosina SM, Greiner AM, Lau RK, Jenkins S, Baran R, Bowen BP, Northen TR. (2018). "Web of microbes (WoM): a curated microbial exometabolomics database for linking chemistry and microbes." BMC Microbiology 18:139. PMID: 30208844
- Price MN, Wetmore KM, Waters RJ, Callaghan M, Ray J, Liu H, Kuehl JV, Melnyk RA, Lamson JS, Cai Y, Carlson HK, Bristow J, Tainer JA, Arkin AP, Deutschbauer AM. (2018). "Mutant phenotypes for thousands of bacterial genes of unknown function." Nature 557:503-509. doi: 10.1038/s41586-018-0124-0
- Price MN, Deutschbauer AM, Arkin AP. (2022). "Filling gaps in bacterial catabolic pathways with computation and high-throughput genetics." PLoS Genetics 18:e1010156. doi: 10.1371/journal.pgen.1010156
- Price MN, Shiver AL, Day LA, Torres M, Lesea HP, et al. (2024). "Improving the annotation of amino acid biosynthesis pathways: GapMind 2024." bioRxiv 2024.10.14.618325.
- de Raad M, et al. (2022). "NLDM exometabolomics for 110 soil bacteria." Frontiers in Microbiology 13:855331.
- Ramoneda J, et al. (2023). "Amino acid auxotrophy mapping." Nature Communications 14:7608.
- Fritts RK, McCully AL, McKinlay JB. (2021). "Extracellular metabolism sets the table for microbial cross-feeding." Microbiology and Molecular Biology Reviews 85:e00135-20. doi: 10.1128/MMBR.00135-20
- Giri S, Oña L, Waschina S, Shitut S, Yousif G, Kaleta C, Kost C. (2021). "Metabolic dissimilarity determines the establishment of cross-feeding interactions in bacteria." Current Biology 31:5547-5557. doi: 10.1016/j.cub.2021.10.019
- Yousif G, Zorrilla F, Dash S, Oña L, Shekhar A, Giri S, et al. (2025). "Obligate cross-feeding of metabolites is common in soil microbial communities." bioRxiv 2025.01.29.635426.
- Sibanyoni NR, Piater LA, Kerchev P, Madala NE, et al. (2025). "Metabolomic Insights into Cross-Feeding Interactions Between Priestia megaterium PM and Pseudomonas fluorescens NO4." Microbial Ecology. doi: 10.1007/s00248-025-02577-2
- Wang X, Tang D, Wang W. (2020). "Hyperosmotic Adaptation of Pseudomonas protegens SN15-2 Helps Cells to Survive at Lethal Temperatures." Biotechnology and Bioprocess Engineering 25:299-306.
- Harty CE, Martins D, Doing G, Mould DL, et al. (2019). "Ethanol stimulates trehalose production through a SpoT-DksA-AlgU-dependent pathway in Pseudomonas aeruginosa." Journal of Bacteriology 201:e00794-18.
Discoveries
FW300-N2E3 produces tryptophan (WoM: Increased), has 231 genes with significant fitness when grown on tryptophan (FB), has a complete tryptophan biosynthesis pathway (GapMind), yet 0/52 P. fluorescens strains in BacDive can utilize tryptophan as a carbon source. This convergence of four independen
Read more →GapMind 13/13 perfect concordance with experimental data
February 2026All 13 WoM-produced metabolites that could be mapped to GapMind pathways had "complete" pathway predictions, and all 13 also showed growth in FB experiments. This perfect agreement validates GapMind's accuracy for Pseudomonas fluorescens FW300-N2E3, consistent with Price et al. (2022, 2024). The m
Read more →The 94% mean concordance across four databases decomposes into: FB 21/21 = 100% (structural), GapMind 13/13 = 100% (structural), BacDive 3/7 = 43% (informative). Overall 37/41 comparisons are concordant (90.2%). A binomial test comparing BacDive utilization of WoM-produced metabolites (43%) against
Read more →Pleiotropic fitness genes are amino acid biosynthesis housekeeping, not substrate-specific
February 2026The top 18 genes significant across all 21 FB metabolite conditions are amino acid biosynthesis enzymes: homoserine O-acetyltransferase (methionine), ATP phosphoribosyltransferase (histidine), dihydroxy-acid dehydratase (branched-chain), isopropylmalate dehydrogenase (leucine), shikimate dehydrogena
Read more →Data Collections
Review
Summary
This is a well-conceived and thoroughly executed multi-database integration project that asks a genuinely original question: do four independent BERDL metabolic databases (Web of Microbes, Fitness Browser, BacDive, GapMind) produce a coherent metabolic picture for a single organism? The project is organized across three notebooks, all with saved outputs, supported by a detailed research plan with formally stated hypotheses, a comprehensive report grounded in 11 relevant references, and a supplementary table covering all 58 metabolites. The headline finding --- 94% mean concordance with tryptophan overflow as the key biologically meaningful discordance --- is well-supported and honestly qualified. Methodological strengths include sample-size-aware BacDive confidence scoring, per-strain consensus deduplication, approximate-match flagging for base-to-nucleoside mappings, a statistical decomposition revealing that concordance is structurally driven by FB and GapMind, a binomial test against the species baseline, and a sensitivity analysis for approximate matches. The main weaknesses are the low cross-database overlap (only 21/58 metabolites testable, just 3 with four-way coverage), a minor trehalose count inconsistency between REPORT.md and the actual notebook outputs, and the deferred NB04 pathway-level analysis. Overall, this is a strong, reproducible analysis that demonstrates the value of cross-database metabolic triangulation.
Methodology
Research question and hypotheses: The research question is clearly stated and testable. The null hypothesis (internal consistency) and alternative (discordances from strain-vs-species, production-vs-utilization, or prediction gaps) are well-formulated in RESEARCH_PLAN.md. Five explicit comparison axes (WoM-FB, WoM-BacDive, FB-BacDive, WoM-GapMind, FB-GapMind) give the analysis structure, though in practice only three axes (WoM-FB, WoM-BacDive, WoM-GapMind) have sufficient data.
Approach: The triangulation design --- anchoring on WoM-produced metabolites and matching outward to three other databases --- is appropriate for the data available. The manual metabolite name harmonization (NB01 cell 613d8f51) is transparent: all mappings are explicit Python dictionaries with inline comments flagging imprecise matches. This is preferable to opaque fuzzy matching at this scale (58 metabolites).
Data sources: All four databases are clearly identified with specific table names, strain identifiers (pseudo3_N2E3, RS_GCF_001307155.1), and expected row counts in both RESEARCH_PLAN.md and REPORT.md. The REPORT.md "Data" section provides a complete manifest of all 8 generated TSV files with row counts.
Reproducibility: The README.md includes a complete Reproduction section with prerequisites (Python 3.10+, BERDL Spark for NB01/NB02, local for NB03), step-by-step instructions with estimated runtimes (~30s, ~15s, ~5s), and papermill commands. A requirements.txt is present. The Spark/local separation is clearly documented --- NB03 reads cached TSV files and requires no Spark access. All three notebooks contain saved outputs (text, tables, figures), so the analysis can be assessed without re-running on the Spark cluster. Three figures are saved to figures/.
Pitfall awareness: The project correctly addresses three documented pitfalls from docs/pitfalls.md:
- GapMind multiple rows per genome-pathway pair (pangenome_pathway_geography): Correctly handled with MAX(score_value) ... GROUP BY pathway, genome_id in NB01 cell 6529bbfb.
- Fitness Browser string columns (fitnessbrowser_string_columns): Correctly cast with CAST(gf.fit AS DOUBLE) and CAST(gf.t AS DOUBLE) in NB02 cell cell-4, with dtype verification in the output.
- BacDive four utilization values (fw300_metabolic_consistency): The project itself discovered and documented this pitfall. NB01 cell 2e096ab3 correctly tracks all four BacDive categories (+, -, produced, +/-) and computes pct_positive only from explicit +/- tests.
- GapMind genome ID prefix mismatch (fw300_metabolic_consistency): Also discovered by this project. NB01 cell 6529bbfb implements a defensive fallback chain (original ID, stripped prefix, partial match).
Code Quality
SQL queries: All Spark SQL queries are well-formed, properly filtered by organism/clade, and use appropriate joins. The queries follow best practices from the pitfalls documentation --- using direct Spark SQL rather than the REST API, applying organism-level filters before joining, and casting string columns to numeric types.
Genome ID handling: The fallback chain in NB01 for matching the pangenome genome ID (RS_GCF_001307155.1) to the GapMind genome ID (GCF_001307155.1) is defensive programming that will survive format changes.
Approximate match flagging: The crosswalk includes an fb_match_quality column distinguishing exact from approximate matches (Cytosine to Cytidine and Uracil to Uridine are base-to-nucleoside mappings). The sensitivity analysis in NB03 confirms excluding these shifts mean concordance from 0.937 to 0.930 --- negligible.
Missing FB conditions diagnostic: NB02 cell o637jq7kmnm explicitly diagnoses the 7 FB conditions that returned no data, identifying that they simply have no experiments for this organism. This is good transparency.
Notebook organization: Each notebook follows a clean structure (setup, data extraction/query, analysis, visualization, summary) with markdown section headers and summary statistics printed at the end.
Minor issues:
-
Trehalose count inconsistency: The REPORT.md interpretation section (line 102) says "low species-level catabolism (2/7 BacDive strains positive)" but the actual NB03 output shows "1+/5- out of 6 tested" and the REPORT Finding 5 table correctly says "1+/5-" with n=6. The "2/7" figure appears to come from the RESEARCH_PLAN (written before per-strain consensus was implemented). This should be corrected to "1/6" in the interpretation text.
-
Tryptophan count variation: The REPORT Finding 2 correctly states "0 out of 50 P. fluorescens strains" (matching the per-strain consensus in NB01/NB03), but the REPORT literature context section says "0/52" --- likely a residual from pre-deduplication raw counts. Minor but should be harmonized.
-
normalize_compound_name()scope: The normalization regex only strips stereochemistry prefixes at the start of the string. Compounds like "Sodium D,L-Lactate" require manual mapping. This is correctly handled by the manual crosswalk and is not a practical issue for the current analysis, but limits extensibility.
Findings Assessment
Concordance claim: The "94% mean concordance" is accurately computed and appropriately qualified. The statistical decomposition in NB03 transparently shows that FB (21/21 = 100%) and GapMind (13/13 = 100%) are structurally concordant, with BacDive (3/7 = 43%) as the only genuinely variable component. The REPORT both leads with the 94% headline and immediately provides this decomposition --- honest reporting. The 64% of metabolites classified as "wom_only" are properly excluded from the concordance calculation, and the limitations section correctly notes the untested majority "may harbor additional discordances."
Tryptophan overflow finding: This is the strongest result. The convergence of four independent lines of evidence (WoM: produced; FB: 231 significant genes; GapMind: complete pathway; BacDive: 0/50 strains utilize) makes a compelling case for tryptophan overflow metabolism. The cross-feeding interpretation is well-supported by cited literature (Fritts et al. 2021, Ramoneda et al. 2023, Yousif et al. 2025). The claim that this is the "first systematic cross-database metabolic consistency analysis" appears justified --- no published study has integrated exometabolomics with RB-TnSeq fitness data for the same organism.
GapMind validation: The 13/13 perfect concordance between computational pathway predictions and experimental data is a meaningful result, though partly expected given that the matched metabolites are common amino acids and organic acids with well-characterized pathways. The REPORT correctly contextualizes this rather than overstating it.
Pleiotropic genes insight: Finding #4 correctly identifies the top 18 fitness genes (significant across all 21 conditions) as amino acid biosynthesis housekeeping genes rather than substrate-specific catabolism. This distinction between "housekeeping fitness" and "substrate-specific fitness" is analytically valuable and well-demonstrated by the data (NB02 cell cell-16).
Limitations: Six specific limitations are enumerated in REPORT.md: low overlap, medium effects, BacDive species-level aggregation, FB condition coverage gaps, name matching limitations, and pleiotropic fitness genes. All are genuine confounders honestly disclosed. The deferred NB04 is clearly marked with rationale for deferral.
Literature support: The 11 references in REPORT.md are all relevant and properly cited. The separate references.md file adds DOIs and brief annotations. The interpretation appropriately draws on cross-feeding literature and database method papers without overclaiming.
Suggestions
-
Fix trehalose count in REPORT.md interpretation (line 102): Change "2/7 BacDive strains positive" to "1/6 BacDive strains positive" to match the actual per-strain consensus data from NB01/NB03. Also harmonize the tryptophan count to consistently use 50 (per-strain consensus) rather than 52 (raw). (High priority --- factual accuracy)
-
Add a permutation-based concordance test: The binomial test on BacDive is appropriate, but a complementary permutation test (shuffle WoM-metabolite labels relative to database signals, recompute concordance 1,000 times) would directly test whether the observed concordance structure is meaningful. The structural decomposition already suggests this would show significance, but it would strengthen the statistical argument. (Moderate impact)
-
Expand BacDive matching via chemical identifiers: The WoM compound table includes
inchi_key,pubchem_id, andsmiles_stringcolumns, though they appear NULL for FW300-N2E3 observations. If populated for other organisms, these could increase the WoM-BacDive overlap beyond 8 metabolites, which is the main bottleneck for concordance power. Noted as a future direction in the REPORT. (Nice-to-have, future work) -
Present the supplementary table more prominently: NB03 cell
clk3s5uhbyigenerates an excellent 58-row summary table of all metabolites with cross-database status. Consider including a condensed version in the REPORT.md Results section (currently it's only in the notebook output). (Nice-to-have --- improves report completeness) -
Consider the deferred NB04 pathway analysis: Mapping the 231 tryptophan fitness genes to specific GapMind pathway steps (biosynthesis vs. catabolism vs. regulatory) would substantially strengthen the overflow metabolism interpretation. While correctly deferred with rationale, this represents the most impactful next step. (Future work, already documented)
This review was generated by an AI system. It should be treated as advisory input, not a definitive assessment.
Visualizations

Concordance Summary

Consistency Heatmap

Fitness Hits Per Metabolite