Contamination Gradient vs Functional Potential in ENIGMA Communities
CompletedResearch Question
Do high-contamination Oak Ridge groundwater communities show enrichment for taxa with higher inferred stress-related functional potential compared with low-contamination communities?
Research Plan
Hypothesis
- H0: After controlling for sampling depth and site structure, contamination level is not associated with inferred stress-related functional potential in community composition.
- H1: Higher contamination is associated with higher inferred stress-related functional potential in community composition.
Approach
- Build a contamination-indexed sample set from ENIGMA geochemistry.
- Build community composition matrices for overlapping samples.
- Map ENIGMA taxa to BERDL pangenome taxa (NCBI taxid and/or normalized taxon name).
- Summarize stress-relevant functional annotation signals per taxon.
- Aggregate functional potential per sample and test association with contamination.
- Run sensitivity checks for mapping ambiguity and low-abundance taxa.
Revision History
- v1 (2026-02-15): Initial plan for ENIGMA contamination-functional potential project.
- v2 (2026-02-15): Added Phase C notebook scaffold (
01_enigma_extraction_qc.ipynb,02_taxonomy_bridge_functional_features.ipynb,03_contamination_functional_models.ipynb) with extraction/QC, taxonomy bridge, and modeling workflow placeholders. - v3 (2026-02-15): Replaced placeholders with concrete extraction, taxonomy-bridge, feature engineering, and modeling code; standardized on built-in
get_spark_session()for BERDL notebook execution. - v4 (2026-02-15): Executed NB01-NB03 end-to-end; optimized NB01 to aggregate ASV counts in Spark before collection (avoids driver OOM) and updated NB03 to use
scipy.stats.linregressdue environment-levelstatsmodelsimport failure. - v5 (2026-02-16): Added strict-vs-relaxed mapping-mode sensitivity analysis and diagnostic visualizations (contamination distribution, mapping coverage); re-executed NB02-NB03 and refreshed outputs.
- v6 (2026-02-16): Added
species_proxy_unique_genussensitivity mode (unique genus->single GTDB clade) to approximate higher-resolution bridging under genus-only ENIGMA taxonomy; re-executed NB02-NB03 and refreshed outputs. - v7 (2026-02-16): Addressed high-priority review rigor items in NB03: predeclared confirmatory vs exploratory analyses, added global BH-FDR q-values across all reported p-values, and added community-fraction robustness models/stratified tests from
sdt_community_name; re-executed NB03 and refreshed outputs. - v8 (2026-02-16): Addressed medium-priority robustness suggestions in NB03: added bootstrap confidence intervals for key effect sizes, contamination-index variant sensitivity (
composite,uranium_only,top3_var_metals,pca1_metals), mapped-coverage decile diagnostics, and explicit model-family sample-count tables; re-executed NB03 and refreshed outputs.
Overview
This project uses ENIGMA CORAL field data to test whether contamination gradients (uranium and co-occurring metals) are associated with shifts in inferred community functional potential. Functional potential is estimated by linking ENIGMA taxa to BERDL pangenome annotations and aggregating stress-relevant functional signals (COG defense/mobilome and related categories) at the site level. The analysis targets community-level ecological filtering rather than gene-level causality.
ENIGMA taxonomy in ddt_brick0000454 is currently available through Genus (no species/strain rows), so higher-resolution analysis is implemented as a species-proxy sensitivity mode using only uniquely resolved genus-to-clade mappings.
Key Findings
Multiplicity and sample-size context (primary panel)
Model-family sample counts from data/model_family_sample_counts.tsv frame how much data each analysis used:
| Mode | Base Spearman n | Adj+Cov n | Adj+Fraction n | High-coverage subset n |
|---|---|---|---|---|
relaxed_all_clades |
108 | 108 | 212 (sample-fraction rows) | 68 |
strict_single_clade |
108 | 108 | 212 (sample-fraction rows) | 68 |
species_proxy_unique_genus |
108 | 108 | 212 (sample-fraction rows) | 1 |
This keeps exploratory interpretation calibrated: coverage-aware models increase effective rows (fraction-level), while species-proxy high-coverage tests are underpowered (n=1).
(Notebook: 03_contamination_functional_models.ipynb; data: model_family_sample_counts.tsv)
Confirmatory Spearman tests remain null with confidence intervals and global FDR
Predeclared confirmatory tests (site_defense_score Spearman vs contamination in genus-level modes) remained non-significant:
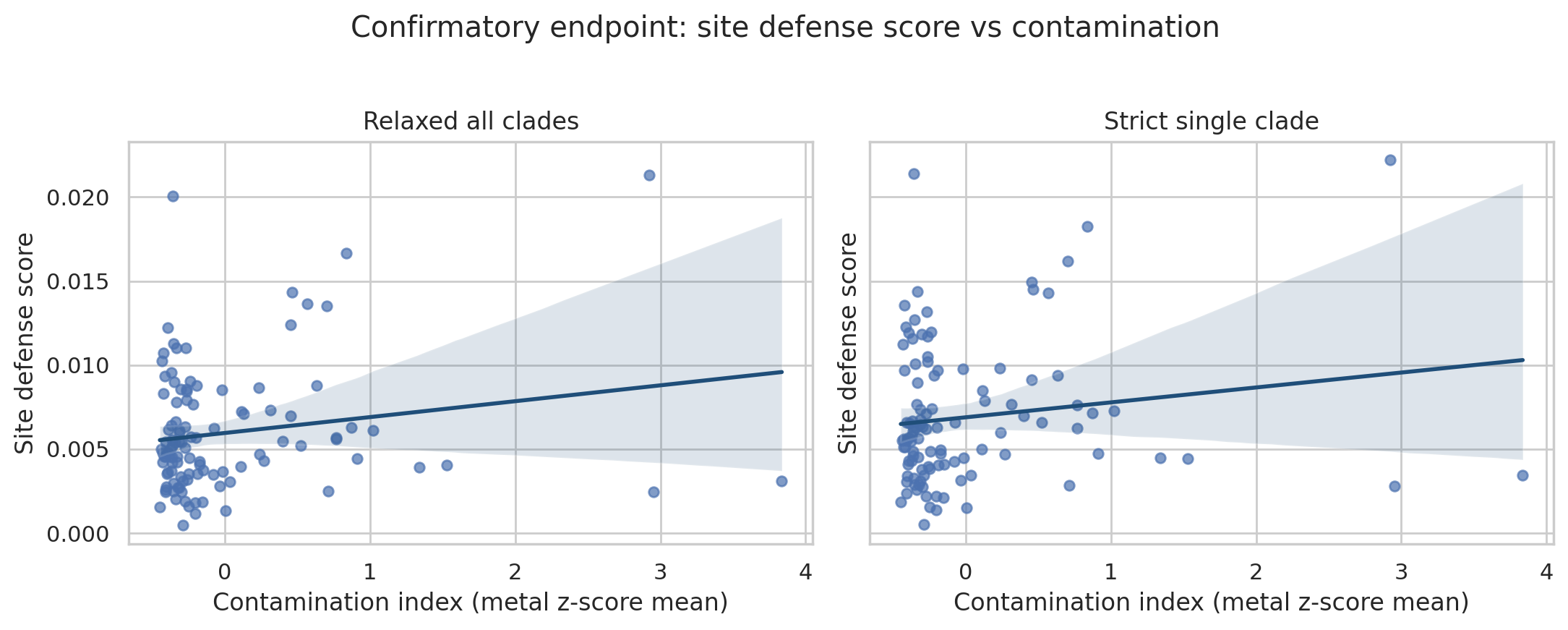
relaxed_all_clades: rho = 0.0587, 95% bootstrap CI [-0.128, 0.250], Spearman p = 0.546, FDR q = 0.862strict_single_clade: rho = 0.0682, 95% bootstrap CI [-0.111, 0.253], Spearman p = 0.483, FDR q = 0.849
Compact summary table (data/confirmatory_family_summary.tsv), where Spearman columns are confirmatory and adjusted-model columns are exploratory sensitivity analyses:
| Mode | Endpoint | n | Confirmatory Spearman rho (95% CI) | Confirmatory Spearman p / q | Exploratory Adj+Cov beta (95% CI) | Exploratory Adj+Cov p / q |
|---|---|---|---|---|---|---|
relaxed_all_clades |
site_defense_score |
108 | 0.0587 [-0.128, 0.250] | 0.546 / 0.862 | 0.000751 [0.000224, 0.001779] | 0.000398 / 0.0462 |
strict_single_clade |
site_defense_score |
108 | 0.0682 [-0.111, 0.253] | 0.483 / 0.849 | 0.000640 [0.000169, 0.001538] | 0.00354 / 0.130 |
(Notebook: 03_contamination_functional_models.ipynb)
Exploratory defense signal remains strongest in coverage-aware models
Exploratory sensitivity models still show strongest positive defense associations in coverage-aware adjustments:
- Coverage-adjusted OLS (contamination + depth + latitude + longitude + mapped_abundance_fraction):
- relaxed_all_clades: beta = 0.000751, 95% bootstrap CI [0.000224, 0.001779], p = 0.000398, FDR q = 0.0462
- strict_single_clade: beta = 0.000640, 95% bootstrap CI [0.000169, 0.001538], p = 0.00354, FDR q = 0.130
- Fraction-aware adjusted model (contamination + mapped_abundance_fraction + C(community_fraction_type)):
- relaxed_all_clades: beta = 0.000607, 95% bootstrap CI [0.000213, 0.001221], p = 0.00144, FDR q = 0.0838
- strict_single_clade: beta = 0.000566, 95% bootstrap CI [0.000214, 0.001113], p = 0.00548, FDR q = 0.130
- High-coverage subset (mapped_abundance_fraction >= 0.25):
- site_defense_score: Spearman p = 0.0207 (relaxed), 0.00980 (strict)
- After global FDR: q = 0.301 (relaxed), 0.189 (strict)
Most non-defense outcomes remained non-significant in these sensitivity tests, with one exploratory exception:
- strict_single_clade, high-coverage subset (mapped_abundance_fraction >= 0.25), site_stress_score: Spearman rho = 0.2489, p = 0.0407.
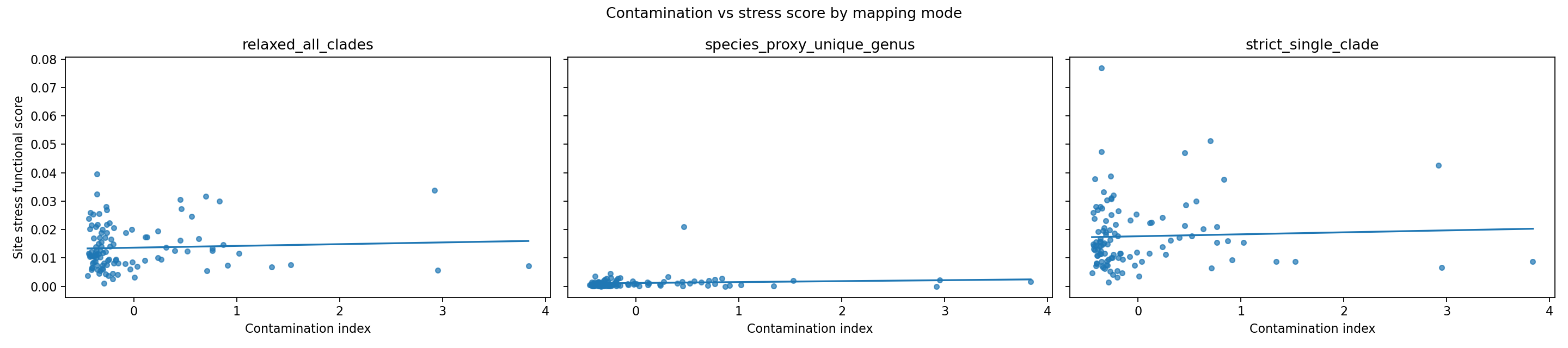
Community-fraction robustness does not show strong within-fraction monotonic signal
To address potential confounding from collapsing multiple community fractions, NB03 now retains community_fraction_type parsed from sdt_community_name and runs fraction-aware robustness analyses.
- Fraction-aware adjusted models use
n = 212sample-fraction rows (106 for0.2_micron_filter, 106 for10_micron_filter). - Within-fraction Spearman tests for defense were non-significant:
relaxed_all_clades: p = 0.767 (0.2micron), p = 0.898 (10micron)strict_single_clade: p = 0.780 (0.2micron), p = 0.793 (10micron)
This supports the conclusion that strong monotonic defense signal is not robustly reproducible inside individual fraction strata.
(Notebook: 03_contamination_functional_models.ipynb)
Contamination-index sensitivity does not change confirmatory outcome
Confirmatory endpoint was re-tested under four contamination index variants:
- Composite all-metals (current primary index)
- Uranium-only
- Top-3 variance metals
- PCA first component of metal z-scores
Across both confirmatory mapping modes, all index variants remained non-significant after FDR (q = 0.546 across the 8 confirmatory-variant tests), including uranium-only.
(Notebook: 03_contamination_functional_models.ipynb; data: contamination_index_sensitivity.tsv)
Species-proxy resolution sensitivity is limited by mapped coverage
To approximate higher taxonomic resolution despite ENIGMA taxonomy stopping at genus, NB02 added species_proxy_unique_genus mode (only genera mapping to exactly one GTDB species clade).
- Unique-clade genera: 150 (vs 530 mapped genera total)
- Ambiguous multi-clade genera: 380
- Mean mapped abundance fraction in species-proxy mode: 0.031 (vs 0.343 in strict/relaxed)
site_defense_scoretrend is positive but not significant: rho = 0.169, Spearman p = 0.081- No high-coverage subset test was feasible at the existing threshold (
mapped_abundance_fraction >= 0.25) due low retained coverage.
This suggests that with current ENIGMA taxonomy granularity, stricter taxonomic resolution sharply reduces analyzable signal and does not strengthen inference yet.
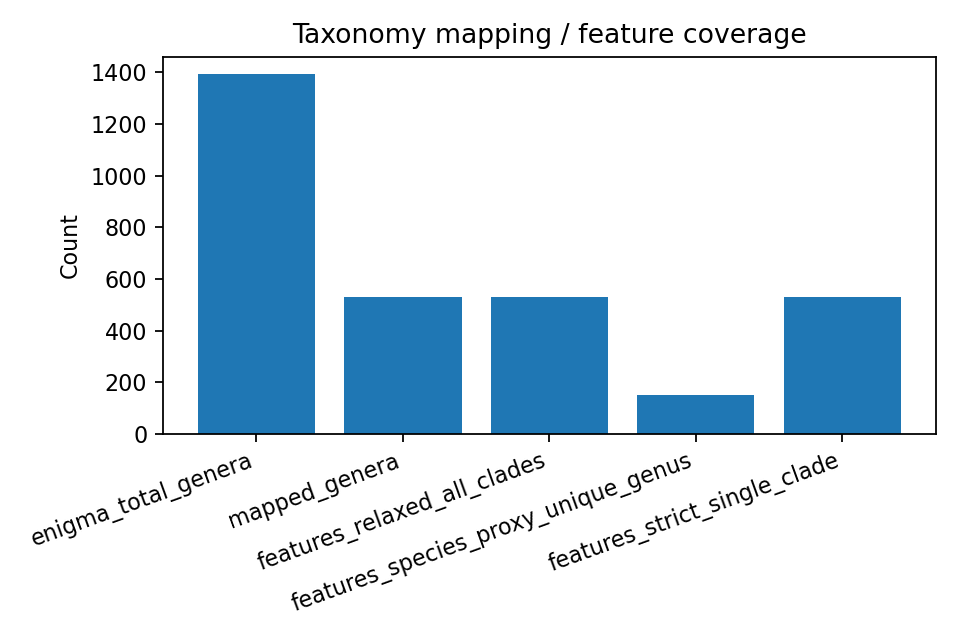
Coverage diagnostics from upstream outputs:
- 1,392 ENIGMA genera observed
- 530 mapped genera, 862 unmapped genera
- Of mapped genera, 150 were unique-clade (species-proxy resolvable) and 380 were multi-clade ambiguous
- Bridge table rows: 8,242 (many-to-many genus-to-clade expansion)
- Functional feature rows: 3,630 across 3 mapping modes (strict_single_clade, relaxed_all_clades, species_proxy_unique_genus)
- Mean mapped abundance fraction: 0.343 in strict/relaxed modes (range 0.031 to 0.854) and 0.031 in species-proxy mode (range 0.0002 to 0.543)
Bridge-quality diagnostics now exported explicitly:
- Summary metrics (data/bridge_quality_summary.tsv): mapped genera 530, unmapped 862, unique-clade 150, ambiguous multi-clade 380.
- Clade-count distribution (data/bridge_clade_count_distribution.tsv): long right tail; max clades per genus = 433.
- Top ambiguous genera (data/bridge_top_ambiguous_genera.tsv): pseudomonas (433), streptomyces (378), prevotella (358), streptococcus (214), mycobacterium (186).
(Notebook: 02_taxonomy_bridge_functional_features.ipynb; data: taxon_bridge.tsv, taxon_functional_features.tsv)
Contamination index was broad but right-skewed
NB03 built contamination index from 8 metal columns (arsenic, cadmium, chromium, copper, lead, nickel, uranium, zinc) using per-metal log1p z-scoring and row-wise mean.
- n = 108 samples
- min = -0.448, max = 3.836
- median = -0.271, IQR = [-0.363, 0.053]
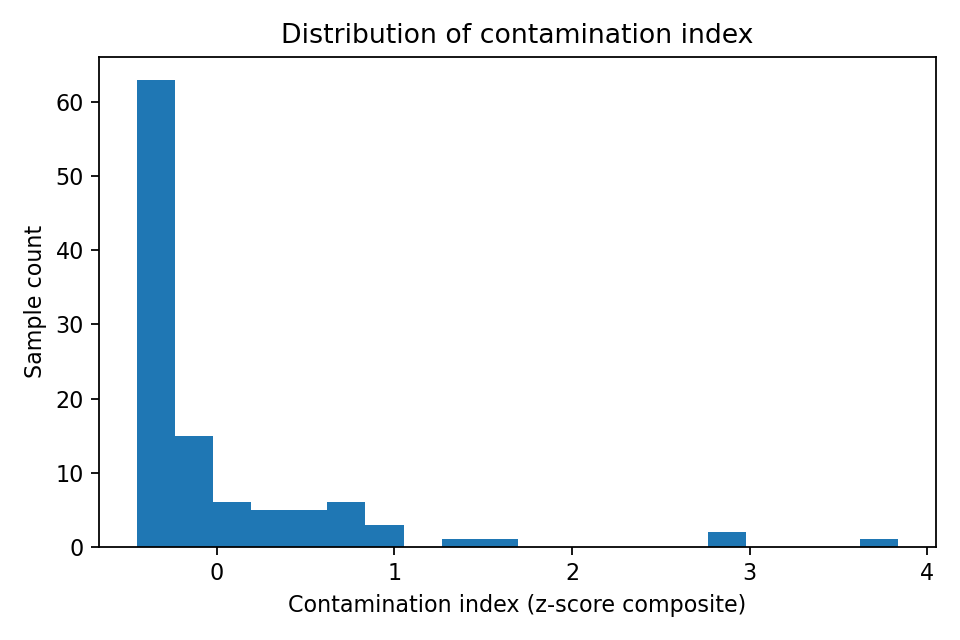
(Notebook: 03_contamination_functional_models.ipynb)
Results
NB01 extracted and QC-filtered overlap data to 108 samples with both geochemistry and community composition:
- Geochemistry matrix shape: (108, 49)
- Community taxon rows: 41,711
- Distinct communities: 212
- Distinct genera: 1,392
NB02 built a genus-normalized bridge to pangenome clades:
- GTDB species rows parsed for genus mapping: 27,690
- Mapped genera: 530
- Unmapped genera: 862
- Strict clades: 530, relaxed clades: 7,380
- Species-proxy clades (unique genus->single clade): 150
- Functional features now computed independently for strict, relaxed, and species-proxy mapping modes.
NB03 produced:
- Site functional score rows: 324 (108 samples x 3 mapping modes)
- Model result rows: 12 (4 outcomes x 3 mapping modes)
- Expanded model diagnostics in model_results.tsv: confirmatory/exploratory labels, bootstrap CIs for key effect sizes, adjusted coordinates/depth models, coverage-adjusted models, site-structure models (location_prefix), fraction-aware models (community_fraction_type), within-fraction correlations, high-coverage subset correlations, and global BH-FDR q-values across all reported p-values.
- Additional robustness tables:
- data/contamination_index_sensitivity.tsv (8 confirmatory variant rows)
- data/mapped_coverage_deciles.tsv (30 decile strata rows)
- data/model_family_sample_counts.tsv (96 model-family count rows)
Overall, confirmatory evidence remains null and robust to index definition; exploratory coverage-aware defense associations persist with positive effect estimates, but are attenuated under global multiple-testing control.
Interpretation
Within this ENIGMA subset, contamination gradients did not translate into a robust community-level shift in inferred stress-related functional potential when features are aggregated at genus resolution from broad COG categories. This is compatible with contamination-driven taxonomic turnover that is functionally redundant or too fine-scale (species/strain/pathway level) for the present mapping.
Literature Context
- Directionally aligned with Hemme et al. (2015; PMID: 26583008), which reported strong contamination-linked taxonomic restructuring in Oak Ridge groundwater with reduced functional breadth in stressed communities. Our confirmatory null Spearman results at genus-aggregated score level are consistent with contamination effects that do not collapse into one robust monotonic community-wide proxy.
- Also consistent with Fan et al. (2025), who reported pronounced compositional shifts but only modest functional-diversity decline in a mixed waste-contaminated Oak Ridge aquifer. This matches the pattern here: ecological differentiation is detectable, while broad functional summaries show weaker confirmatory monotonic response.
- Relative to Carlson et al. (2019; PMID: 30523276), this project extends from taxa-abundance association to reproducible bridge-aware functional sensitivity modeling (strict, relaxed, and species-proxy modes), quantifying where inference remains coverage-limited.
- The bridge/annotation stack depends on GTDB and eggNOG conventions (Parks et al. 2022; Cantalapiedra et al. 2021); at genus-level aggregation, broad COG-style signals can dilute pathway-specific metal-response effects.
Novel Contribution
This project contributes a reproducible ENIGMA-to-BERDL functional inference workflow with independent strict-vs-relaxed feature construction, mapped-coverage diagnostics, a species-proxy unique-clade sensitivity mode, and multi-tier modeling beyond basic univariate association tests.
Limitations
- ENIGMA taxonomy table
ddt_brick0000454currently provides labels throughGenus(no species/strain labels), so true species-level bridge testing is not directly possible in this dataset slice. - Genus-level mapping may mask strain-level adaptation.
- 862/1,392 observed genera were unmapped to current pangenome bridge.
- COG-fraction proxies are coarse summaries, not curated resistance pathways.
- Exploratory sensitivity-model significance is concentrated in defense and depends on coverage/covariate specification; broader effect across outcomes is not yet established.
- Site structure is represented by coarse
location_prefixeffects rather than full hierarchical/random-effects modeling.
Future Directions
- Replace COG-fraction proxies with curated metal-stress gene sets and pathway-level summaries.
- Increase taxonomic resolution where possible (species/strain-level bridge from metagenomic or higher-resolution ENIGMA taxonomy) and quantify gains over genus and species-proxy modes.
- Fit adjusted models with explicit covariates (depth, location cluster, sampling date) and compositional modeling controls.
- Investigate unmapped genera contribution to contamination gradients and bridge expansion opportunities.
- Add mixed-effects or hierarchical site models (well/location-level random effects) to test whether defense associations persist under richer structure control.
Data
Sources
| Collection | Tables Used | Purpose |
|---|---|---|
enigma_coral |
ddt_brick0000010, ddt_brick0000459, ddt_brick0000454, sdt_sample, sdt_community, sdt_location |
Geochemistry, community composition, taxonomy linkage, and sample metadata |
kbase_ke_pangenome |
gtdb_species_clade, pangenome, gene_cluster, eggnog_mapper_annotations |
Taxonomy bridge and COG-derived functional feature construction |
Generated Data
| File | Rows | Description |
|---|---|---|
data/geochemistry_sample_matrix.tsv |
108 | Per-sample geochemistry matrix used for contamination index |
data/community_taxon_counts.tsv |
41,711 | Sample-community-genus abundance table |
data/sample_location_metadata.tsv |
108 | Location/depth metadata for overlapping samples |
data/taxon_bridge.tsv |
8,242 | Genus-to-GTDB species-clade bridge with mapping tier |
data/taxon_functional_features.tsv |
3,630 | Per-genus functional proxy features across three mapping modes |
data/site_functional_scores.tsv |
324 | Site-level functional scores by mapping mode |
data/model_results.tsv |
12 | Confirmatory/exploratory labels, Spearman/permutation, adjusted model coefficients with bootstrap CIs, fraction-aware/coverage/site sensitivities, and global FDR q-values |
data/contamination_index_sensitivity.tsv |
8 | Confirmatory endpoint results across contamination index variants |
data/mapped_coverage_deciles.tsv |
30 | Defense association diagnostics by mapped-coverage decile and mapping mode |
data/model_family_sample_counts.tsv |
96 | Sample-count table for each modeling family and endpoint |
data/confirmatory_family_summary.tsv |
2 | Compact confirmatory endpoint summary with effect sizes, CIs, p, q, and n |
data/bridge_quality_summary.tsv |
6 | Bridge-level mapped/unmapped/ambiguity summary metrics |
data/bridge_clade_count_distribution.tsv |
71 | Distribution of species-clade counts per mapped genus |
data/bridge_top_ambiguous_genera.tsv |
20 | Most ambiguous mapped genera by species-clade multiplicity |
References
- Carlson HK et al. (2019). The selective pressures on the microbial community in a metal-contaminated aquifer. ISME J 13:937-949. PMID: 30523276.
- Hemme CL et al. (2010). Metagenomic insights into evolution of a heavy metal-contaminated groundwater microbial community. ISME J 4:660-672. DOI: 10.1038/ismej.2009.154.
- Hemme CL et al. (2015). Comparative metagenomics reveals impact of contaminants on groundwater microbiomes. Front Microbiol 6:1205. PMID: 26583008.
- Fan Y et al. (2025). Modest functional diversity decline and pronounced composition shifts of microbial communities in a mixed waste-contaminated aquifer. Microbiome 13:106. DOI: 10.1186/s40168-025-02105-x.
- Parks DH et al. (2022). GTDB: an ongoing census of bacterial and archaeal diversity through a phylogenetically consistent, rank normalized and complete genome-based taxonomy. Nucleic Acids Res 50:D785-D794.
- Cantalapiedra CP et al. (2021). eggNOG-mapper v2: Functional annotation, orthology assignments, and domain prediction at the metagenomic scale. Mol Biol Evol 38:5825-5829.
Discoveries
Primary contamination-functional tests are null, but defense signal appears in coverage-aware sensitivity models
February 2026In the ENIGMA contamination-functional project (108 overlap samples), primary univariate associations were non-significant across defense, stress, metabolism, and mobilome outcomes in both mapping modes. However, after adding mapped-coverage-aware sensitivity models, contamination-defense associatio
Read more →Independent strict vs relaxed feature construction materially changes mode-specific outputs
February 2026When strict and relaxed mapping modes were computed independently in NB02 (instead of reusing strict outputs), 1,140 of 1,590 genus-feature pairs differed between modes (~71.7%), and site stress scores differed for all 108 samples. Strict-vs-relaxed sensitivity is therefore analytically meaningful a
Read more →enigma_coral.ddt_brick0000454 currently provides taxonomy through Genus (no species/strain rows). A species-proxy mode (species_proxy_unique_genus) that keeps only unique genus->single GTDB clade mappings retained 150 genera, but mapped abundance collapsed to mean 0.031 (vs 0.343 in strict/rel
Confirmatory defense tests remain null after global FDR; strongest signal remains exploratory and coverage-dependent
February 2026After adding explicit confirmatory vs exploratory labels in NB03 and applying global BH-FDR across all reported p-values, confirmatory defense Spearman tests in genus-level modes remained null (p=0.546/0.483; q=0.862/0.849). The strongest surviving signal was exploratory: relaxed coverage-adjusted d
Read more →Re-testing confirmatory defense endpoint under four index constructions (all-metals composite, uranium-only, top-3 variance metals, PCA-PC1) did not produce significant confirmatory associations after FDR correction (all q=0.546 across 8 confirmatory-variant tests). This indicates the null confirmat
Read more →Data Collections
Review
Summary
This project is methodologically strong and reproducible, with clear confirmatory vs exploratory framing, complete staged notebooks (01 extraction, 02 bridge/features, 03 modeling), and all expected data/figure artifacts present. I did not find a critical implementation bug in the current code or outputs. The main remaining risk is interpretability drift where readers may still over-weight exploratory adjusted-model significance relative to the null confirmatory endpoint, plus minor notebook hygiene noise from retained comment-only code cells.
Methodology
The research question and hypothesis are clearly defined in README.md and RESEARCH_PLAN.md, and the approach is testable with explicit sensitivity analyses. Data provenance is clear across ENIGMA and kbase_ke_pangenome sources, and notebook sequencing is coherent.
Reproducibility is good:
- README.md has a concrete ## Reproduction section with Spark/local separation, execution order, and runtime expectations.
- requirements.txt is present.
- Notebooks contain saved outputs (NB01: 7/7 code cells with outputs; NB02: 6/6; NB03: 5/7, with 2 comment-only code cells).
- Artifacts are complete and synchronized with README expectations (data/: 14 TSV files, figures/: 4 PNG files).
Pitfall handling from docs/pitfalls.md is largely correct:
- explicit numeric casting in extraction/modeling (notebooks/01_enigma_extraction_qc.ipynb cell 3, cell 4; notebooks/03_contamination_functional_models.ipynb cells 2-4)
- Spark-first aggregation before collection (notebooks/01_enigma_extraction_qc.ipynb cell 4)
- correct eggNOG join key (notebooks/02_taxonomy_bridge_functional_features.ipynb cell 7, gc.gene_cluster_id = e.query_name)
Code Quality
Notebook organization is clear and defensive. In NB03, status labeling for insufficient/constant/invalid conditions is implemented and global BH-FDR is applied across reported p-values (notebooks/03_contamination_functional_models.ipynb cell 5). SQL in NB01/NB02 is consistent with documented schema usage.
I did not find a reproducible logic error that would invalidate current outputs. The dominant technical risk remains inferential rather than computational: genus-to-clade ambiguity is high (data/bridge_top_ambiguous_genera.tsv, data/bridge_clade_count_distribution.tsv) and species-proxy coverage is low (data/site_functional_scores.tsv, mapped abundance fraction near zero for many rows).
Findings Assessment
Conclusions in REPORT.md are supported by outputs: confirmatory defense Spearman tests are null in primary mapping modes (data/model_results.tsv, data/confirmatory_family_summary.tsv), contamination-index sensitivity remains null after FDR (data/contamination_index_sensitivity.tsv), and species-proxy mode is coverage-limited (data/model_family_sample_counts.tsv, high-coverage subset n=1).
Limitations are explicitly acknowledged and generally aligned with evidence. The principal communication risk is that statistically significant adjusted exploratory estimates in the same summary context as confirmatory results can be over-interpreted by non-technical readers.
Suggestions
- [Medium] In
REPORT.md, further separate confirmatory and exploratory inferential statements where adjusted-model q-values are shown (for example, keep confirmatory claims tied only to predeclared Spearman endpoint rows fromdata/confirmatory_family_summary.tsv). - [Low] Convert NB03 comment-only code cells to markdown (or remove them) to reduce execution-trace noise and avoid confusion during reruns (
notebooks/03_contamination_functional_models.ipynb, cells 6-7). - [Low] Add one brief sentence in
README.mdorREPORT.mdwarning that species-proxy high-coverage subset analyses are not interpretable at current sample size (n=1indata/model_family_sample_counts.tsv).
This review was generated by an AI system. It should be treated as advisory input, not a definitive assessment.
Visualizations

Confirmatory Defense Vs Contamination

Contamination Index Distribution

Contamination Vs Functional Score

Mapping Coverage By Mode
Notebooks
Data Files
| Filename | Size |
|---|---|
bridge_clade_count_distribution.tsv |
0.4 KB |
bridge_quality_summary.tsv |
0.2 KB |
bridge_top_ambiguous_genera.tsv |
0.4 KB |
community_taxon_counts.tsv |
3433.0 KB |
confirmatory_family_summary.tsv |
0.6 KB |
contamination_index_sensitivity.tsv |
1.2 KB |
geochemistry_sample_matrix.tsv |
103.6 KB |
mapped_coverage_deciles.tsv |
3.2 KB |
model_family_sample_counts.tsv |
6.6 KB |
model_results.tsv |
11.1 KB |
sample_location_metadata.tsv |
14.5 KB |
site_functional_scores.tsv |
51.3 KB |
taxon_bridge.tsv |
1058.4 KB |
taxon_functional_features.tsv |
314.9 KB |