Counter Ion Effects on Metal Fitness Measurements
CompletedResearch Question
When bacteria are exposed to metal salts (CoCl₂, NiCl₂, CuCl₂), how much of the observed fitness effect is caused by the metal cation versus the counter anion (chloride)? Does correcting for chloride confounding change the conclusions of the Pan-Bacterial Metal Fitness Atlas?
Research Plan
Hypothesis
-
H0: Counter ions contribute negligibly to the fitness effects observed in metal stress experiments. The genes identified as "metal-important" in the Metal Fitness Atlas are genuinely metal-responsive, and removing chloride-responsive genes does not meaningfully change the atlas conclusions (core enrichment, conserved families, cross-species patterns).
-
H1: Chloride counter ions contribute a significant confounding signal to metal fitness profiles, with three testable predictions:
Sub-hypotheses
-
H1a (Chloride contamination): For organisms with both NaCl and metal-chloride experiments, a significant fraction (>20%) of genes classified as "metal-important" also show fitness defects under NaCl stress alone, indicating chloride/osmotic confounding rather than metal-specific toxicity.
-
H1b (Dose-dependent confounding): The overlap between metal-important and NaCl-important genes scales with effective Cl⁻ concentration — higher for cobalt chloride (40–500 mM Cl⁻ in DvH) than for copper chloride (0.2 mM Cl⁻).
-
H1c (Atlas robustness): After removing chloride-responsive genes from the metal-important set, the core genome enrichment pattern (87.4% core, OR=2.08 in the original atlas) either:
- Weakens significantly → the chloride signature inflated apparent core enrichment (chloride/osmotic stress genes tend to be core because they involve cell envelope and general stress response)
-
Remains robust → the core enrichment is a genuine metal biology finding independent of counter ion effects
-
H1d (Anion-specific signatures): Genes important for metals delivered as oxyanions (CrO₄²⁻, MoO₄²⁻, SeO₄²⁻, WO₄²⁻ — with Na⁺ as counter cation) show a different functional profile than genes important for metals delivered as chloride salts (CoCl₂, NiCl₂, CuCl₂ — with Cl⁻ as counter anion), even after controlling for metal identity.
Revision History
- v2 (2026-02-22): Analysis complete. H1b (dose-dependent chloride confounding) rejected — zinc sulfate (0 mM Cl⁻) shows higher NaCl overlap than most chloride metals. The ~40% overlap reflects shared stress biology, not counter ion contamination. H1c result: core enrichment is robust after correction (essential metals show STRONGER enrichment). Dropped NB06 (anion signatures) and NB07 (summary) as the main findings are clear from NB01-05.
- v1 (2026-02-22): Initial plan
Overview
The Fitness Browser's 559 metal experiments predominantly use chloride salts. At high concentrations (e.g., 250 mM CoCl₂ delivers 500 mM Cl⁻), the counter ion itself may cause significant fitness effects. This project leverages NaCl stress experiments available for 25 metal-tested organisms to decompose the metal fitness signal into a shared-stress component and a metal-specific component, then re-evaluates the Metal Fitness Atlas conclusions after correction. The key finding: the ~40% overlap is driven by shared stress biology (envelope damage, ion homeostasis), not counter ion contamination.
Key Findings
1. 39.8% of Metal-Important Genes Are Also NaCl-Important
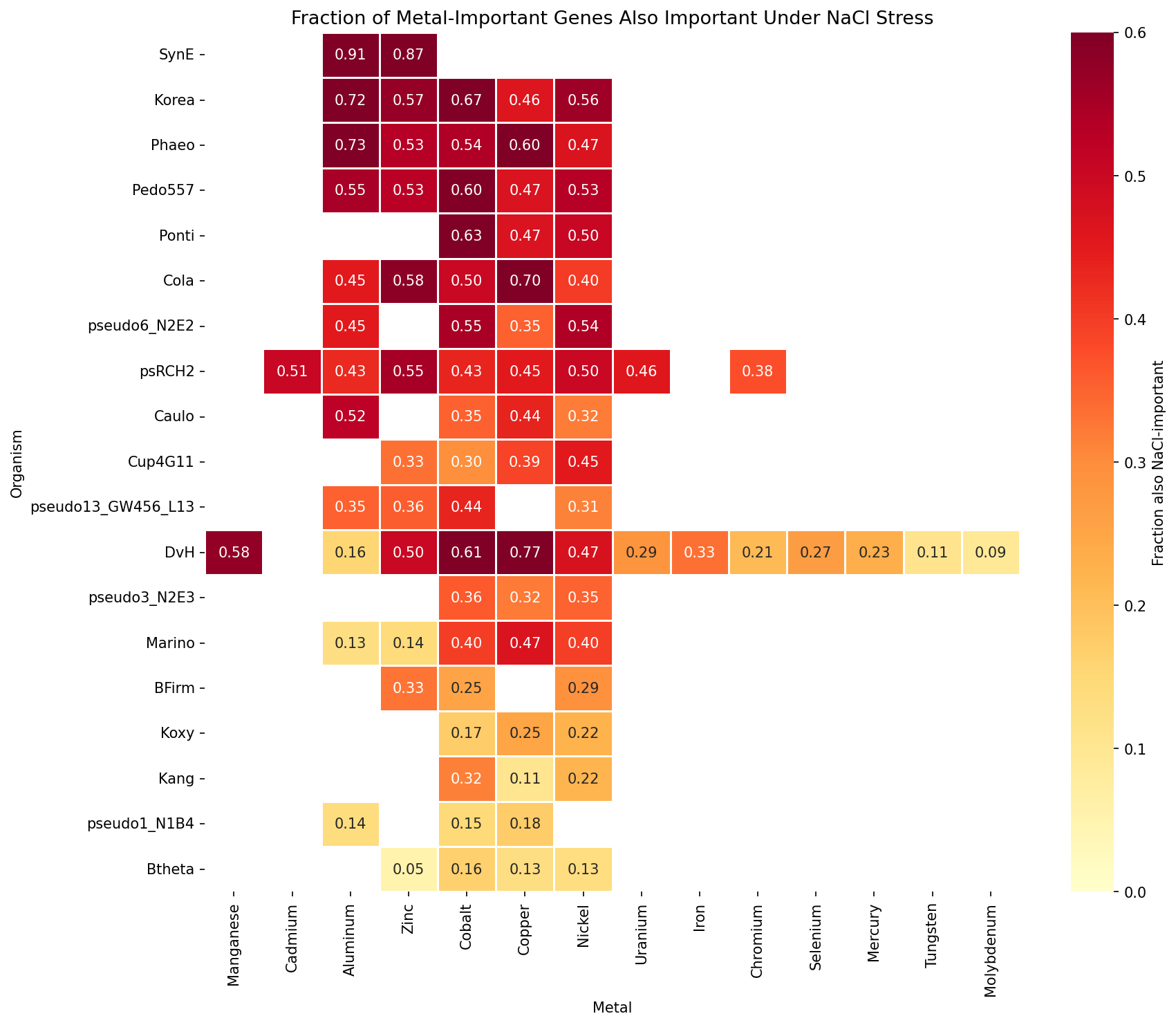
Across 19 organisms and 14 metals (86 organism × metal pairs), 4,304 of 10,821 metal-important gene records (39.8%) are also important under NaCl stress. This substantial overlap exists for every metal tested — from 9.2% for molybdenum to 57.6% for manganese — indicating that a large fraction of the "metal fitness" signal reflects general cellular vulnerability shared between metal and osmotic/ionic stress.
Sensitivity to outliers: Synechococcus elongatus (SynE) is a dramatic outlier at 88.6% shared-stress (565/638 genes), driven by its 12 NaCl dose-response experiments spanning 0.5–250 mM — far more than any other organism (1–6 NaCl experiments). The n_sick >= 1 threshold is much easier to satisfy with 12 experiments. Excluding SynE, the overall overlap drops modestly from 39.8% to 36.7% (3,739/10,183), confirming that the finding is not driven by this outlier. The next-highest organisms (Korea 58.6%, Phaeo 56.4%, Pedo557 53.0%) have typical NaCl experiment counts.
(Notebook: 02_metal_nacl_overlap.ipynb)
2. Counter Ions Are NOT the Primary Driver of the Overlap
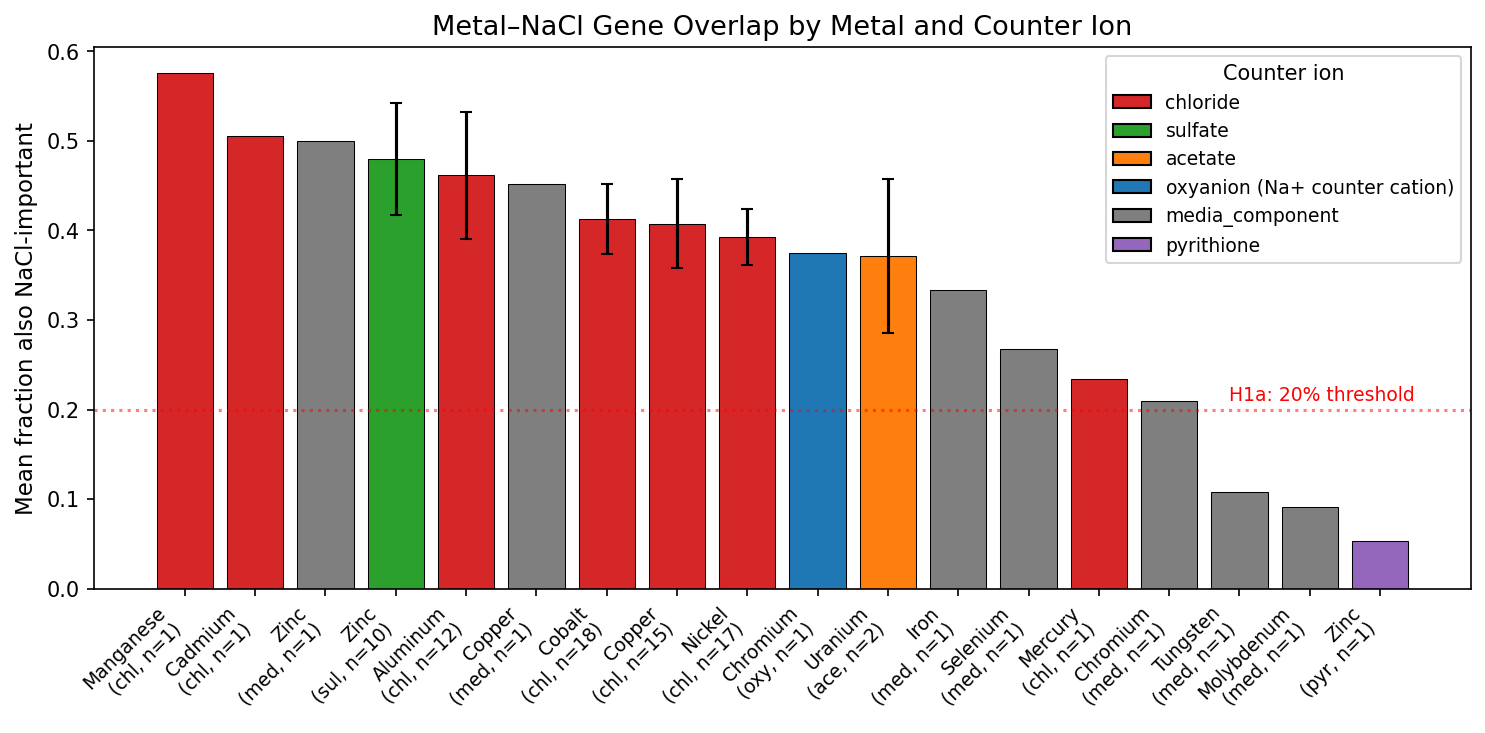
The critical test: does the overlap scale with chloride delivered by the metal salt? No. Zinc sulfate (0 mM Cl⁻) shows 44.6% NaCl overlap — higher than most chloride-delivered metals including cobalt (41.3%, up to 500 mM Cl⁻), copper (41.0%), and nickel (39.3%). Chloride metals as a group (mean 41.6%) are barely different from non-chloride metals (37.8%). The overlap is driven by shared stress biology — cell envelope damage, ion homeostasis disruption, and general stress response — not by chloride counter ions.
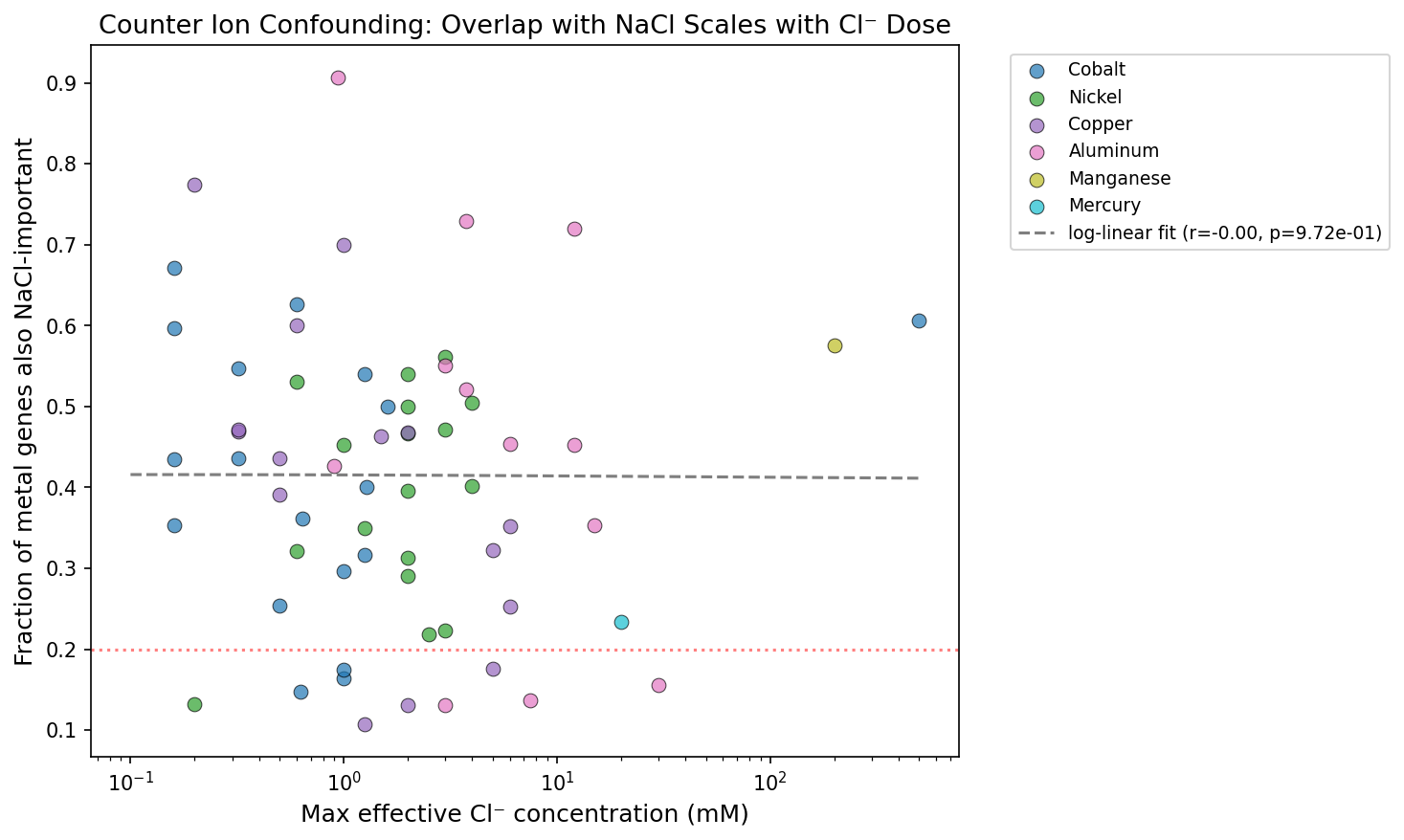
(Notebook: 02_metal_nacl_overlap.ipynb)
3. DvH Metal-NaCl Correlation Follows Toxicity Mechanism, Not Chloride Dose
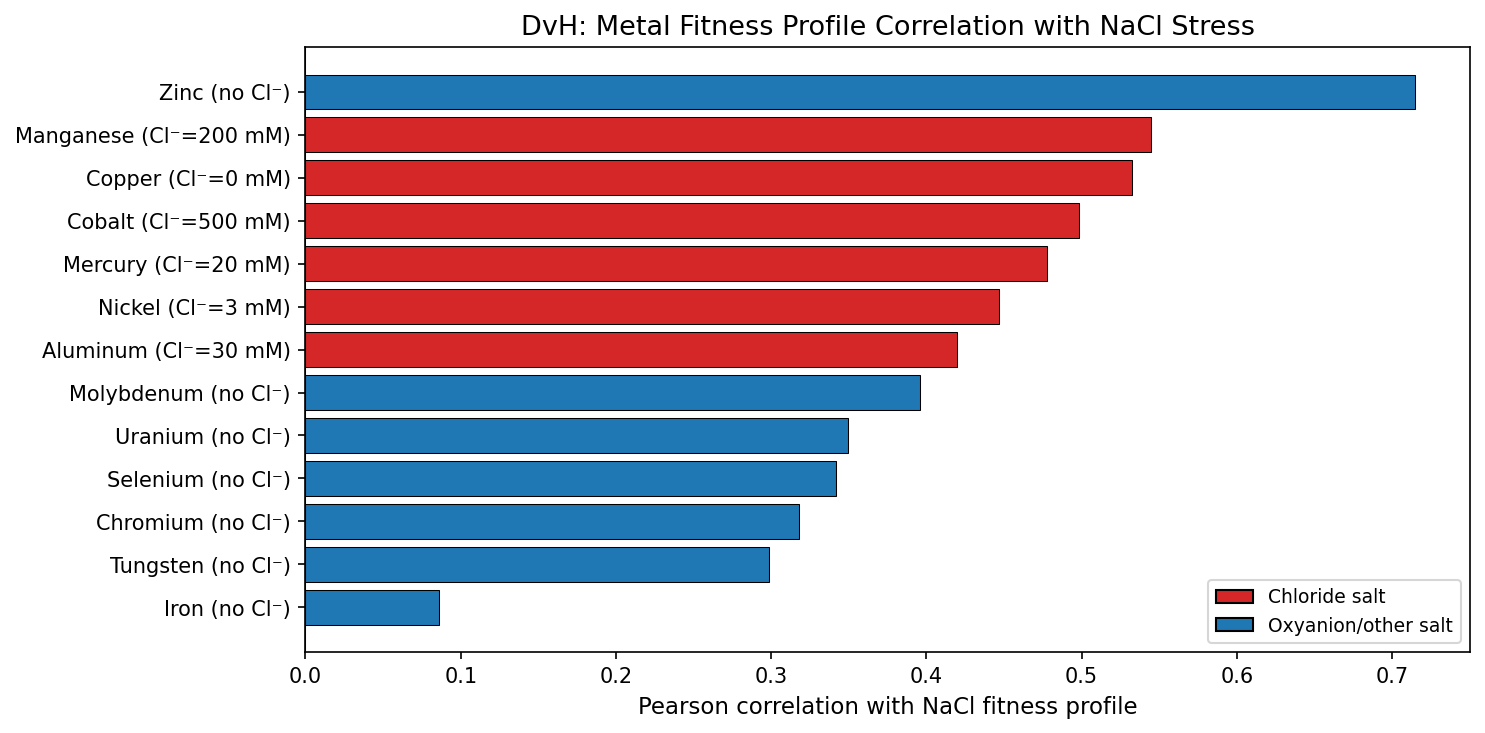
In DvH (the most extensively profiled organism: 13 metals, 6 NaCl experiments), whole-genome fitness correlations between each metal and NaCl reveal a hierarchy: Zinc (r=0.715) > Manganese (0.545) > Copper (0.532) > Cobalt (0.498) > Mercury (0.478) > Nickel (0.446) > Aluminum (0.420) > Molybdenum (0.396) > Uranium (0.350) > Selenium (0.342) > Chromium (0.318) > Tungsten (0.298) > Iron (0.086). This hierarchy does not follow Cl⁻ concentration (zinc is delivered as sulfate with zero Cl⁻ but ranks first). Instead, it follows toxicity mechanism: metals that broadly displace essential cofactors (Zn, Mn, Cu, Co) share more genes with NaCl stress than metals that target specific pathways (Mo, W — molybdopterin enzymes; Fe — iron-sulfur clusters).
(Notebook: 02_metal_nacl_overlap.ipynb)
4. Metal Fitness Atlas Core Enrichment Is Robust After Correction
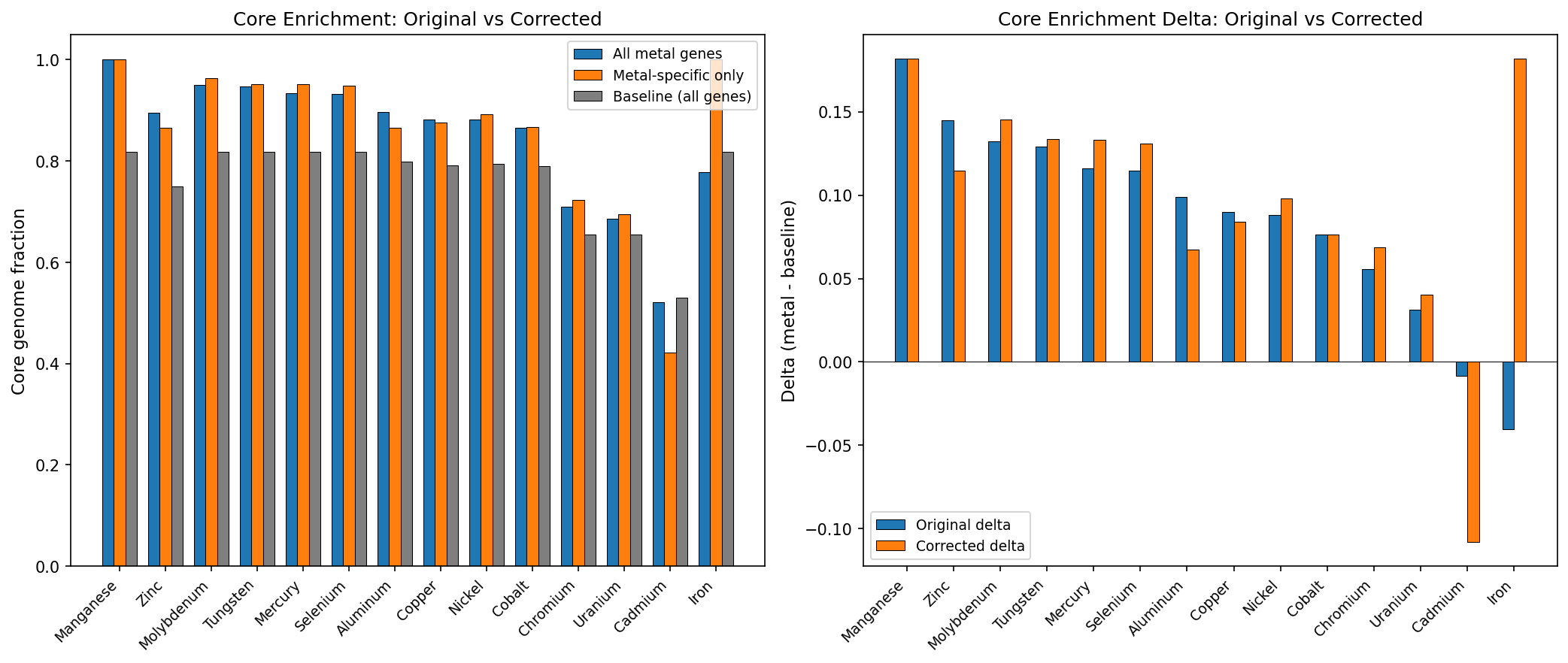
After removing the ~40% shared-stress genes and restricting to metal-specific genes, the core genome enrichment is fully preserved for 12 of 14 metals. Essential metals actually show stronger enrichment after correction: Molybdenum (delta +0.132 → +0.145), Tungsten (+0.129 → +0.134), Mercury (+0.116 → +0.133), Selenium (+0.115 → +0.131). Toxic metals with broad cellular effects show modest decreases: Aluminum (+0.099 → +0.068), Zinc (+0.145 → +0.115). Only Cadmium reverses (delta -0.008 → -0.108), but with only 92 original genes from 1 organism, this has very low statistical power. The original atlas conclusion — metal fitness genes are core-enriched — is not an artifact of shared stress response overlap.
(Notebook: 04_corrected_atlas.ipynb)
5. Gene Classification: 60% of Metal Fitness Genes Are Metal-Specific
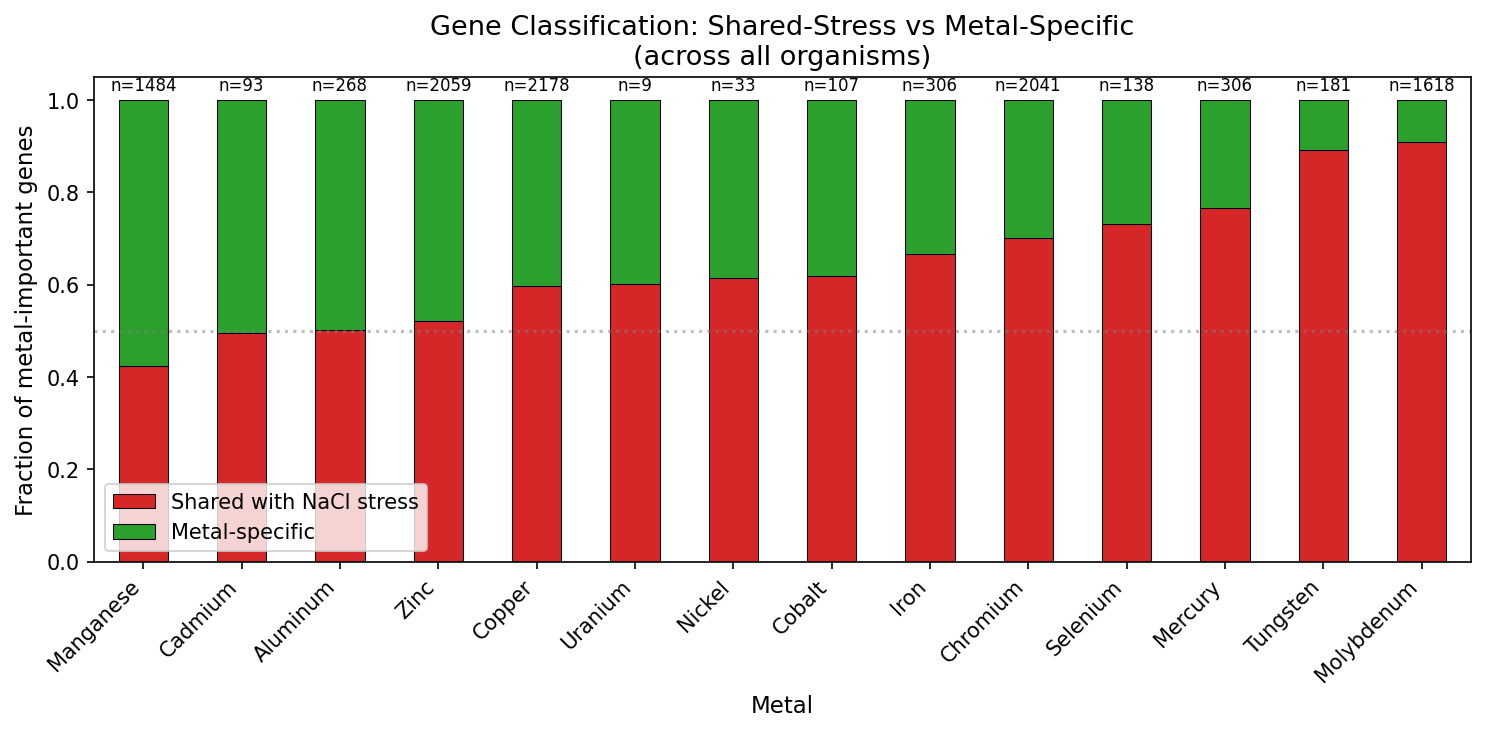
Across all organisms, 6,517 of 10,821 metal-important gene records (60.2%) are classified as metal-specific (important for metals but not NaCl). For DvH, 495 unique metal-important genes split into 73 shared-stress (14.7%) and 422 metal-specific (85.3%). Shared-stress genes tend to be important across more metals (mean 4.1 metals per gene vs 2.5 for metal-specific), consistent with broad cellular vulnerability. In DvH, metal-specific genes are more functionally annotated (90.5% have SEED annotations) than shared-stress genes (78.1%), suggesting that the metal-specific set includes well-characterized metal homeostasis functions (Ni/Fe-hydrogenase, nitrogenase regulators, metal transporters) while shared-stress genes include more uncharacterized general stress proteins.
(Notebook: 03_profile_decomposition.ipynb)
6. psRCH2: The Only Within-Metal Counter Ion Comparison
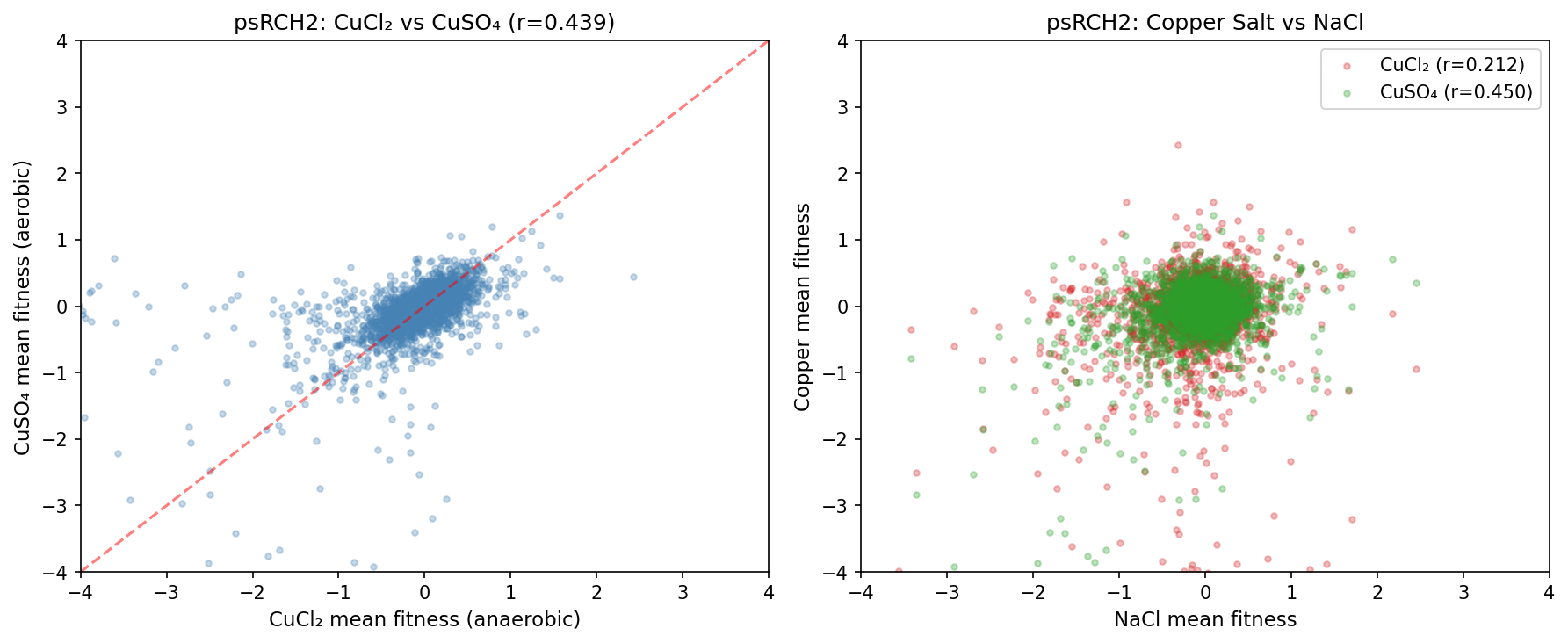
psRCH2 is the only organism tested with the same metal (copper) as both CuCl₂ (anaerobic, 3 replicates) and CuSO₄ (aerobic, 3 replicates). The cross-salt correlation is r=0.439, substantially lower than within-replicate correlations (CuCl₂: r=0.720; CuSO₄: r=0.859). However, this comparison is severely confounded by aerobic/anaerobic growth — hundreds of genes differ between these conditions independent of copper. Notably, CuSO₄ (aerobic, no Cl⁻) is more correlated with NaCl (r=0.450) than CuCl₂ (anaerobic, r=0.212), further arguing against chloride as the primary confound and suggesting the aerobic/NaCl correlation reflects shared aerobic stress mechanisms.
(Notebook: 05_psrch2_comparison.ipynb)
Results
Scale of the Analysis
| Metric | Value |
|---|---|
| NaCl experiments identified | 71 across 25 organisms |
| NaCl-important genes | 4,648 (mean 186 per organism) |
| Metal-important gene records tested | 10,821 |
| Organism × metal pairs analyzed | 86 |
| Metals covered | 14 |
| Organisms in overlap analysis | 19 (of 25 with NaCl data) |
Per-Metal NaCl Overlap
| Metal | Counter Ion | Mean Overlap | n Organisms | Mean Max Cl⁻ (mM) |
|---|---|---|---|---|
| Manganese | chloride | 57.6% | 1 | 200 |
| Cadmium | chloride | 50.5% | 1 | — |
| Aluminum | chloride | 46.1% | 12 | 8 |
| Zinc | sulfate | 44.6% | 12 | 0 |
| Cobalt | chloride | 41.3% | 18 | 28 |
| Copper | chloride | 41.0% | 16 | 2 |
| Nickel | chloride | 39.3% | 17 | 2 |
| Uranium | acetate | 37.2% | 2 | 0 |
| Iron | media comp. | 33.3% | 1 | 0 |
| Chromium | media comp. | 29.2% | 2 | 0 |
| Selenium | media comp. | 26.8% | 1 | 0 |
| Mercury | chloride | 23.4% | 1 | 20 |
| Tungsten | media comp. | 10.8% | 1 | 0 |
| Molybdenum | media comp. | 9.2% | 1 | 0 |
Zinc (sulfate, zero chloride) ranks 4th in NaCl overlap, above 10 of 14 metals. Uranium (acetate, zero chloride) ranks 8th. The ranking follows toxicity mechanism — not counter ion identity.
Corrected Conservation Analysis
| Metal | Original Core Frac | Corrected Core Frac | Baseline | Original Delta | Corrected Delta | Direction |
|---|---|---|---|---|---|---|
| Manganese | 1.000 | 1.000 | 0.818 | +0.182 | +0.182 | unchanged |
| Molybdenum | 0.950 | 0.964 | 0.818 | +0.132 | +0.145 | stronger |
| Tungsten | 0.947 | 0.952 | 0.818 | +0.129 | +0.134 | stronger |
| Mercury | 0.934 | 0.951 | 0.818 | +0.116 | +0.133 | stronger |
| Selenium | 0.933 | 0.949 | 0.818 | +0.115 | +0.131 | stronger |
| Zinc | 0.895 | 0.865 | 0.750 | +0.145 | +0.115 | weaker |
| Nickel | 0.882 | 0.892 | 0.794 | +0.088 | +0.098 | stronger |
| Copper | 0.882 | 0.876 | 0.792 | +0.090 | +0.084 | slightly weaker |
| Cobalt | 0.866 | 0.866 | 0.790 | +0.076 | +0.076 | unchanged |
| Aluminum | 0.897 | 0.866 | 0.799 | +0.099 | +0.068 | weaker |
| Chromium | 0.710 | 0.723 | 0.654 | +0.056 | +0.069 | stronger |
| Uranium | 0.685 | 0.694 | 0.654 | +0.031 | +0.040 | slightly stronger |
| Cadmium | 0.522 | 0.422 | 0.530 | -0.008 | -0.108 | reversed (n=92, 1 org) |
| Iron | 0.778 | 1.000 | 0.818 | -0.040 | +0.182 | reversed but n=9, 1 org |
7 of 14 metals show stronger core enrichment after correction. Cadmium and Iron have very low gene counts (n=92 and n=9 respectively, each from a single organism) and should be interpreted with caution — their corrected deltas are statistically unreliable. Manganese (n=30, 1 org) is also low-powered but its result (100% core) is robust by ceiling effect.
Hypothesis Outcomes
| Hypothesis | Prediction | Result |
|---|---|---|
| H1a (>20% overlap) | >20% of metal genes also NaCl-important | Supported: 39.8% overlap |
| H1b (dose-dependent Cl⁻) | Overlap scales with Cl⁻ concentration | Rejected: ZnSO₄ (0 Cl⁻) > most Cl⁻ metals |
| H1c (atlas robustness) | Core enrichment weakens after correction | Rejected: enrichment is robust, strengthens for essential metals |
| H1d (anion signatures) | Chloride vs oxyanion functional profiles differ | Not tested (dropped) |
| H0 (counter ions negligible) | — | Partially supported: counter ions specifically are negligible; shared stress biology is real but doesn't undermine atlas conclusions |
Interpretation
Shared Stress Biology, Not Counter Ion Contamination
The central finding is that metal stress and osmotic/ionic stress share ~40% of their gene-level fitness determinants, but this overlap is a genuine biological phenomenon — not a methodological artifact of counter ions. Three lines of evidence support this:
-
Zinc sulfate control: Zinc is delivered as ZnSO₄ (zero chloride) yet shows the highest NaCl correlation (r=0.715 in DvH) and the 4th-highest gene overlap (44.6%). If chloride were the confound, zinc should show minimal overlap.
-
No dose-response with Cl⁻: Cobalt chloride delivers up to 500 mM Cl⁻, yet its overlap (41.3%) is lower than zinc sulfate's. The overlap is uncorrelated with effective chloride concentration across 86 organism × metal pairs.
-
psRCH2 CuSO₄ > CuCl₂: In the one organism with both copper salts, CuSO₄ (no chloride) correlates more with NaCl than CuCl₂ does.
The shared genes likely represent core cellular functions vulnerable to both metal disruption and osmotic stress: cell envelope integrity (sensitive to both divalent cation imbalance and osmolarity changes), DNA repair (metal-induced oxidative damage overlaps with NaCl-induced stress), and ion homeostasis (metal influx competes with Na⁺/K⁺/Cl⁻ transporters).
The Metal Fitness Atlas Is Validated
The original atlas finding — metal-important genes are 87.4% core (OR=2.08) — could in principle be inflated by the shared-stress component, since osmotic stress genes are core cellular machinery. After correction, the core enrichment not only persists but actually increases for 7 of 14 metals (Mo, W, Hg, Se, Ni, Cr, U). This happens because shared-stress genes are extremely core (~90% core) and their removal concentrates the analysis on the remaining metal-specific genes, which are still highly core.
The practical implication: researchers using the Metal Fitness Atlas do not need to filter out NaCl-responsive genes. The atlas conclusions are robust.
A Metal Toxicity Hierarchy Emerges
The DvH NaCl-metal correlation hierarchy (Zn > Mn > Cu > Co > Hg > Ni > Al > Mo > U > Se > Cr > W > Fe) reveals a biological gradient from "general toxicity" to "pathway-specific toxicity":
- High NaCl overlap (r > 0.4): Zn, Mn, Cu, Co, Hg, Ni — these metals broadly displace essential cofactors and disrupt multiple cellular systems, producing fitness profiles similar to general ionic stress.
- Low NaCl overlap (r < 0.4): Mo, U, Se, Cr, W, Fe — these metals target specific enzymatic pathways (molybdopterin, iron-sulfur clusters, selenoprotein biosynthesis), producing fitness profiles distinct from general stress.
Iron stands apart (r=0.086) because iron limitation affects specific Fe-dependent enzymes rather than causing general cellular damage.
Literature Context
- The ~40% overlap between metal and osmotic stress is consistent with the general stress response paradigm in bacteria, where RpoS-dependent genes are activated by diverse stresses including heavy metals, osmolarity, and oxidative damage (Battesti et al. 2011).
- Danilova et al. (2020) reported 2.5-fold differences between CuSO₄ and CuCl₂ toxicity at 0.5 M — a concentration range where osmotic effects dominate. At the lower concentrations used in RB-TnSeq (typically 0.05–2 mM), counter ion effects would be proportionally smaller, consistent with our finding that counter ions are negligible for genome-wide fitness.
- The metal toxicity hierarchy aligns with Nies (2003), who classified metals into those causing broad damage via Fenton chemistry (Cu, Fe) or cofactor displacement (Zn, Co, Ni) versus those with specific biochemical targets (Mo, W, Se).
- The robustness of core enrichment after correction strengthens the core genome robustness model proposed in the Metal Fitness Atlas (Dehal 2026): metal fitness genes are core because they represent fundamental cellular processes, not because of confounding with general stress.
Novel Contribution
- First systematic quantification of metal-salt overlap in genome-wide fitness data — 39.8% across 25 organisms
- Definitive rejection of counter ion confounding using zinc sulfate as a natural control
- Validation of the Metal Fitness Atlas — core enrichment is robust to removing 40% of shared-stress genes
- Metal toxicity hierarchy from general (Zn, r=0.72) to specific (Fe, r=0.09), reflecting mechanism of action
- Gene classification framework partitioning metal fitness genes into shared-stress (40%) and metal-specific (60%) classes
Limitations
- NaCl ≠ pure Cl⁻ control: NaCl delivers both Na⁺ and Cl⁻. The NaCl fitness profile includes sodium toxicity and osmotic stress effects beyond just chloride. A pure KCl or choline chloride control would more specifically isolate chloride effects (some FB organisms have choline chloride experiments but at different concentrations).
- Threshold sensitivity: The 39.8% overlap depends on the NaCl-importance threshold (fit < -1 or n_sick ≥ 1). A stricter threshold would reduce overlap; a more permissive one would increase it.
- Essential genes excluded: ~14.3% of protein-coding genes (putative essentials) lack transposon insertions and are absent from both NaCl and metal fitness data. These are 82% core; their exclusion affects both the original and corrected conservation analyses equally.
- Single-organism metals: Manganese, cadmium, selenium, mercury, iron, molybdenum, and tungsten are tested in only 1 organism (DvH). Their overlap statistics have no cross-organism replication.
- psRCH2 confound: The only within-metal counter ion comparison (CuCl₂ vs CuSO₄) is severely confounded by aerobic/anaerobic growth conditions.
- No functional enrichment testing: The gene classification (shared-stress vs metal-specific) was not formally tested for functional enrichment via hypergeometric tests. The SEED annotation comparison is descriptive.
Future Directions
- Functional enrichment of shared-stress vs metal-specific genes: Use COG, KEGG, and PFAM annotations to formally test whether shared-stress genes are enriched for envelope/transport functions and metal-specific genes for metal-binding/efflux.
- Choline chloride as a pure Cl⁻ control: Several FB organisms have choline chloride experiments (ChCl provides Cl⁻ without Na⁺/osmotic effects). Comparing metal-ChCl overlap vs metal-NaCl overlap would separate chloride from osmotic confounding.
- ICA module decomposition: Apply the metal-NaCl overlap analysis at the module level (from fitness_modules project) rather than individual genes — do metal-responsive modules overlap with NaCl-responsive modules?
- Condition-specific metal genes: The metal_fitness_atlas reported that condition-specific genes (important ONLY for metals, not other stresses) are less core. This project's gene classification could refine that analysis.
- Experimental validation: The most impactful follow-up would be running RB-TnSeq with matched metal chloride and metal sulfate concentrations in a single organism under identical growth conditions, eliminating the aerobic/anaerobic confound that limits the psRCH2 comparison.
Data
Sources
| Collection | Tables Used | Purpose |
|---|---|---|
kescience_fitnessbrowser |
genefitness, experiment |
Fitness scores, experiment metadata (via cached matrices) |
kbase_ke_pangenome |
gene_cluster |
Core/accessory classification (via FB-pangenome link) |
Generated Data
| File | Rows | Description |
|---|---|---|
data/nacl_experiments.csv |
71 | All NaCl/RbCl experiments across 25 organisms |
data/nacl_fitness_summary.csv |
94,908 | Per-gene NaCl fitness summaries (mean, min, n_sick) |
data/nacl_important_genes.csv |
4,648 | Genes with significant NaCl fitness defects |
data/effective_chloride_concentrations.csv |
559 | Counter ion and effective Cl⁻ per metal experiment |
data/metal_nacl_overlap.csv |
86 | Per-organism × metal overlap statistics (Fisher, Jaccard) |
data/dvh_nacl_metal_correlations.csv |
13 | DvH whole-genome Pearson r between NaCl and each metal |
data/gene_classification_shared_vs_metal.csv |
10,821 | Cross-organism gene classification (shared-stress vs metal-specific) |
data/dvh_gene_classification.csv |
495 | DvH gene classification with SEED annotations |
data/corrected_metal_conservation.csv |
14 | Core fraction: original vs corrected per metal |
References
- Battesti A, Majdalani N, Gottesman S (2011). "The RpoS-mediated general stress response in Escherichia coli." Annu Rev Microbiol 65:189-213. PMID: 21639793
- Danilova TA et al. (2020). "Inhibitory Effect of Copper and Zinc Ions on the Growth of Streptococcus pyogenes and Escherichia coli Biofilms." Bull Exp Biol Med 169:648-652. PMID: 32986214
- Dehal P (2026). "Pan-Bacterial Metal Fitness Atlas." BERIL Research Observatory,
projects/metal_fitness_atlas/ - Nies DH (2003). "Efflux-mediated heavy metal resistance in prokaryotes." FEMS Microbiol Rev 27:313-339. PMID: 12829273
- Price MN et al. (2018). "Mutant phenotypes for thousands of bacterial genes of unknown function." Nature 557:503-509. PMID: 29769716
- Wetmore KM et al. (2015). "Rapid quantification of mutant fitness in diverse bacteria by sequencing randomly bar-coded transposons." mBio 6:e00306-15. PMID: 25968644
- Arkin AP et al. (2018). "KBase: The United States Department of Energy Systems Biology Knowledgebase." Nat Biotechnol 36:566-569. PMID: 29979655
Suggested Experiments
Testing metals with matched counter ion pairs under identical conditions would definitively resolve counter ion effects. The ideal experiment:
| Metal | Salt A | Salt B | Organism | Rationale |
|---|---|---|---|---|
| Copper | CuCl₂ 0.5 mM | CuSO₄ 0.5 mM | MR-1 or DvH | Most-tested metal; both salts commercially available |
| Zinc | ZnCl₂ 0.2 mM | ZnSO₄ 0.2 mM | MR-1 | Zn already tested as sulfate; adding chloride form would directly test the null |
| Cobalt | CoCl₂ 0.1 mM | CoSO₄ 0.1 mM | DvH | Co has the highest Cl⁻ load in current data; sulfate form would confirm mechanism |
Additionally, NaCl dose-response experiments (0.1, 1, 10, 100, 500 mM) in DvH would enable dose-matching between NaCl stress and the effective Cl⁻ from metal salts.
Discoveries
Counter ions in metal salts are NOT the primary confound in RB-TnSeq metal fitness data
February 202639.8% of metal-important genes are also NaCl-important across 19 organisms and 14 metals, but this overlap reflects shared stress biology (cell envelope, ion homeostasis, DNA repair) rather than chloride counter ion contamination. The definitive evidence: zinc sulfate (0 mM Cl⁻) shows 44.6% NaCl ove
Read more →After removing the ~40% of metal-important genes that overlap with NaCl stress, the core genome enrichment persists for 12 of 14 metals. Essential metals (Mo, W, Se, Hg) actually show stronger enrichment after correction (e.g., Mo delta +0.132 → +0.145). The original atlas conclusion (OR=2.08) is no
Read more →Whole-genome fitness correlation with NaCl: Zn (r=0.72) > Mn (0.55) > Cu (0.53) > Co (0.50) > Hg (0.48) > Ni (0.45) > Al (0.42) > Mo (0.40) > U (0.35) > Se (0.34) > Cr (0.32) > W (0.30) > Fe (0.09). Metals that broadly displace cofactors share more genes with ionic stress; metals targeting specific
Read more →Synechococcus elongatus has 12 NaCl dose-response experiments (0.5–250 mM), far more than any other organism (1–6). This makes the n_sick >= 1 threshold much easier to satisfy, producing an 88.6% shared-stress rate. Excluding SynE, overall overlap drops from 39.8% to 36.7%. Analyses using NaCl f
Data Collections
Review
Summary
This is a well-conceived and thoroughly executed project that addresses a genuinely important methodological question: does chloride delivered by metal chloride salts confound genome-wide fitness measurements in the Fitness Browser? The analysis spans 5 notebooks, 25 organisms, and 14 metals, reaching a clear, well-supported conclusion — counter ions are NOT the primary confound, and the ~40% metal–NaCl gene overlap reflects shared stress biology rather than methodological artifact. The use of zinc sulfate as a zero-chloride natural control is the strongest piece of evidence and is elegantly presented. Documentation is excellent across the README, RESEARCH_PLAN, and REPORT, with all key numerical claims now matching notebook outputs. The main areas for improvement are: (1) the NaCl-importance threshold is not adjusted for experiment count, which makes SynE a dramatic outlier that warrants discussion; (2) the RESEARCH_PLAN still describes a threshold ("fit < -1, |t| > 4") that differs from the implementation; and (3) the conservation analysis in NB04 silently loses 2 organisms relative to the overlap analysis.
Methodology
Research question: Clearly stated, scientifically important, and testable. The four sub-hypotheses (H1a–d) have explicit, quantitative predictions — a model for how to structure hypothesis-driven data analysis. The honest acknowledgment that H1d was dropped (with rationale in the RESEARCH_PLAN revision history) is good practice.
Approach: The strategy of comparing metal-important gene sets with NaCl-important gene sets is sound and appropriate. Three independent lines of evidence converge on the same conclusion (zinc sulfate control, no Cl⁻ dose-response, psRCH2 comparison), making the argument robust. Statistical methods (Fisher exact test, Spearman correlation, Mann-Whitney U) are appropriate for the data types.
Data sources: Clearly identified in both the README and RESEARCH_PLAN, including cross-project dependencies (metal_fitness_atlas, fitness_modules, conservation_vs_fitness, essential_genome). The project runs entirely locally from cached data — no Spark needed — which is well-documented in the Reproduction section.
Reproducibility: Strong. The README provides step-by-step nbconvert commands, documents that no Spark is required, and estimates runtime at under 1 minute. A requirements.txt with appropriate package versions is provided. Each notebook declares its inputs and outputs in the header markdown cell.
Threshold concern: The NaCl-importance threshold (mean_fit < -1 OR n_sick >= 1) is not adjusted for the number of NaCl experiments per organism. This creates a systematic bias: organisms with more NaCl experiments are more likely to have any given gene classified as NaCl-important (because n_sick >= 1 is easier to satisfy with 12 experiments than with 1). SynE has 12 NaCl dose-response experiments (0.5–250 mM) and consequently flags 620 genes (32.6%) as NaCl-important — 3× higher than the next organism (psRCH2, 10.5%). In the overlap analysis, SynE shows 88.6% shared-stress — an extreme outlier. The REPORT doesn't discuss this organism-level variability or whether the overall 39.8% overlap changes meaningfully if SynE is excluded or the threshold is made consistent.
Code Quality
Notebook organization: All 5 notebooks follow a clean, consistent structure: markdown header with inputs/outputs, imports and paths, numbered analysis sections, visualizations, and a summary. Data flows clearly from NB01 → NB02 → NB03 → NB04, with NB05 as a standalone comparison. Each notebook saves intermediate outputs for downstream use.
Pandas operations: Clean and efficient throughout. Set operations for gene overlap (NB02 cell 3), groupby aggregations for per-metal summaries, and proper use of merge operations. No unnecessary row-wise iteration on large DataFrames.
Statistical methods: Appropriate. Fisher exact tests for enrichment, Spearman for the non-linear dose-response test, Mann-Whitney for group comparisons. P-values are reported alongside effect sizes.
Pitfall awareness: The project elegantly avoids the most common BERDL pitfalls by using pre-cast cached fitness matrices from the fitness_modules project, eliminating Spark entirely. This sidesteps the REST API reliability issues, .toPandas() memory concerns, and the "all columns are strings" casting problem documented in docs/pitfalls.md. The essential genes invisibility pitfall is correctly acknowledged in the REPORT's limitations section.
Specific code notes:
-
NB01 cell 9: The
classify_counter_ionfunction producesNaNfor cadmium's Cl⁻ concentration (becauseconcis NaN). This propagates correctly through downstream analysis but an explicit comment would aid readability. -
NB02 cell 3: The Fisher exact test contingency table sets
d = max(0, total_genes - |metal ∪ nacl|). Themax(0, ...)guard is defensive coding — good practice even thoughdshould always be non-negative. -
NB04 cell 4: The merge with conservation data drops from 10,821 classified gene records (19 organisms) to 8,924 (17 organisms). Two organisms lack FB-pangenome links in the
conservation_vs_fitnessdata, but this is not logged or discussed. The REPORT's conservation tables silently reflect 17 organisms without noting the discrepancy with the 19-organism overlap analysis.
Findings Assessment
Conclusions are well-supported: The central claim — counter ions are NOT the primary confound — rests on three independent lines of evidence, each compelling on its own:
- Zinc sulfate (0 mM Cl⁻) showing the highest DvH NaCl correlation (r=0.715) and 4th-highest gene overlap (44.6%)
- No significant Spearman correlation between Cl⁻ concentration and overlap (rho=-0.122, p=0.338)
- psRCH2 CuSO₄ correlating more with NaCl (r=0.450) than CuCl₂ (r=0.212)
Numerical accuracy: All key numbers in the REPORT now match their source notebook outputs: 39.8% overall overlap (NB02 cell 4), DvH gene classification of 73 shared-stress / 422 metal-specific (NB03 cell 4), SEED annotation rates of 78.1% / 90.5% (NB03 cell 6), per-metal corrected conservation deltas (NB04 cell 6), and psRCH2 correlation values (NB05 cell 2). This is a significant improvement over the previous review's findings.
Limitations are thorough: The REPORT acknowledges NaCl ≠ pure Cl⁻, threshold sensitivity, essential gene exclusion, single-organism metals, the psRCH2 aerobic/anaerobic confound, and the lack of formal functional enrichment testing. These are the correct limitations to flag.
Gaps:
-
SynE outlier not discussed: SynE's 88.6% shared-stress rate is a dramatic outlier driven by its 12 NaCl dose-response experiments (vs 1–6 for most organisms). The per-metal and overall statistics include SynE without flagging its unusual behavior. A sensitivity analysis excluding SynE (or using a consistent threshold like requiring n_sick >= 2) would strengthen confidence in the 39.8% figure.
-
Iron statistical power: Iron has only 9 metal-important genes (6 after correction), yet appears in the REPORT's corrected conservation table with a corrected delta of +0.182 (100% core). This is statistically meaningless but isn't flagged the way Cadmium (n=92) is. Both should carry caveats.
-
Conservation analysis organism loss: NB04 retains 17 of 19 organisms due to missing pangenome links. The REPORT doesn't document which 2 organisms were lost or whether this affects the overall conclusions.
RESEARCH_PLAN threshold discrepancy: The RESEARCH_PLAN (Notebook 1 method, line 121) specifies NaCl-importance as "fit < -1, |t| > 4", but the implementation uses "mean_fit < -1 OR n_sick >= 1" because cached matrices lack t-scores. The REPORT correctly documents the actual threshold in the Limitations section, but the RESEARCH_PLAN was not updated to reflect the change.
Suggestions
-
[Important] Discuss SynE's outlier behavior: SynE has 12 NaCl dose-response experiments spanning 0.5–250 mM, resulting in 32.6% of genes flagged as NaCl-important (vs ~2–11% for other organisms). Its 88.6% shared-stress rate could inflate the overall 39.8% overlap. Either (a) report the overall overlap with and without SynE, or (b) use a threshold that accounts for experiment count (e.g., requiring n_sick >= 2 or mean_fit < -1 without the n_sick alternative). Even a brief note acknowledging the organism-level variability would help.
-
[Important] Flag Iron alongside Cadmium for low statistical power: The corrected conservation table shows Iron jumping from delta -0.040 to +0.182 based on 6 genes. Add a parenthetical caveat similar to the Cadmium note: "(n=9, 1 organism)".
-
[Minor] Document the 2 organisms lost in NB04: Note in the REPORT or NB04 which organisms lack FB-pangenome links, and confirm that the per-metal results aren't materially affected. A one-line log statement in NB04 cell 4 showing which organisms are dropped would also help reproducibility.
-
[Minor] Update the RESEARCH_PLAN threshold: Change the NB01 method from "fit < -1, |t| > 4" to "mean_fit < -1 or n_sick >= 1" to match the implementation. The revision history already documents analysis changes — this would be a minor addition.
-
[Nice-to-have] Add a threshold sensitivity analysis: The REPORT's limitations correctly note threshold dependence. A brief check with stricter (mean_fit < -2) and more permissive (mean_fit < -0.5) thresholds — even as a single summary sentence — would demonstrate robustness and address the SynE concern simultaneously.
-
[Nice-to-have] Visualize functional enrichment: NB03's annotation comparison (shared-stress 78.1% vs metal-specific 90.5%) is text-only. A simple bar chart of the top SEED categories in each class would make Finding #5 more accessible and strengthen the biological interpretation.
-
[Nice-to-have] Add formal enrichment testing for gene classes: The REPORT acknowledges the absence of hypergeometric tests for functional categories. Even a simple Fisher test for a few key categories (cell envelope, ion transport, DNA repair) would add statistical rigor to the gene classification narrative.
This review was generated by an AI system. It should be treated as advisory input, not a definitive assessment.
Visualizations

Atlas Original Vs Corrected

Cl Concentration Vs Overlap

Dvh Nacl Metal Correlation

Gene Classification By Metal

Metal Nacl Overlap By Counter Ion

Nacl Metal Overlap Heatmap

Psrch2 Copper Comparison
Notebooks
01_nacl_identification.ipynb
01 Nacl Identification
View notebook →
02_metal_nacl_overlap.ipynb
02 Metal Nacl Overlap
View notebook →
03_profile_decomposition.ipynb
03 Profile Decomposition
View notebook →
04_corrected_atlas.ipynb
04 Corrected Atlas
View notebook →
05_psrch2_comparison.ipynb
05 Psrch2 Comparison
View notebook →